-
Coronavirus Disease 2019 (COVID-19) is a pulmonary inflammatory disease caused by infection with the highly pathogenic new coronavirus (SARS-CoV-2)[1]. To date, COVID-19 has spread to at least 210 countries and territories, infected more than 10.94 million people, and killed more than 519,000 people, according to Johns Hopkins University. On April 15, 2020, the COVID-19 outbreak was declared a global health emergency[2]. SARS-CoV-2 is the third novel coronavirus able to infect humans that has crossed to humans from another species in the past two decades. The other two were severe acute respiratory syndrome-related coronavirus (SARSr-CoV) in 2002–2003, and middle east respiratory syndrome coronavirus (MERS-CoV) in 2014. All of the coronaviruses mentioned above belong to the beta coronavirus subgroup, which has obvious person-to-person transmission characteristics[3].
Recent studies have found that SARS-CoV-2 can successfully infect host cells through the SARS-CoV receptor angiotensin converting enzyme 2 (ACE2)[4]. Using X-ray diffraction technology, researchers have successfully solved the crystal structure of the complex between the receptor molecule ACE2 and the surface spike glycoprotein (S protein) of SARS-CoV-2, and found the interaction sites between SARS-CoV-2 receptor binding domain (RBD) and ACE2[5]. Several research teams have analyzed this crystal structure of the surface spike glycoprotein of SARS-CoV-2 in complex with ACE2. As shown in Figure 1A, residues Q24, E35, E37, D38, Y41, Q42, Y83, K353, and R393 in ACE2 form hydrogen bonds with residues N487, Q493, Y505, Y449, T500, N501, Q498, G446, N489, Y495, G496, G502, and Y505 in SARS-CoV-2, respectively. Residues D30 in ACE2 and K417 in SARS-CoV-2 interact via a disulfide bond. We collected the ACE2 amino acid sequences of 18 different species in the NCBI database to perform comparative analyses of the secondary structures of ACE2 proteins from different species using the above observations from the existing structure. We then constructed a phylogenetic tree by comparing these sequences. The accession numbers for ACE2 sequences included in the manuscript are as follows: Homo sapiens ACE2, BAD99266.1; Mus musculus ACE2, AAH26801.1; Sus scrofa ACE2, NP_001116542.1; Macaca mulatta ACE2, NP_001129168.1; Bos taurus ACE2, NP_001019673.2; Gallus gallus ACE2, QEQ50331.1; Canis lupus familiaris ACE2, ACT66277.1; Ovis aries ACE2, XP_011961657.1; Anas platyrhynchos ACE2, XP_012949915.2; Equus caballus ACE2, XP_001490241.1; Rattus norvegicus ACE2, NP_001012006.1; Rhinolophus sinicus ACE2, ADN93475.1; Manis javanica ACE2, XP_017505746.1; Felis catus ACE2, NP_001034545.1; Xenopus tropicalis ACE2, XP_002938293.2; Danio rerio ACE2, NP_001007298.1; Oncorhynchus mykiss ACE2, XP_021433278.1; Oryctolagus cuniculus ACE2, QHX39726.1. The conservation at each of these 10 loci in ACE2 is shown in Figure 1B and 1C. The residue D30 in ACE2, which can form a disulfide bond with SARS-CoV-2, displays the most variability among species of the 10 ACE2 loci examined. ACE2 in fish, amphibians, and poultry differs from human ACE2 at more than 40% of examined interaction sites. Monkeys, pigs, cattle, sheep, and rabbits display identical residues at more than 80% of the examined interaction sites. Overall sequence similarities among ACE2 amino acid sequences of the chosen species were also analyzed. The ACE2 amino acid sequences of all mammals and reptiles share 80.6%–94.9% identity with their human counterpart. Poultry and aquatic organisms share only 55.5%–65.2% and 55.3%–56.9% identity with their human counterpart, respectively (Supplementary Table S1 available in www.besjournal.com).
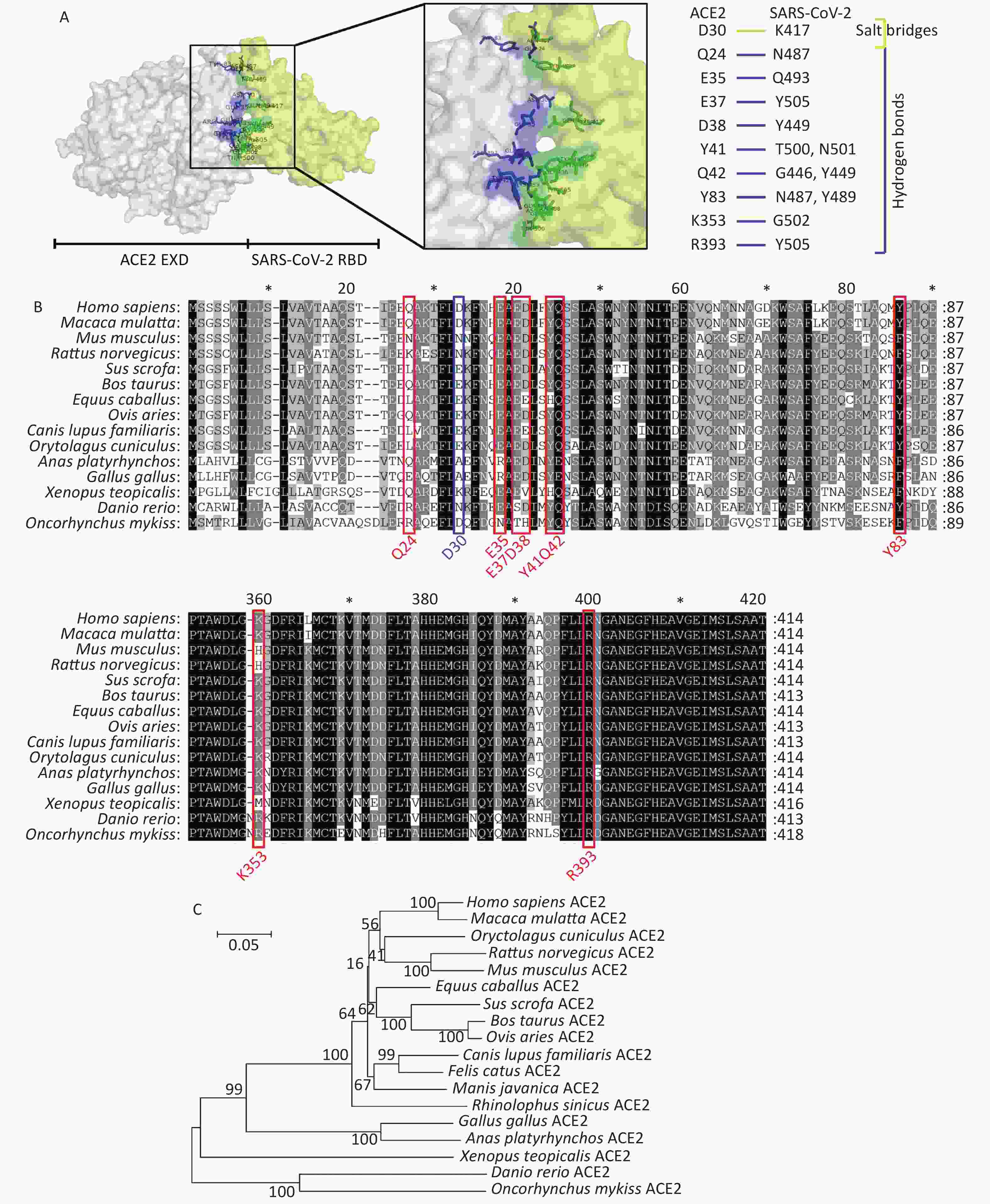
Figure 1. Comparative analyses of secondary structure in COVID-19 receptor molecule ACE2 proteins of different species. (A) The crystal structure of the human ACE2 SARS-CoV-2 S protein complex, illustrating energetically favorable interactions between ACE2 and the SARS-CoV-2 S protein. (B) Alignment of ACE2 amino acid sequences from human and 14 other species. Residues outlined in red and blue form hydrogen bonds and a disulfide bond, respectively, at the SARS-CoV-2/ACE2 interface. (C) Phylogenetic analysis of ACE2 from 18 different species. The unrooted phylogenetic tree was constructed via the neighbor-joining method based on the alignment of multiple amino acid sequences (CLUSTAL W).
Species B C D E F G H I J K L M N O P Q R A 94.9 82.1 82.4 81.3 80.8 86.8 81.6 80.6 85.2 83.3 84.8 84.8 64.2 65.2 55.5 55.3 56.9 B 81.9 82.1 80.8 81.3 86.5 81.9 80.3 84.4 83.1 85.0 84.9 64.2 64.6 54.9 54.6 56.6 C 90.4 80.6 80.2 82.8 80.0 78.0 81.7 81.1 83.3 82.7 65.2 66.0 57.4 55.6 57.4 D 80.0 80.1 82.1 80.0 77.6 81.6 80.6 83.6 82.6 65.1 65.0 56.0 55.8 57.1 E 87.3 87.0 87.4 80.1 83.9 81.9 82.4 82.6 65.1 65.3 54.5 54.8 57.2 F 85.0 97.1 79.7 83.1 81.9 82.6 81.6 63.5 65.0 55.4 54.3 55.8 G 85.3 84.6 88.0 87.0 85.7 88.0 65.5 66.5 55.3 55.7 57.6 H 80.5 83.3 81.7 82.4 82.2 64.1 65.6 55.3 54.7 56.0 I 83.8 81.2 79.2 82.7 64.4 65.5 55.4 55.0 56.9 J 90.5 84.1 87.3 65.5 67.0 56.2 55.6 56.7 K 82.9 86.2 65.0 65.5 55.7 54.8 56.6 L 84.1 65.0 64.5 56.1 55.7 57.1 M 65.1 65.3 56.6 55.9 57.9 N 86.0 56.6 54.37 56.4 O 56.9 53.8 56.4 P 52.2 54.0 Q 66.4 Note. A, Homo sapiens (human) ACE2; B, Macaca mulatta (rhesus monkey) ACE2; C, Mus musculus (mouse) ACE2; D, Rattus norvegicus (rat) ACE2; E, Sus scrofa (pig) ACE2; F, Bos Taurus (cattle) ACE2; G, Equus caballus (horse) ACE2; H, Ovis aries (sheep) ACE2; I, Rhinolophus sinicus (bat) ACE2; J, Felis catus (cat) ACE2; K, Canis lupus familiaris (dog) ACE2; L, Oryctolagus cuniculus (rabbit) ACE2; M, Manis javanica (pangolin) ACE2; N, Anas platyrhynchos (duck) ACE2; O, Gallus gallus (chook) ACE2; P, Xenopus tropicalis (amphibian) ACE2; Q, Danio rerio (zerafish) ACE2; R, Oncorhynchus mykiss (trout) ACE2. Table S1. Percentages (%) of amino acid sequence identity for the ACE2 amino acid sequences in different species
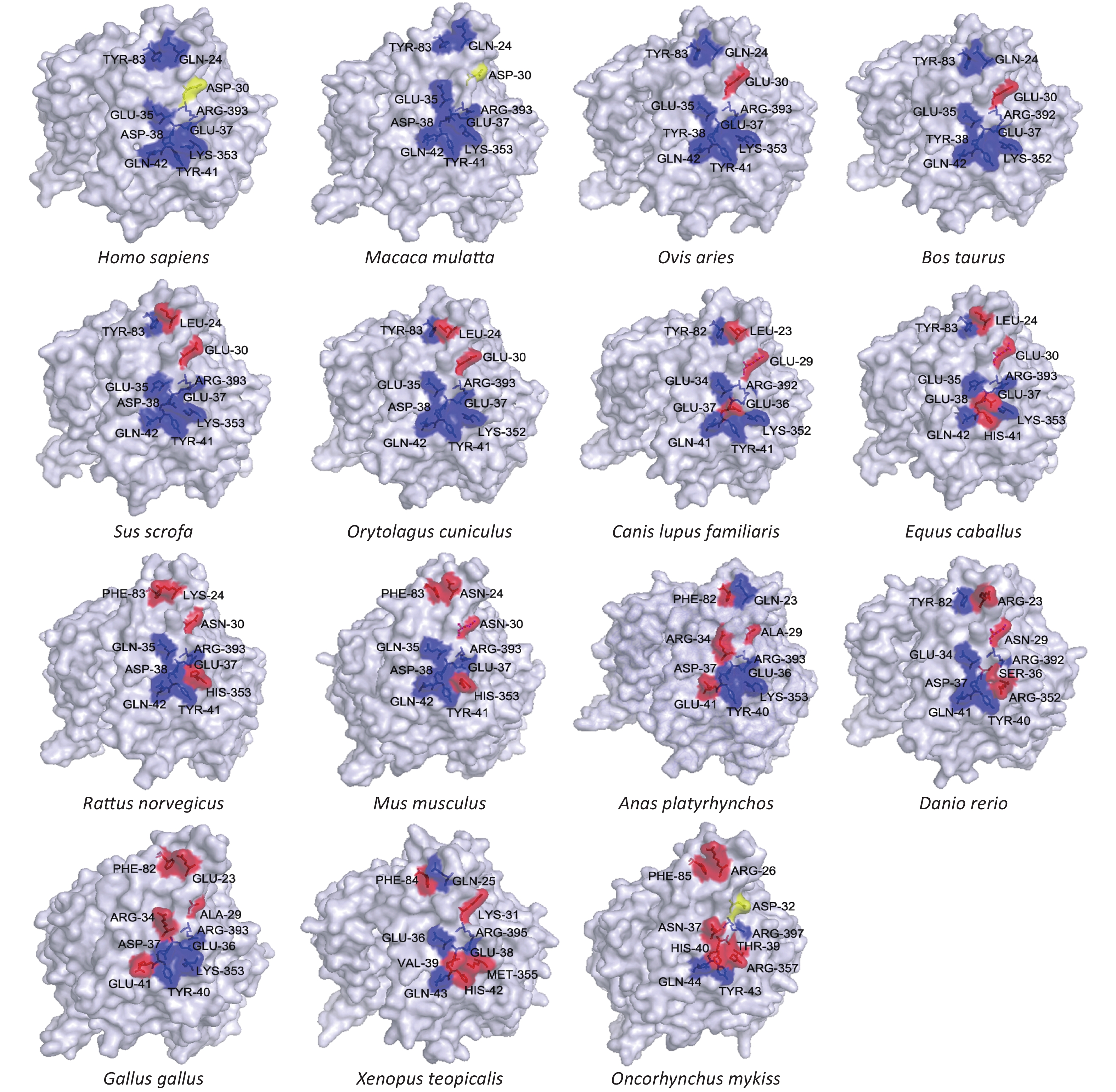
We also modeled the structure of the ACE2 protein from each of the 18 different species. Since binding of ACE2 to the virus S protein cannot be directly observed in two-dimensional representations, we modeled each enzyme’s tertiary structure. We assessed the locations of 10 key binding sites (hydrogen and disulfide) in the human ACE2 crystal structure on the surface of the ACE2 models from the 18 selected species. The results are shown in Figure 2 and Supplementary Table S2 available in www.besjournal.com. Macaca mulatta (rhesus monkey) and Homo sapiens (human) have the same ACE2 key binding sites, suggesting that SARS-CoV-2 infection in monkeys should be similar to that in humans. This conclusion is supported by the results from some research groups who have constructed a SARS-CoV-2 infection model using rhesus monkeys, and described the characteristics of rhesus monkey infection with COVID-19[6].
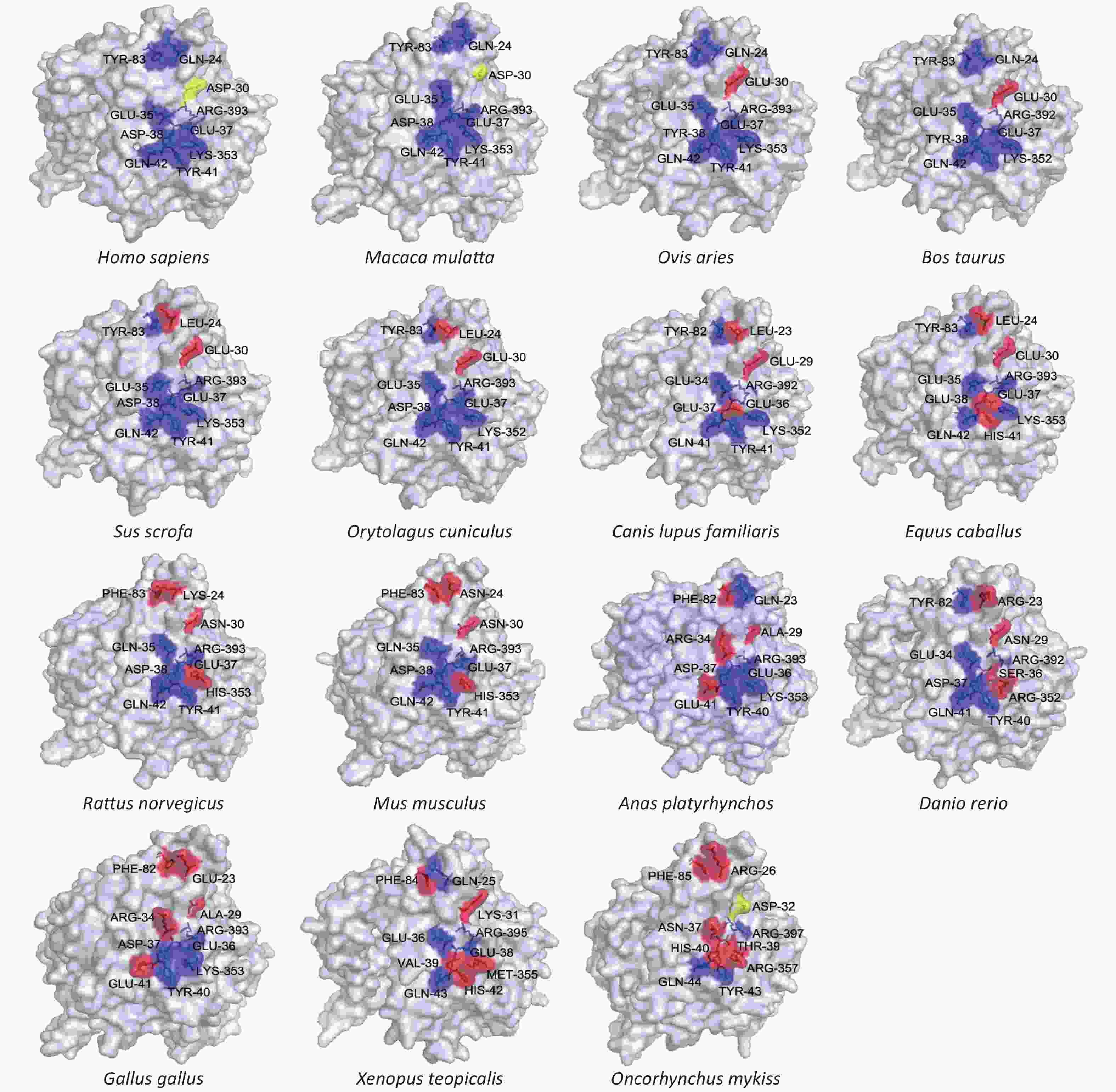
Figure 2. Tertiary structures of the extracellular domains of ACE2 from different species. Structures were predicted by homology modeling using multiple solved human ACE2 N-terminal extracellular domains as templates. Key binding sites (hydrogen and disulfide) on the surfaces of different species’ ACE2 structures were compared, using solved structures of human ACE2 as criteria. Blue regions represent conserved binding sites, and red regions represent absent binding sites.
Species Protein Binding Sites Q24 D30 E35 E37 D38 Y41 Q42 Y83 K353 R393 Human 1 1 1 1 1 1 1 1 1 1 Monkey 1 1 1 1 1 1 1 1 1 1 Bat 0 1 1 1 1 1 1 1 1 1 Sheep 1 0 1 1 1 1 1 1 1 1 Cattle 1 0 1 1 1 1 1 1 1 1 Cat 0 0 1 1 1 1 1 1 1 1 Pangolin 0 0 1 1 1 1 1 1 1 1 Pig 0 0 1 1 1 1 1 1 1 1 Rabbit 0 0 1 1 1 1 1 1 1 1 Dog 0 0 1 1 0 1 1 1 1 1 Horse 0 0 1 1 0 0 1 1 1 1 Rat 0 0 1 1 1 1 1 0 0 1 Mouse 0 0 1 1 1 1 1 0 0 1 Duck 1 0 0 1 1 1 0 0 1 1 Zerafish 0 0 1 0 1 1 1 1 0 1 Chook 0 0 0 1 1 1 0 0 1 1 Amphibian 1 0 1 1 0 0 1 0 0 1 Trout 0 1 0 0 0 1 1 0 0 1 Note. 0 represents different amino acid binding site, and 1 represents the same amino acid binding site. Table S2. Statistics of COVID-19 and ACE2 protein binding sites
For Ovis aries (sheep), Bos Taurus (cattle), and Sus scrofa (pig), the observed bonds between human ACE2 and SARS-CoV-2 are all present except at D30, whereas D30 is the key site for the formation of disulfide bonds during the binding process. No binding or very weak binding between Human ACE2 and SARS-CoV-2 indicates that loss of this binding epitope may decrease infection in these species. Sus scrofa (pig) and Oryctolagus cuniculus (rabbit) appear to be missing two binding sites, including the disulfide bond forming site.
In ACE2 from Canis lupus familiaris (dog), the one disulfide bond site, and two hydrogen bond sites present in human ACE2 are absent, which could destabilize SARS-CoV-2 binding to canine ACE2, reducing the possibility of infection in dogs. This result could explain a recent study which found that SARS-CoV-2 could be weakly detected in dogs by RT-PCR, but results were unable to conclusively show that a dog carried the virus. Equus caballus (horse), Rattus norvegicus (rat), Mus musculus (mouse), Anas platyrhynchos (duck), and Danio rerio (zebrafish) have 4 unconserved residues among the 10 assessed binding sites. Therefore, their likelihood of SARS-CoV-2 infection should be similar and low.
More than half of the sites in Gallus gallus (chicken), Xenopus tropicalis (amphibian), Danio rerio (zebrafish), and Oncorhynchus mykiss (trout) are inconsistent with SARS-CoV-2 binding epitopes in human ACE2, which implies a low possibility of infection for these species. Even if medical wastewater and virus-containing domestic sewage flows into external rivers, the probability of infection among poultry and aquatic organisms is probably very small.
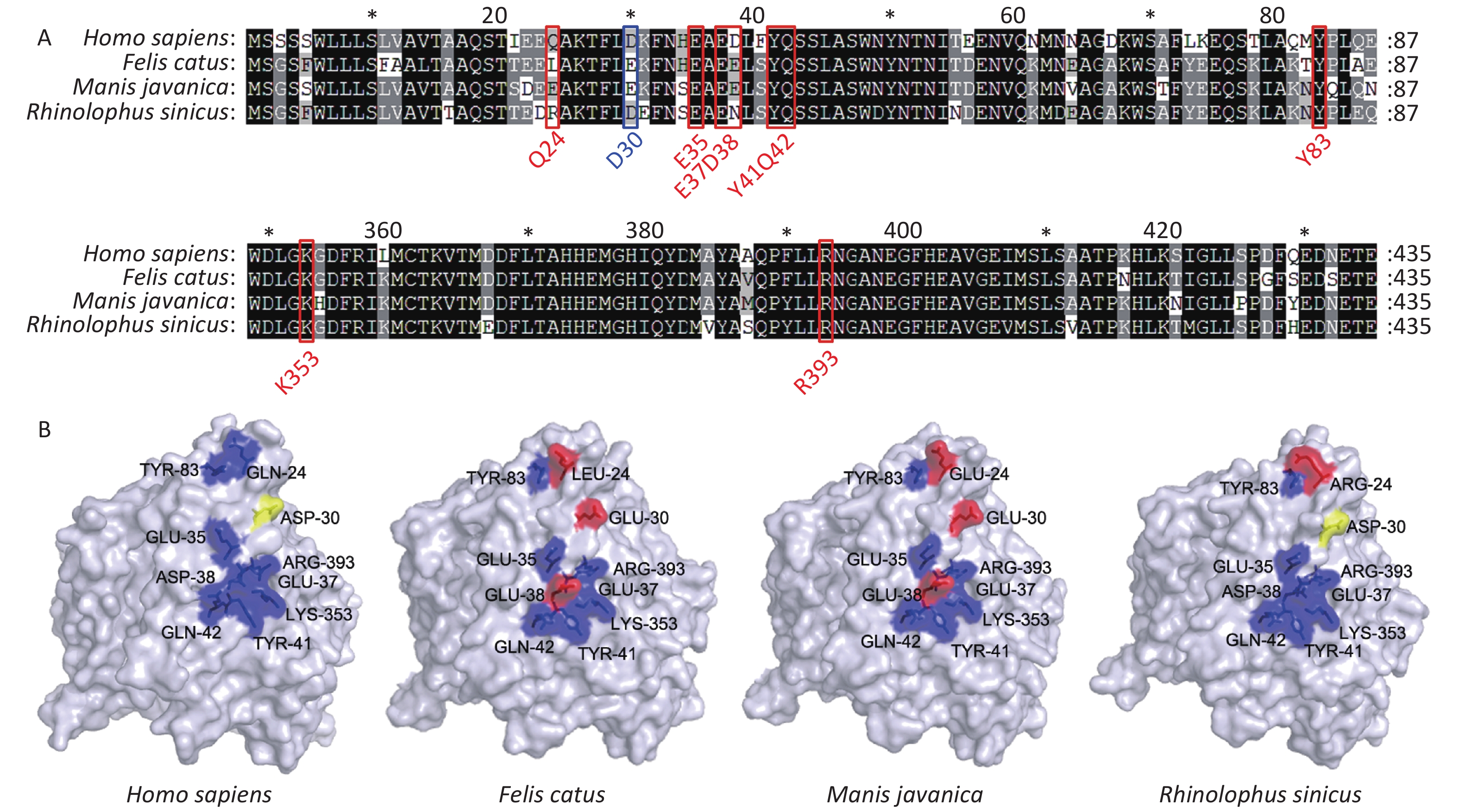
SARS-CoV-2 spread rapidly from animal species [possible sources include Manis javanica (pangolin) or Rhinolophus sinicus (bat)] to humans[7]. We thus analyzed ACE2 sequences from these putative intermediate hosts, and found that they have a high degree of similarity to human ACE2 at known binding sites. Bats display higher similarity to humans at examined binding sites than pangolins. Felis catus (cat) ACE2 displays the same residues as pangolins at all binding sites (Figure 3A–B).
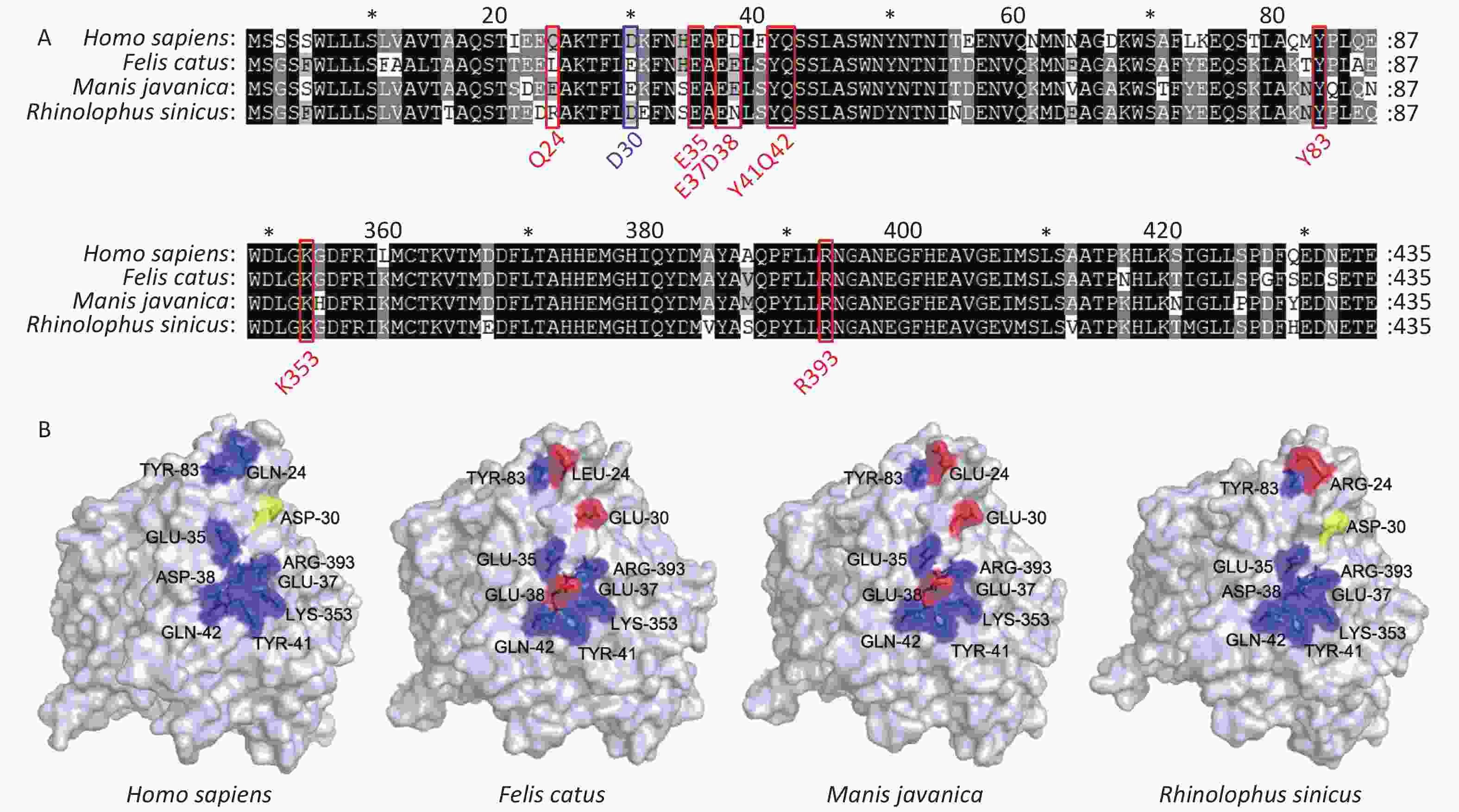
Figure 3. Comparison of Secondary and three-dimensional structures of pangolin, bat, cat, and human ACE2. (A) Alignment of the amino acid sequences of pangolin, bat, cat, and human ACE2. Residues outlined in red and blue form hydrogen bonds or a disulfide bond, respectively, with SARS-CoV-2 S protein. Blue, conserved relative to human; yellow, able to form a disulfide bond; red, not conserved relative to human. (B) Tertiary structures of the extracellular domains of ACE2 from pangolin, bat, cat, and human.
In summary, we used bioinformatics methods (protein secondary structure analysis, three-dimensional structure analysis, and phylogenetic tree construction) to analyze differences among 18 species in binding sites between ACE2 and the SARS-CoV-2 S protein. Our results suggest varying probabilities of SARS-CoV-2 infection among different species. Specifically, the probabilities of SARS-CoV-2 infection can be ranked as follows: Homo sapiens (human) ≈ Macaca mulatta (rhesus monkey) > Rhinolophus sinicus (bat) ≈ Ovis aries (sheep) ≈ Bos taurus (cattle) > Felis catus (cat) ≈ Manis javanica (pangolin) ≈ Sus scrofa (pig) ≈ Oryctolagus cuniculus (rabbit) > Canis lupus familiaris (dog) > Equus caballus (horse) ≈ Rattus norvegicus (rat) ≈ Mus musculus (mouse) ≈ Anas platyrhynchos (duck) ≈ Danio rerio (zerafish) > Gallus gallus (chook) ≈ Xenopus tropicalis (amphibian) ≈ Oncorhynchus mykiss (trout). This study provides a theoretical basis for research into whether SARS-CoV-2 may infect livestock, pets, poultry, and aquatic organisms that humans often come into contact with, and provides frames of reference for strengthening epidemic control through regulating exposure of non-human factors.
Previous studies have overexpressed the ACE2 receptor, the DDP4 receptor (MERS virus binding receptor), or the APN receptor (coronavirus HCoV-229E binding receptor) in the cell line BHK-21, which is not normally susceptible to SARS-CoV-2 infection. These studies showed that SARS-CoV-2 could only infect BHK-21 cells with the ACE2 receptor. ACE2 has been widely recognized as an essential receptor in viral invasion[4]. This essential role of ACE2 in viral infection allows similarities among species at ACE2 residues involved in SARS-CoV-2 binding to be used to predict the susceptibility of different species to SARS-CoV-2. Moreover, CD147, integrins, the endosomal cysteine protease cathepsin L (CTSL), and the serine protease TMPRSS2 can also facilitate SARS-CoV-2 entry into target cells[4, 8, 9]. Each different species will vary in its susceptibility to SARS-CoV-2 based on differences in the above factors. Recent research shows that SARS-CoV-2 replicates poorly in dogs, pigs, chickens, and ducks, but cats are permissive to infection[10]. Our results can guide the future construction of animal models of SARS-CoV-2 infection, which are expected to contribute to a better understanding of the pathophysiological mechanisms of SARS-CoV-2. As animal models continue to be constructed, we can distinguish which species may be infected with SARS-CoV-2, and take measures to cut off routes of animal transmission to humans. These models also provide researchers with convenient tools for choosing appropriate species to design optimal SARS-CoV-2 infection experiments. Subsequently, with further research, other novel factors, targeted genes, and proteases affecting the SARS-CoV-2 infection will also be revealed. Ultimately, we can find an effective method to control the spread of SARS-CoV-2.
Contributors XU XG and MAO GX designed this study; XU XG, WANG SY, and ZHANG J acquired the data; XU XG and SUN C drafted the manuscript; YAN J and WANG H critically revised the manuscript for important intellectual content; XU XG performed the statistical analysis assisted by WEI Ming Xiang.
Competing Interests No conflict of interest to declare.
Predictive Analysis of Susceptibility of Different Species to SARS-CoV-2 based on ACE2 Receptors
doi: 10.3967/bes2020.120
- Received Date: 2020-04-15
- Accepted Date: 2020-06-28
| Citation: | XU Xiao Gang, WANG Huan, WANG San Ying, ZHANG Jing, SUN Chuan, WEI Ming Xiang, YAN Jing, MAO Gen Xiang. Predictive Analysis of Susceptibility of Different Species to SARS-CoV-2 based on ACE2 Receptors[J]. Biomedical and Environmental Sciences, 2020, 33(11): 877-881. doi: 10.3967/bes2020.120 |








 Quick Links
Quick Links
 DownLoad:
DownLoad: