-
Acute pancreatitis, one of the most common gastroenterological diseases, is commonly caused by self-digestion of trypsin. Acute pancreatitis usually has a mild and self-limiting clinical outcome and relapses occur more frequently in younger individuals[1]. Previous studies have reported that approximately 20% to 30% of acute pancreatitis patients will develop systemic inflammatory response syndrome and intestinal bacteria translocation, which can increase acute pancreatitis severity[2-3]. Acute pancreatitis can also develop into chronic pancreatitis, especially in alcoholics[1].
The gastrointestinal tract is colonized by a vast community of symbionts; there are at least 1, 000 different microbial species and approximately 1, 014 microorganisms in the human intestinal wall. The intestinal microbiota plays an important role in the development of the intestinal architecture and function, the gut-associated immune system, and epithelial cell functions. Dysbiosis of the intestinal microbiota has been reported to be associated with intestinal barrier dysfunction and the high mortality rates observed in acute pancreatitis patients with systemic inflammatory response syndrome and intestinal bacterial translocation[4]. Previous studies have demonstrated reduced richness and diversity of the predominant fecal microbiota in most acute pancreatitis cases, even when symptoms are mild, indicating that dysbiosis of the intestinal microbiota occurred prior to systemic inflammatory response syndrome[5]. Microbial translocation occurs in acute pancreatitis and is in accordance with disease severity[6-7]; thus, it is important to maintain intestinal barrier function by restoring intestinal microbiota during the treatment of acute pancreatitis. Although there are a few studies on changes in intestinal microbial community in patients with pancreatitis[6], the effects of probiotics are controversial. Therefore, additional studies regarding microbiota community changes in acute pancreatitis patients are urgently needed to provide more information for clinical treatment.
In this study, we compared the differences in intestinal microbial communities between acute pancreatitis patients and healthy volunteers; intestinal microbial community was determined by high-throughput 16S rRNA gene amplicon sequencing.
A total of 89 participants were enrolled in this study including 45 acute pancreatitis patients (group A) and 44 healthy volunteers (group B) who underwent physical examination at the same hospital. The patients were diagnosed as acute pancreatitis at the department of Gastroenterology, the First Hospital of Nanjing. Patients with two of the following factors were diagnosed with acute pancreatitis: 1) Acute pancreatitis characteristic abdominal pain; 2) Serum amylase and/or lipase ≥ 3-times higher than the upper limit of normal; 3) Computed tomography showing acute pancreatitis. The exclusion criteria for the two groups were as follows: 1) Individuals with digestive system diseases other than acute pancreatitis; 2) individuals recently treated with antibiotics; 3) Individuals with other serious diseases that could not cooperate with sample collection. Pancreatitis patients suffering from digestive system diseases were excluded. The study was conducted according to the guidelines of Helsinki Declaration. Ethical approval for this study was obtained from the Ethics Committee of First Hospital of Nanjing. Written informed consent was obtained from all the participants.
Fecal samples were collected from acute pancreatitis patients prior to treatment. Following coloclyster with physiological saline, fresh fecal samples (approximately 1 g) were collected using steam-sterilized tubes, transported immediately from our hospital to the laboratory on an ice pack, and stored in a -80 ℃ freezer. The healthy volunteer samples were collected on the same day as the acute pancreatitis patient samples.
Each fecal sample (200 mg) was processed for DNA extraction using the QIAamp Media MDx Kit (Qiagen, German) according to the manufacturer's recommendations. DNA concentration was measured using a spectrophotometer DNA library construction, including workflows as cluster generation, template hybridization, isothermal amplification, linearization, blocking, denaturing, and hybridization of the sequencing primers, was performed according to the manufacturer's instructions (Illumina). In addition, paired-end sequencing was performed with 23, 100 base pairs (bp) for all libraries. The same insert size inferred by Agilent 2, 100 was used for all libraries (ranging from 275 to 450).
High-throughput 16S rRNA gene amplicon sequencing[8] was used to analyze microbial composition in fecal samples collected from the 45 acute pancreatitis patients and 44 healthy volunteers. Subsequent to bacterial DNA extraction and detection, the V3 hypervariable 16S rRNA region was amplified using universal PCR primers 338F (5'-CCTACGGGAGGCAGCAG-3') and 518R (5'-ATTACCGCGGCTGCTGG-3'). Following sequencing library construction, 16S rDNA PCR products with an amplicon size of 180 bp were sequenced using double-terminal sequencing on an Illumina MiSeq sequencing platform.
Because sequencing errors, such as point mutations, usually occur in high-throughput sequencing and the quality of the end of the sequence is relatively low, we first optimized the raw sequencing data in order to obtain higher quality and more accurate bioinformatic analysis results. Step one, all raw sequencing files were merged according to the sequencing file name; step two, the merged sequencing file was matched according to the R1/R2 file of the corresponding samples and the two sequences were aligned according to the matching overlapping area. An overlap of at least 10 bp was guaranteed at the time of splicing and the sequence containing N in the spliced result was then removed; step three, the samples were split according to the barcode information; step four, the primer and linker sequence, bases with a mass value of < 20 at both ends, and sequences < 120 bp in length were all removed; step five, sequences from the above spliced filter were compared with the database to remove chimera sequences and the final valid data were obtained.
Operational taxonomic unit (OTU) analysis, species annotation, α diversity index analysis, and UniFrac were analyzed using the QIIME software (1.9.1). Statistical analysis was carried out using commercially available software (SAS software, version 9.4; SAS Institute Inc., Cary, NC, USA). Normally distributed measurement data were described by means ± standard deviations and the means were compared using the Student's t-test. Skewed measurements were described by median (Q25, Q75) and the differences between the two groups were tested using a nonparametric test. Categorical variables were described as percentages and their differences were compared using the chi-squared test. Principal co-ordinates analysis (PcoA) of microbial communities in fecal samples from acute pancreatitis patients and healthy volunteers was performed with the QIIME software.
A total of 89 participants were eligible for inclusion in this study; 45 acute pancreatitis patients and 44 healthy volunteers. Fifty-one of the participants were male and there was no significant difference between acute pancreatitis patients and healthy volunteers in terms of gender (P = 0.232). The age of the acute pancreatitis patients and the healthy volunteers was 53.52 ± 13.64 years and 53.59 ± 9.18 years, respectively (P = 0.960). The acute pancreatitis patients exhibited higher white blood cell counts (P < 0.001), neutrophil ratio (P < 0.001), glucose (P < 0.001), aspartate transaminase (P < 0.001), alanine aminotransferase (P = 0.001), total bilirubin (P < 0.001), and CA199 (P < 0.001). In contrast, the hemoglobin (P < 0.001), packed cell volume (P = 0.010), and total cholesterol (P = 0.001) of acute pancreatitis patients were lower (Table 1).
Variable Total (n = 89) Acute Pancreatitis Patients (n = 45) Healthy Volunteers (n = 44) T/Χ2 P Gender, n (%) Male 51 (57.30) 23 (45.10) 28 (54.90) 1.427 0.232 Female 38 (42.70) 22 (57.89) 16 (42.11) Age (years) 53.52 ± 13.64 53.44 ± 17.02 53.59 ± 9.18 21.063 0.960 Total number of white blood cells (× 109) 7.54 (5.65, 10.89) 10.74 (7.86, 12.85) 6.37 (4.93, 7.39) 21.079 < 0.001 Neutrophil ratio (%) 65.00 (52.60, 83.40) 83.40 (76.10, 90.50) 52.50 (50.29, 57.60) 39.044 < 0.001 Hemoglobin (g/L) 141.75 ± 18.76 136.36 ± 21.26 147.27 ± 14.01 71.277 < 0.001 Packed cell volume (%) 41.30 (38.90, 44.50) 40.10 (38.70, 43.00) 42.70 (40.19, 44.90) 80.205 0.010 Platelet count (× 109) 189 (163, 230) 203 (155, 239) 187 (165, 225) 9.815 0.614 Glucose (mmol/L) 5.81 (5.16, 7.03) 7.00 (5.90, 8.70) 5.23 (4.98, 5.82) 21.707 < 0.001 Blood urea nitrogen (mmol/L) 4.80 (3.90, 6.00) 4.70 (3.90, 5.49) 5.10 (4.01, 6.12) 20.012 0.312 Aspartate transaminase (U/L) 26.0 (20.0, 40.0) 38.0 (20.0, 142.0) 24.0 (20.0, 28.0) 6.512 < 0.001 Alanine aminotransferase (U/L) 33.0 (20.0, 64.0) 44.5 (20.5, 234.0) 28.0 (20.0, 34.0) 6.213 0.001 Total bilirubin (µmol/L) 13.60 (11.11, 20.60) 20.30 (12.85, 27.10) 11.90 (10.50, 14.60) 7.335 < 0.001 Total cholesterol (mmol/L) 4.54 (4.00, 5.22) 4.15 (3.62, 4.87) 4.76 (4.42, 5.28) 37.202 0.001 Triacylglycerol (mmol/L) 1.66 (0.96, 2.82) 1.09 (0.65, 3.14) 2.02 (1.53, 2.54) 4.402 0.027 CA19-9 (U/mL) 11.38 (8.29, 21.12) 21.94 (12.77, 38.16) 9.07 (7.92, 11.38) 4.299 < 0.001 Note.Normally distributed measurement data were described by means ± standard deviations. Skewed measurements were described by median (Q25, Q75). Table 1. Clinical Characteristics and Laboratory Results of All Participants
The comparison between the alpha diversity indices of pancreatitis patients and healthy volunteers is shown in Table 2. After filtering out low-quality, chimeric, and non-microbial sequences, there was no significant difference between the two groups in total tags after the quality control (P = 0.083). The average number of acute pancreatitis patient tags used to construct the OTUs was significantly higher than that of healthy volunteers (P = 0.044); however, the difference in the number of OTUs was not significant between the two groups (P = 0.599). The healthy volunteers had a higher Chao1 index (P < 0.001) and Shannon index (P = 0.003) than the acute pancreatitis patients.
Variable Acute Pancreatitis Patients Healthy Volunteers P Total Tags 14271.00 (6293.00, 39358.00) 8373.00 (4350.75, 26885.00) 0.083 Tags (Genus) 13191.00 (5234.50, 35701.00) 6408.50 (3417.25, 23175.00) 0.044 OTUs* 1497.00 (806.00, 2992.00) 1350.50 (798.00, 2344.50) 0.599 Chao1 889.05 (801.85, 985.31) 1156.12 (968.66, 1430.22) < 0.001 Shannon 6.32 ± 0.54 6.62 ± 0.38 0.003 Note.*Operational taxonomic units. Normally distributed measurement data were described as means ± standard deviations and abnormally distributed data were designated as median (Q25, Q75). Table 2. Alpha Diversity Index of Pancreatitis Patients and Healthy Volunteers
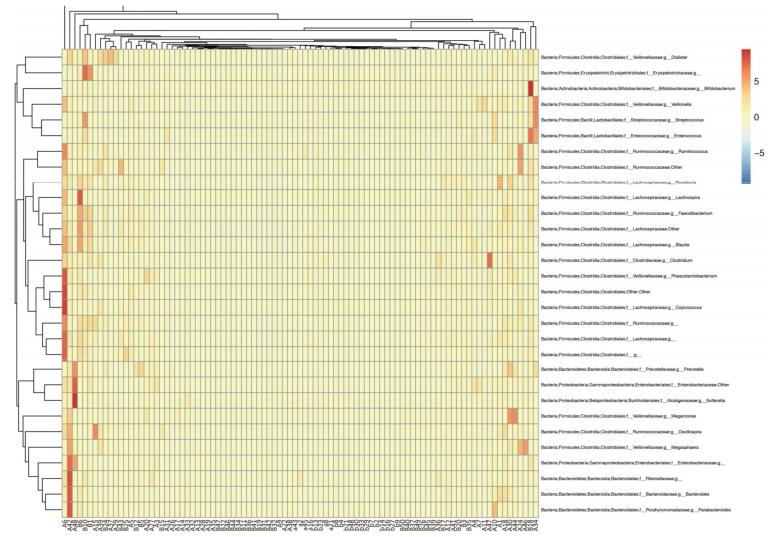
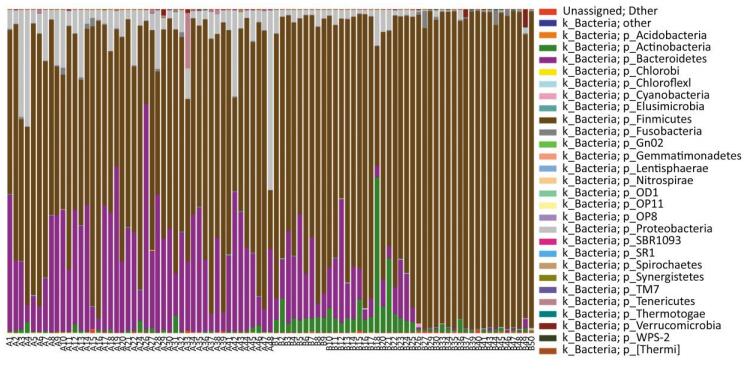
A total of 27 microbial phyla were detected and the samples of pancreatitis patients contained fewer phyla. Moreover, acute pancreatitis samples contained more Bacteroidetes and Proteobacteria and fewer Firmicutes and Actinobacteria compared with the samples of healthy volunteers (Supplementary 1 available at www.besjournal.com). Table 3 details the main phyla present in the intestinal microbial communities of the two groups. Species and sample clustering were performed based on the phyla of each sample and a clustered heat map was constructed (Figure 1). Different colors represent the relative abundance of each species in the sample.
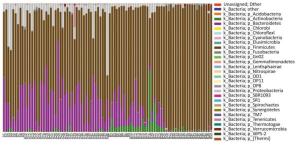
Figure Supplementary 1. Distribution of bacteria at phylum level. Microbes in the samples of pancreatitis patients contained fewer phyla and acute pancreatitis samples contained more Bacteroidetes and Proteobacteria compared with healthy volunteers, while less Firmicutes were obtained. A group means pancreatitis patients and B group means healthy volunteers.
Acute Pancreatitis Patients Healthy Volunteers Phylum Percent (%) Phylum Percent (%) Firmicutes 61.84 Firmicutes 84.67 Bacteroidetes 26.35 Bacteroidetes 7.66 Proteobacteria 9.77 Actinobacteria 4.52 Actinobacteria 0.77 Proteobacteria 2.07 Tenericutes 0.58 Fusobacteria 0.47 Table 3. The Main Phyla Present in the Intestinal Microbial Communities of Healthy Volunteers and Acute Pancreatitis Patients
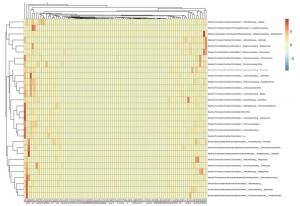
Figure 1. Clustered heat map of acute pancreatitis patients and healthy volunteers at the genus level. Group A, acute pancreatitis patients; group B, healthy volunteers. Different colors indicate the relative abundance of each species in the sample.
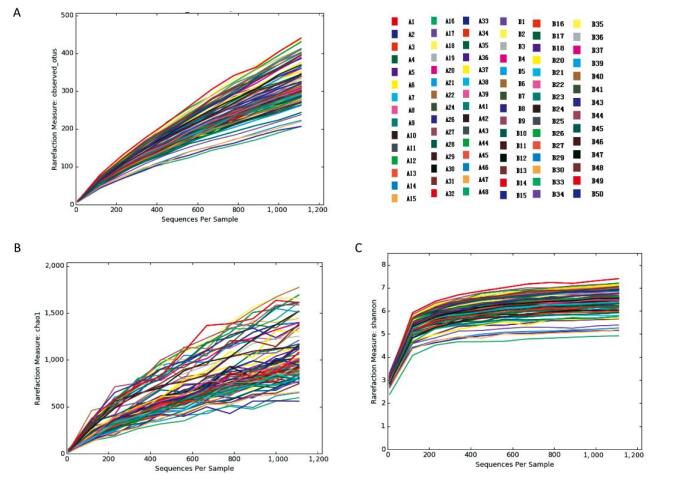
The trends of the rarefaction curves and Chao1 rarefaction curves were gradually gentle and there was an inflection point at approximately 150 in the Shannon rarefaction curve. These results suggest that the sequencing depth and the richness of the species in the samples were sufficient for subsequent analysis. (Supplementary 2 available at www.besjournal.com)
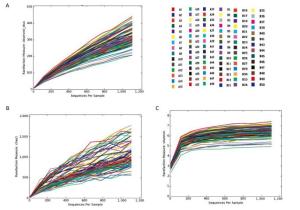
Figure Supplementary 2. Rare faction curves, Chao1 rare faction curves and Shannon rarefaction curves of observed OTUs. (A) Rare faction curves of observed OTUs. (B) Chao1 rarefaction curves of observed OTUs. (C) Shannon rarefaction curves of observed OTUs. A group means pancreatitis patients and B group means healthy volunteers.
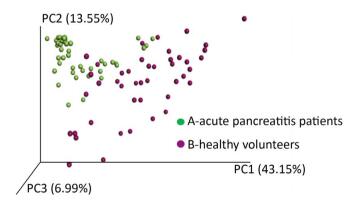
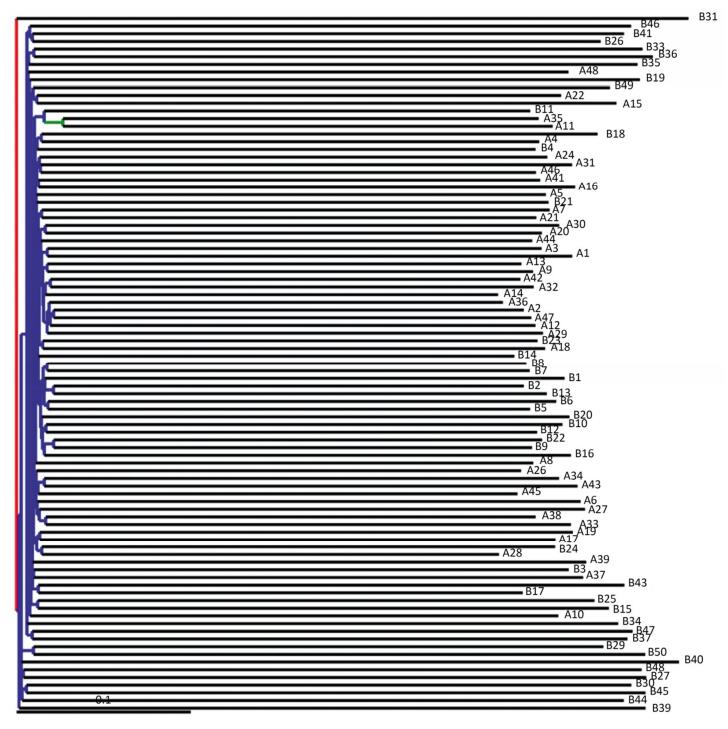
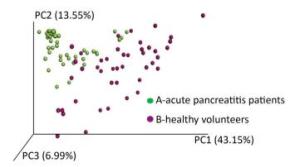
A phylogenetic tree was constructed using representative OTU sequences from different environmental samples. The UniFrac metric measures the difference between two different environmental samples based on the length of the constructed phylogenetic tree. The weighted UniFrac PCoA analysis was shown in Figure 2. The proportion of PC1, PC2, and PC3 was 43.15%, 13.55%, and 6.99%, respectively. The PCoA analyses could effectively distinguish the fecal microbial communities in acute pancreatitis patients from those in healthy volunteers. Next, we used the unweighted pair group method with arithmetic mean (UPGMA) clustering method; sample clustering was based on the unweighted UniFrac distance matrices. UPGMA could also distinguish between the microbial communities of acute pancreatitis patients and healthy volunteers (Figure 3).
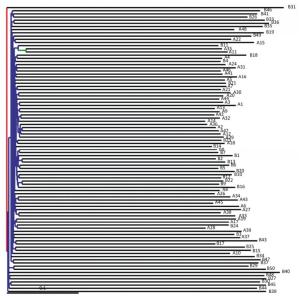
Figure 3. Unweighted UniFrac UPGMA tree. Each branch in the graph represents a sample and the inner nodes separate the bootstrap values of different intervals by color. Red denotes bootstrap values between 75%-100%; yellow indicates bootstrap values between 50%-75%; green signifies bootstrap values between 25%-50%; and blue represents bootstrap values < 25%.
Metagenomics is a new concept and method for studying microbial diversity and developing novel physiologically active substances based on microbial genomics. In this study, we analyzed the microbial community composition and diversity of fecal samples from acute pancreatitis patients and healthy volunteers using the Illumina MiSeq high-throughput sequencing platform. The fecal microbial communities of acute pancreatitis patients and healthy volunteers were distinct and the intestinal microbial community composition is altered along the course of acute pancreatitis.
A previous study reported that urinary trypsinogen-2 and serum amylase have similar value in diagnosing acute pancreatitis[9]. Aufenanger et al. showed that pancreatic phospholipase A2 activity in serum is strongly elevated in severe acute pancreatitis, thus enabling discrimination between mild and severe forms of this disease in its early phase[10]. Resistin has also been reported as a new, inexpensive, and effective indicator for predicting acute pancreatitis severity and the presence of necrosis[11]. Our finding that acute pancreatitis samples contained more Bacteroidetes and Proteobacteria and fewer Firmicutes and Actinobacteria compared with healthy volunteers suggests that some intestinal microbiota may lead to bacterial translocation and result in secondary infection, which exacerbates acute pancreatitis.
The intestinal microbiota is very important in the prevention of bacterial overgrowth and secondary infectious complications. Changes in the intestinal microbial community lead to alteration of the intestinal barrier function, resulting in bacterial overgrowth and impaired immunity[12]. Lutgendorff et al. reported that probiotic pre-treatment beginning five days prior to the induction of acute pancreatitis diminished acute pancreatitis-induced intestinal barrier dysfunction and prevented oxidative stress via mechanisms mainly involving mucosal glutathione biosynthesis in rats[13]. Although further studies are required, our findings suggest that modulation of intestinal microbes may constitute an alternative strategy for acute pancreatitis therapy.
Our study has several limitations. First, according to previous studies[6], the microbiota in colonic mucosal sites is quite different than fecal microbial communities; however the mucosal microbiota was not included in this study. Second, there was no obvious inflection point in the rarefaction curves and Chao1 rarefaction curves of all samples, indicating that the richness of the samples was not sufficient although the trends were gradually gentle.
In conclusion, the intestinal microbes of acute pancreatitis patients are different from those of healthy volunteers. The occurrence of acute pancreatitis can cause intestinal microbial disorders without any treatment; any treatments that could normalize the intestinal microbes of acute pancreatitis patients should be promoted.
The authors declare that no potential conflicts of interest exist.
HTML
Nanjing Medical Technology and Development General Subject YKK14092







 Quick Links
Quick Links
 DownLoad:
DownLoad: