HTML
-
Candida albicans is one of the most common opportunistic human fungal pathogens[1]. The rate of infection by C. albicans has increased with the improvement and use of advanced medical treatments, such as those applied to organ and bone marrow transplants, cancer, and immunodeficiency virus infections[2, 3]. The immune suppression accompanying these treatments increases a patient's susceptibility to infection. C. albicans can colonize mucosal surfaces on the oral cavity, digestive tract, and genitourinary system, leading to mucosal and systemic infections, especially in immunocompromized individuals[4]. Treatment of these infections usually includes the application of antifungal azoles, such as fluconazole (FLU) or itraconazole (ITRA); however, increased treatment and improper use of azoles have led to increased antifungal resistance[5, 6].
Several mechanisms have been found to cause azole resistance in C. albicans. One such mechanism involves overexpression of efflux pump genes. Candida drug resistance 1, CDR1, an ATP binding cassette (ABC) transporter gene, and multidrug resistance 1, MDR1, a multidrug transporter-encoding gene, can use ATP and a proton gradient, respectively, to move antifungal drugs across the cytoplasmic membrane[7]. CDR1 and MDR1 upregulation are controlled by the transcription factors TAC1 and MRR1, respectively[8]. DDR48 is a stress-associated gene and the DDR48/ddr48 heterozygote strain is susceptible to ITRA, FLU and ketoconazole[9]. Previous studies indicate that CDR1 and MDR1 are upregulated in azole-resistant C. albicans strains[6]. Azole antifungals inhibit the biosynthetic pathway of ergosterol, which is the major sterol component of the plasma membrane of fungi[10]. Ergosterol synthesis is inhibited and the fungal membrane is ultimately disrupted when azole antifungals bind to the ergosterol biosynthesis enzyme lanosterol 4-α-sterol demethylase (ERG11), which is responsible for the conversion of lanosterol to ergosterol[6]. Overexpression of ERG11 may lead to azole resistance in C. albicans[8]. Regulation of ERG11 is controlled by the fungal transcriptional regulator UPC2, a Zn2Cys6 cluster transcription factor[11]; if UPC2 is deleted, loss of ERG11 expression, which contributes to the high susceptibility of C. albicans to azole, occurs[12].
The limited selection of antifungal drug classes has contributed to the widespread emergence of resistant Candida isolates[13], especially toward the azole family[13], making candidiasis a challenging infection to treat. Possible solutions to counteract antifungal resistance include the development of new antifungal drugs, increasing the dosage of existing antifungal drugs, and treatment with combinations of antifungal drugs[14]. Compared with the increasing patient population, however, the rate of discovery of new drugs is slow, and only one new antifungal drug class has been discovered for clinical use in the past 20 years[15]. Because new drug discovery is impractical and increased drug dosage could lead to even greater resistance, treatment with combinations of antifungal drugs is likely the most promising approach[16]. The advantages of using antifungal combinations in treating resistant fungal isolates include enhanced rates of microbial killing, shortening of the duration of therapy, and possible prevention or delayed emergence of drug-resistant mutants[17, 18].
Recent reports have shown the effectiveness of antifungal drug combinations[13, 19]. Dual combinations of intravenous amphotericin B or oral FLU in combination with flucytosine are specifically mentioned in the practice guidelines for candidiasis[17]. Drug combinations studies have included azoles, such as FLU and amiodarone[20], FLU and doxycycline[21], and FLU and chloroquine[22]. FLU has also been combined with various types of non-antifungal agents such as antibacterials, calcineurin inhibitors, calcium homeostasis regulators, heat shock protein inhibitors, and traditional chinese medicines, due to the increased FLU resistance of C. albicans[23]. Indeed, many of these treatment combinations exert synergistic effects against antibiotic-resistant C. albicans isolates. The mechanisms behind these synergistic effects obtained using different compounds include increased permeability of the cell membrane[24], inhibition of the efflux of antifungal drugs[25], decreased transporter activity[26], and blockage of biofilm formation[27] Among these mechanisms, efflux is broadly recognized as a major component of resistance to antimicrobial drugs[28]. Future studies examining drug interactions to reverse fungal resistance to antimicrobial treatment will hopefully result in new approaches to eliminate drug resistance and shed light on routes toward novel antifungal drug discovery.
Cis-2-dodecenoic acid (BDSF), which is produced by Burkholderia cenocepacia, regulates the morphological transition of C. albicans and inhibits biofilm formation[29, 30]. Biofilm formation is one of the foundations of drug resistance, and BDSF is effective in enhancing antifungal treatment and decreases C. albicans cell adherence, the initial phase of biofilm formation, to plates and catheters by 4- and 25-fold, respectively[31]. Because BDSF decreases adhesion and morphological transition, the two major factors contributing to Candida pathogenicity, it should be considered a potential therapeutic agent to treat diseases caused by Candida species.
In a previous report, the signaling molecule BDSF showed efficient inhibitory effects on all clinical isolates tested, regardless whether the isolates were resistant to FLU[32]. In the present study, two C. albicans strains resistant to FLU and ITRA were chosen to investigate the effects of combining either of the antifungal drugs FLU or ITRA with BDSF and their possible mechanisms. While candidiasis has become a refractory disease with the widespread emergence of azole-resistant Candida strains, here, we show that the combination treatment of BDSF with azoles is a new innovative therapy.
-
Two C. albicans strains resistant to FLU and ITRA (strain Nos. 99 and 108) were selected from 60 clinical C. albicans isolates[32]. The laboratory strains ATCC 22019 and ATCC 6258 were used in susceptibility tests (Clinical and Laboratory Standards Institute, 2008) as controls[33]. Isolates were stored in LB broth with 15% glycerol at -80 ℃. Each strain was inoculated and subcultured on Sabouraud dextrose agar (SDA)[10] at 35 ℃ for 24 h before testing.
-
FLU and ITRA were purchased from Sigma- Aldrich (St Louis, MO, USA), and BDSF was synthesized as described previously[30]. Stock solutions of the following were prepared for subsequent dilution: FLU prepared using sterile water at 5, 120 μg/mL, ITRA prepared using dimethylsulfoxide at 1, 600 μg/mL, and BDSF prepared in equal volumes of sterile water and methanol at 30, 000 μmol/L. All stock solutions were then stored at -20 ℃. Further dilutions of the drugs were prepared by the double-dilution method in liquid medium. Cisplatin and 3-(4, 5-dimethyl thiazol-2-yl)-2, 5-diphenyl tetrazolium bromide (MTT; Sigma-Aldrich) were used to estimate the relative cytotoxicity of BDSF.
-
The minimum inhibitory concentrations (MICs) of FLU, ITRA, and BDSF against C. albicans strain Nos. 99 and 108 were tested following the broth microdilution method in RPMI 1640 medium as specified in Clinical and Laboratory Standards Institute (CLSI, formerly NCCLS) method M27-A3 (Clinical and Laboratory Standards Institute, 2008)[33] and reported by Chen et al.[34]. The final drug concentrations of ITRA and FLU were 0.03-16 μg/mL and 0.12-64 μg/mL, respectively. MICs were defined as 90% inhibition of growth when compared with controls after 24 h of inoculation and determined by spectrophotometry at 530 nm.
-
The antifungal susceptibilities of C. albicans strain Nos. 99 and 108 were also measured by the E-test diffusion method using E-test strips (AB Biodisk, Sweden). Isolates were suspended in 0.85% sterile saline, diluted to an optical density of 0.1 at 530 nm, and then plated with a sterile swab onto RPMI-glucose agar plates containing RPMI 1640 medium, 2% glucose, and 0.165 mol/L 4-morpholinepropanesulfonic acid (MOPS) under pH 7.0 condition. After complete absorption of extra moisture, E-test strips were applied to the surface, then 0.03-16 μg/mL ITRA solutions and 0.12-64 μg/mL FLU solutions from the broth dilution method was added to E-test strips, respectively. After the plates were incubated at 37 ℃ for 24 h, the MICs of the strains were read.
-
C. albicans strain Nos. 99 and 108 were used to investigate the in vitro interaction between BDSF and FLU or ITRA via the microdilution checkerboard method in 96-well plates[21]. Each strain was inoculated in RPMI 1640 buffered with 0.165 mol/L MOPS to pH 7.0 with a starting inoculum density of 105 cfu/mL. Each drug was diluted to the desired final concentration, ensuring the final concentration to be at least 2-fold of the MIC of each strain against the respective drug. Plates were inoculated for 24 h at 35 ℃. Using a spectrophotometer at 530 nm, the MICs of the drug combinations were determined as the lowest concentration showing 90% inhibition compared with the growth control[35]. Each strain was evaluated in triplicate on different days.
To assess the relation between BDSF and FLU or ITRA, the fractional inhibitory concentration index (FICI) was used. The equations for the FICI are as follows:
$$ \text{FICI = FICA + FICB }\!\!~\!\!\text{ } $$ (1) $$ \begin{align} &\text{FICA = MIC of drug A in combination / MIC of } \\ &\text{drugA alone} \\ \end{align} $$ (2) $$ \begin{align} &\text{FICB = MIC of drug B in combination / MIC of } \\ &\text{drug B alone} \\ \end{align} $$ (3) When the FICI values are ≤ 0.5, the drug interactions are classified as synergistic; when the FICI values are ≥ 4, the drug interactions are considered antagonistic. FICI values between 1 and 4 are classified as indifferent[17].
Heat maps were created using Excel 2007 (Microsoft, USA) to illustrate the inhibition percentages of C. albicans growth compared with those of the controls[14].
-
Clinical multi-azole-resistant C. albicans strains (Nos. 99 and 108) were prepared in triplicate for experiments as described previously (RPMI 1640 medium with 0.165 mol/L MOPS to pH 7.0, 1 × 105 cfu/mL starting density). Each inoculum was incubated with antifungal drugs alone (at $ {}^{1}\!\!\diagup\!\!{}_{2}\;$ MIC), BDSF alone (at $ {}^{1}\!\!\diagup\!\!{}_{2}\;$ MIC), or the combination of each antifungal drug (at $ {}^{1}\!\!\diagup\!\!{}_{2}\;$ MIC) with BDSF (at $ {}^{1}\!\!\diagup\!\!{}_{2}\;$ MIC) at 35 ℃. At 0, 2, 6, 12, 24, 36, or 48 h, an aliquot of 100 μL was extracted from each tube, serially diluted in sterile water, and then streaked and subcultured on SDA plates. The colony number was counted after incubation at 35 ℃ for 24 h to estimate log10 values. Any increase or decrease in 2log10 value after 24 h was defined respectively as synergism or antagonism compared with FLU, ITRA, or BDSF treatment alone. A change in value < 2log10 was defined as indifferent.
-
The multi-azole-resistant C. albicans strains were cultured in YPD broth (1% yeast extract, 2% peptone, 1% dextrose) at 35 ℃. After overnight culture, the cells were obtained and diluted in PBS solution (1 × 106 cfu/mL). A mouse candidiasis model was constructed via lateral tail vein injection of C. albicans (dose: 1 × 105 cells) for 4 h. Subsequently, infected female BALB/c mice were randomly divided into four groups (five mice per group) and individually treated with PBS, FLU [1 mg/(kg·day)], BDSF [10 mg/(kg·day)], or FLU [0.5 mg/(kg·day)] + BDSF [10 mg/(kg·day)] for 4 days. The mice were sacrificed, and their kidneys were harvested aseptically and weighed after the end of treatment. To measure the organ fungal burden, kidneys were homogenized in 5 mL of sterile PBS solution, and the homogenates were 10-fold serially diluted and inoculated on SDA plates at 35 ℃ for 24 h for further colony counting. All animal experiments were performed in accordance with the guidelines of the National Institutes of Health on animal care, and the protocol was approved by the School of Pharmaceutical Science, Nanjing Tech University.
-
Primers for the genes DDR48, ERG1, ERG11, UPC2, CDR1, MDR1, TAC1, and MRR1, as well as those for the housekeeping gene ACT1, are listed in Table 1. The azole-resistant C. albicans isolates were subcultured on SDA at 35 ℃ for 24 h and then inoculated into YPD broth with OD = 0.1 (read at 530 nm). The inocula were diluted in YPD broth and cultured with FLU or ITRA ($ {}^{1}\!\!\diagup\!\!{}_{2}\;$ MIC), BDSF ($ {}^{1}\!\!\diagup\!\!{}_{2}\;$ MIC), or their combinations ($ {}^{1}\!\!\diagup\!\!{}_{2}\;$ MIC of the antifungal drug and $ {}^{1}\!\!\diagup\!\!{}_{2}\;$ MIC of BDSF) at 35 ℃ for 12 h with shaking at 220 rpm. Total RNA was extracted using the EASYspin Yeast RNA Fast Extraction Kit (Qiagen Co.). After treatment with DNase, the total RNA was reversed-transcribed with the RevertAid First Strand cDNA Synthesis Kit (Roche) according to the manufacturer's instruction. SYBR Green-based qRT-PCR was performed with a Stratagene Mx3000p system using the following temperature parameters: 95 ℃ for 10 min, followed by 40 cycles of 95 ℃ for 15 s, 57 ℃ for 25 s, and 72 ℃ for 25 s. ACT1 was used as the internal control gene to normalize the qRT-PCR results and measure changes in expression of the chosen gene. Experiments were performed thrice with different extractions of total RNA samples. Data are presented as fold-changes in mRNA expression level normalized against the housekeeping gene ACT1.
Gene Name PCR Product (bps) Forward Reverse DDR48 108 TTCGGTAAAGACGACGACAAAGA GCCAAATGAAGAGGATCCATAAGA ERG1 70 AGAATGTGTTAACGGGCCAATT ATGGTTGAATAACAACATTGGGAAT ERG11 131 GAATCCCTGAAACCAAT AGCAGCAGTATCCCATC UPC2 71 ATTATGGATTCGTTAGCCAATGC CATGCAGAAGCTGGCAAAGA CDR1 179 GCATTGATGGGAGCATCTGG GTAGTGGTTTCCAAATGAACGTCTT MDR1 280 GGAGTTAAACATTTCACCCTCGTT ATGCACCAGAAGCAGTAGTAGCAG MRR1 230 AACGCTGGTTATGGGTGA TTTGCTGTTGGGCTTCTT TAC1 71 TGGCAATGTATTTAGCAGATGAGG TGCTTGAACTGAGGTGAATTTTG ACT1 152 TTGACCAAACCACTTTCAACTC AGAAGATGGAGCCAAAGCAG Table 1. Primers Used in This Study
-
MTT assay was employed using the normal foreskin fibroblast cell line to estimate the cytotoxicity of BDSF[31]. Cisplatin, a cytotoxic drug, was used as the positive control for these tests. Cells (5 × 103) prepared in 0.2 mL of Dulbecco's modified Eagle's medium (DMEM) containing 10% fetal bovine serum were seeded onto a 96-well plate and treated with BDSF or cisplatin at 37 ℃ for 72 h. Next, the medium was removed from the wells, and 25 μL of MTT solution (5 mg/mL in PBS) and 75 μL of DMEM were added to each well. The plate was then incubated at 37 ℃ for another 2 h. Following the second incubation, an aliquot of 0.1 mL of MTT lysis buffer was added to each well, and the plate was incubated once more at 37 ℃ for 4 h. Finally, optical densities (A570) were determined on a BioRad plate reader at 570 nm. This experiment was replicated thrice, and the percentage of cell relative viability was estimated as: (A570 of the treated sample) / (A570 of the untreated sample) × 100. Statistical methods were carried out using SPSS (version 17.0; SPSS Inc., USA) and statistical significance was defined as P < 0.05.
Candida Strains and Growth Conditions
Antifungal Preparation
Minimum Inhibitory Concentration (MIC) Determination by Broth Dilution
MIC Determination by E-test
Checkerboard Testing and Heat Map Plotting
Time-kill Curves
In vivo Antifungal Synergy
Quantitative Reverse-transcription Polymerase Chain Reaction (qRT-PCR)
Cytotoxicity Testing
-
The drug susceptibility of two selected clinical strains, Nos. 99 and 108, toward the drugs FLU and ITRA was measured using the broth dilution method and E-test. According to the recommendations of the NCCLS[33], the two strains were cultured with the antifungal agents at 35 ℃ for 24 h. Strain No. 99 showed resistance to FLU (MIC ≥ 64 μg/mL) and ITRA (MIC = 8 μg/mL). Strain No. 108 also showed resistance to FLU (MIC ≥ 64 μg/mL) and ITRA (MIC = 16 μg/mL). To verify these results, we tested these antifungal-resistant strains with E-test strips and confirmed the strong resistance of both strains to FLU and ITRA (Table 2). Because these two strains had a high level of azole resistance to both FLU and ITRA, we chose them as representative strains for further experimental studies.
Drugs aMIC (μg/mL) of Each Strain No. 99 Strain No. 108 Alone* Combination FIC FICI (Outcome) Alone* Combination FIC FICI (Outcome) BDSF ≥ 256 64 ≤ 0.25 ≤ 0.28 ≥ 256 16 ≤ 0.06 ≤ 0.08 FLU ≥ 256 8 ≤ 0.03 (SYN) ≥ 256 4 0.02 (SYN) BDSF ≥ 256 8 ≤ 0.03 ≤ 0.09 ≥ 256 64 ≤ 0.25 ≤ 0.28 ITRA 4 0.25 0.06 (SYN) 8 0.25 0.03 (SYN) Note. Indices were calculated as described in the text. Experiments were carried out thrice. aThe MICs (μg/mL) of BDSF and selected azole drugs were determined using the microdilution checkerboard technique and revealed synergistic activity against azole-resistant C. albicans. *Agreement of E-test MICs with broth dilution MICs. For strain No. 99, the FICIs were ≤ 0.28 for the combination of BDSF and FLU and ≤ 0.09 for the combination BDSF and ITRA. For strain No. 108, the FICIs were ≤ 0.08 for the combination of BDSF and FLU and ≤ 0.28 for the combination BDSF and ITRA. All FICIs ≤ 0.5 show synergy. Table 2. Minimum Inhibitory Concentration (MIC) Indices
-
Table 2 shows the susceptibility of the chosen strains (Nos. 99 and 108) to BDSF in combination with FLU and ITRA using the checkerboard technique. Based on the FICI method, both strains showed synergistic interactions between BDSF and FLU/ITRA. The MICs of FLU, ITRA, and BDSF against strain No. 99 were 256, 4, and ≥ 256 μg/mL, respectively. When BDSF was combined with FLU, the MIC of the former decreased 4-fold to 64 μg/mL, while the MIC of the latter decreased 32-fold to 8 μg/mL. The FICI of this combination was ≤ 0.28. When BDSF was combined with ITRA, the MIC of the former decreased 32-fold to 8 μg/mL, while that of the latter decreased 16-fold to 0.25 μg/mL. The FICI of this combination was ≤ 0.09. As both FICI values obtained are ≤ 0.5, the drug interactions observed are synergistic (Table 2).
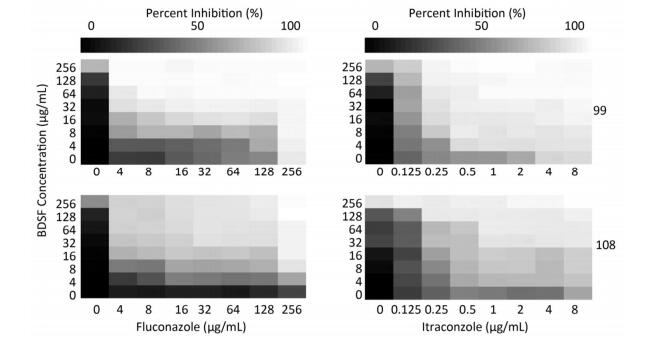
For strain No. 108, when BDSF was combined with FLU, the MIC of the former decreased 16-fold from ≥ 256 μg/mL to 16 μg/mL, while that of the latter dropped 64-fold from ≥ 256 μg/mL to 4 μg/mL. When BDSF was combined with ITRA, the MIC of the former decreased 4-fold from ≥ 256 μg/mL to 64 μg/mL, while that of the latter decreased 32-fold from 8 μg/mL to 0.25 μg/mL. These checkerboard results demonstrate that the MICs of the two tested antifungals alone against C. albicans are significantly higher than their MICs when combined with BDSF. The FICIs of the combination of FLU and ITRA with BDSF were ≤ 0.08 and ≤ 0.28, respectively, indicating synergistic interactions (≤ 0.5) (Table 2). The heat map in Figure 1 shows the synergistic interactions of BDSF in combination with FLU or ITRA. In addition, BDSF in combination with the azoles decreased the amounts of these drugs required to inhibit C. albicans (Table 2).
-
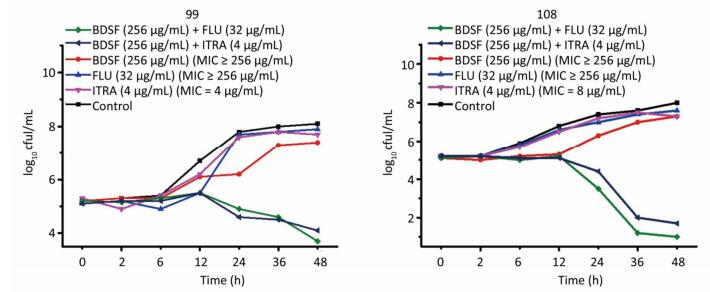
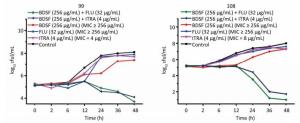
Time-kill studies were performed using BDSF and FLU/ITRA against strain Nos. 99 and 108 (Figure 2). BDSF alone at 256 μg/mL minimally affected yeast cell growth after 48 h; however, growth was significantly reduced in the multi-azole-resistant strains when BDSF in combination with antifungal azoles was added. The combination of FLU and BDSF showed potent fungicidal activity, resulting in a 4.4-log cfu/mL decrease in growth levels of strain No. 99 and a 6.4-log cfu/mL decrease in growth of strain Nos. 108 compared with that induced by 256 μg/mL FLU alone after incubation for 48 h. The combination of ITRA and BDSF yielded 3.4- and 6.9-log cfu/mL decreases in the strain Nos. 99 and 108, respectively. Thus, the proposed BDSF–azole synergy is consistent with the data obtained from the checkerboard technique (Figure 2).
-
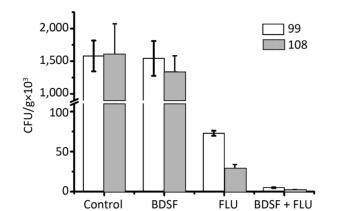
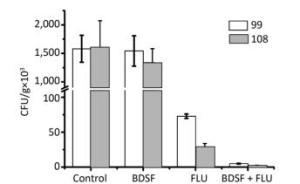
In vitro experimental results confirmed the synergistic effects between BDSF and FLU against multi-azole-resistant C. albicans, further indicating that the combination can be used for clinical application. The effect of this combination on candidiasis in BALB/c mice was estimated by infecting individual mice with strain Nos. 99 and 108, treating them with PBS, FLU, BDSF, or FLU + BDSF, and assessing their kidney fungal burden. Figure 3 shows that the combination of FLU [0.5 mg/(kg·day)] + BDSF [10 mg/(kg·day)] could effectively eliminate the fungal burden in the kidney when compared with FLU [1 mg/(kg·day)] or BDSF [10 mg/(kg·day)] alone. This result indicates that the synergistic effects of BDSF and azole can strongly reduce the dosage of azole required to treat candidiasis in mice.
-
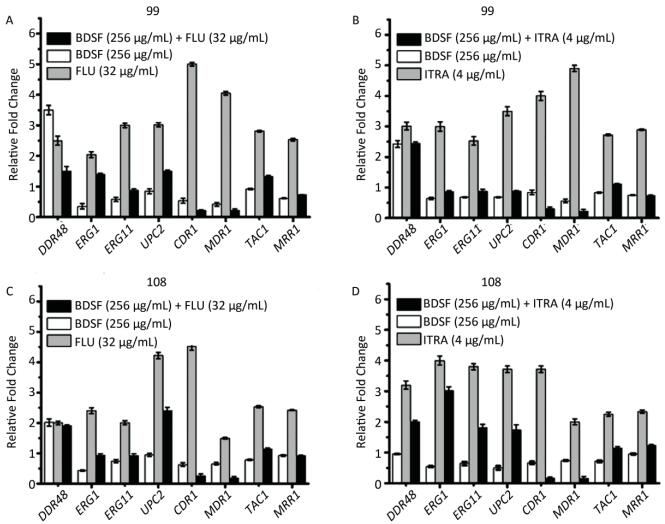
To investigate the synergistic mechanism of FLU/ITRA and BDSF against multi-azole-resistant C. albicans, qRT-PCR was conducted (Figure 4). The mRNA expression of ERG1, ERG11, UPC2, CDR1, MDR1, TAC1, and MRR1 in BDSF-only treated strains consistently showed a decrease in expression compared with that of the standard ACT1; by comparison, the mRNA expression of DDR48 was upregulated or invariable when compared with that of the standard ACT1. The mRNA levels of DDR48, ERG1, ERG11, UPC2, CDR1, MDR1, TAC1, and MRR1 in multi-azole-resistant strains were upregulated by 1.5–5.1-fold in strains treated with FLU (Figure 4A, 4C) or ITRA (Figure 4B, 4D) alone. In strain No. 99, the combination of BDSF and FLU inhibited the expression of CDR1 and MDR1 by 10-fold, respectively, when compared with FLU alone (Figure 4A), while the combination of BDSF with ITRA inhibited the expression of CDR1 and MDR1 by 8- and 11-fold, respectively, when compared with ITRA alone (Figure 4B). In strain No. 108, the combination of BDSF with FLU inhibited the expression of CDR1 and MDR1 by 7.5- and 5-fold, respectively, compared with FLU alone (Figure 4C). The combination of BDSF with ITRA inhibited the expression of CDR1 and MDR1 by 6.3- and 4-fold, respectively, compared with ITRA alone (Figure 4D).
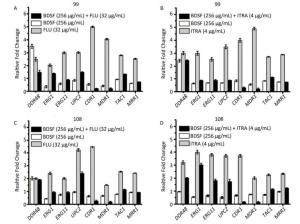
Figure 4. mRNA expression levels of DDR48, ERG1, ERG11, UPC2, CDR1, MDR1, TAC1, and MRR1 determined by qRT-PCR experiments after C. albicans (strain Nos. 99 and 108) were treated with BDSF alone, FLU or ITRA alone, or BDSF + azole.
The expressions of CDR1 and MDR1 in the combination group decreased obviously when compared with those in the BDSF-alone group. By contrast, the mRNA expression of DDR48, ERG1, ERG11, and UPC2 in the combination group, except for DDR48 in the group in which strain No. 99 was with FLU + BDSF, was upregulated or invariable when compared with that in the BDSF-alone group (Figure 4). These results indicate that the combination of BDSF with azole does not affect the ergosterol synthesis pathway and the expression of DDR48. Previous studies indicate that the expressions of CDR1 and MDR1 are modulated by the transcription factors TAC1 and MRR1, respectively[8]. As shown in Figure 4, the expressions of TAC1 and MRR1 were 1.9-2.8 fold upregulated, as expected, in azole-resistant strains under the condition of drug induction. Interestingly, addition of BDSF (256 μg/mL) to samples with FLU (32 μg/mL) or ITRA (4 μg/mL) downregulated the mRNA expression levels of the TAC1 and MRR1 genes. Based on the above results, the combination of BDSF and FLU or ITRA strongly represses CDR1 and MDR1 gene expression via suppression of the expression levels of TAC1 and MRR1, respectively, in the multi-azole-resistant strains when compared with FLU/ITRA treatment alone.
-
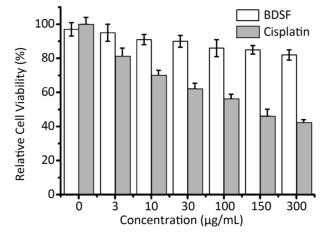
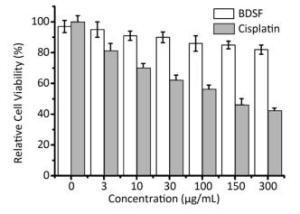
BDSF could potentially be used against infection for its ability to inhibit some virulence factors in C. albicans[31, 36]. The cytotoxicity of BDSF was estimated against a normal human fibroblast (foreskin) cell line by MTT assay. The cell line showed no significant cytotoxicity at BDSF testing levels of up to 300 μg/mL (Figure 5). By comparison, cisplatin was more toxic to fibroblast cells, resulting in only 40% viability at 300 μg/mL (Figure 5). This result indicates that BDSF may be safe for human use at low doses.
Drug Susceptibility of Multi-azole-resistant Strains by Broth Dilution and E-test
Synergistic Interaction of BDSF in Combination with FLU or ITRA Using Checkerboard Test
BDSF in Combination with FLU or ITRA Inhibited Growth in Azole-resistant Strains
Synergy between BDSF and FLU in Vivo
Combination of BDSF with FLU/ITRA Inhibited the Expression of CDR1 and MDR1 Genes
BDSF Exhibits No Significant Cytotoxicity
-
A number of small signaling molecules from bacteria regulate the morphological transition of C. albicans, which corresponds to the upward trend in antifungal drug development. Among the current stratagems available, synergy is an effective method for combination therapy since it decreases drug dosages and reduces the emergence of drug resistance[37]. Herein, we chose a signaling molecule, BDSF, from B. cenocepacia, to individually combine with antibiotics (FLU or ITRA) and evaluate the resulting in vitro effects against two multi-azole-resistant C. albicans strains. No antagonism (FICI ≥ 4) was found among these combinations, and treatment of the multi-azole-resistant strains with both BDSF + FLU and BDSF + ITRA revealed apparent synergistic results (FICI values ≤ 0.5). In addition, combination treatment exhibited stronger antifungal performance compared with treatment by the azoles or BDSF alone, thus indicating that maintaining the same efficacy via combination therapy can decrease the antibiotic dosage and side effects, as well as the development of drug resistance[17]. BDSF mediates its synergistic effect with azoles by inhibiting efflux pump expression in FLU-resistant C. albicans isolates rather than by interfering with ergosterol biosynthesis, one of the usual mechanisms of azole drugs. Previous studies show that many resistant strains of C. albicans overexpress CDR1 and MDR1, two efflux pump genes affecting the transportation of antifungal drugs[12]; these effects have led to azole resistance in Candida[38]. Development of resistance to azoles is a complex process, and increased efflux pump activity to expel azole molecules from the cell is likely the main mechanism involved[12]. The use of BDSF in combination with azoles, as shown in this study, is able to overcome this mechanism of drug resistance in C. albicans.
The emergence of drug-resistant strains is likely the result of the abuse of antifungal drugs in the clinical setting. Resistance to azoles widely used in antifungal therapy can occur in C. albicans by overexpression of some genes. The expression of these genes is controlled by transcription factors, such as TAC1, which is involved in the control of CDR1, MRR1, which is involved in the control of MDR1, and UPC2, which is involved in the control of ERG11, which, in turn, encodes azole target genes[8]. Our qRT-PCR results showed decreased mRNA levels of the efflux pump gene family members CDR1 and MDR1, the transcriptional activator of the CDR gene TAC1, and the multidrug resistance regulator gene MRR1; these changes are correlated with decreased drug resistance. Since TAC1 is responsible for CDR1 upregulation, MRR1 is required for the constitutive activation of the MDR1 promoter in azole-resistant strains[39, 40]. We propose that the combination of BDSF with azoles mainly reduces CDR1 and MDR1 efflux pump activity via downregulating the mRNA expression levels of TAC1 and MRR1, respectively, rather than UPC2-mediated ergosterol levels, which are the usual target of azole drugs in fungi. Overexpression of such transporters increases the ability of fungal cells to expel harmful substances, such as FLU or ITRA; however, BDSF may efficiently downregulate the expression of CDR1 and MDR1 by altering the gene expression patterns of regulators governing the expression of efflux pumps in the presence of the antifungals, resulting in decrease intrinsic azole resistance. Previous studies show that antifungal-resistant strains of C. albicans overexpress CDR1 and MDR1[12]. The synergy observed in the combination of BDSF and FLU or ITRA provides an available approach to conquer antifungal drug resistance. Further in vivo experiment results indicate that this combination therapy provides an effective therapeutic approach to treat candidiasis.
Farnesol, a structural analogue of BDSF, comes from C. albicans and is synergistic to azoles and amphotericin by mediating the ABC superfamily of transporters[41]. Interestingly, the mRNA expression levels of ERG9, ERG11, and ERG20 are inhibited by 200 μmol/L farnesol, while those of ERG9, ERG11, and ERG20 are restored, when C. albicans is treated with FLU and farnesol[41]. In this study, we found that BDSF alone downregulates ERG9 and ERG11 and upregulates the same when combined with azoles. This phenomenon may be caused by the dominant effects of the azoles over BDSF. We discovered that BDSF also acts synergistically with azoles at low concentrations by interfering with efflux pumps. Efflux pump factors affect azole resistance, which is a key solution to treat fungal infections[25, 38]. Inhibition of drug efflux pumps is considered a feasible strategy to overcome clinical antifungal resistance[25, 42]. BDSF is a cis-monounsaturated fatty acid, and daily intake of unsaturated fatty acids protects against cardiovascular disease and enhances immune function[43]. In vitro cytotoxicity analysis indicated that BDSF has good biocompatibility, making it an ideal drug for combination antifungal therapy.
This study is the first of its kind to discover that BDSF may be a new antibiotic-synergistic prototype against fungal pathogens. The inhibitory effect of BDSF toward CDR1 and MDR1 efflux pump activity is a very important and significant finding for further development of new BDSF-azole combination strategies to treat azole-resistant Candida infections.
-
The authors declare no conflict of interest.
-
We thank Professor LIAO Wan Qing in Changhai Hospital (Shanghai, China) for providing C. albicans clinical strains.







 Quick Links
Quick Links
 DownLoad:
DownLoad: