-
Periodontitis is a common oral disease that is characterized by the progressive destruction of periodontal supporting tissue and ultimately leads to tooth loss[1]. Diabetes mellitus is a common chronic disease characterized by elevated blood glucose levels as a result of reduced insulin secretion and/or insulin function[2]. Diabetes not only increases the degree of damage to periodontal tissue but also impairs the repair of periodontal tissue[3]. Current therapies, such as periodontal curettage and guided periodontal tissue regeneration, have a limited capacity for functional periodontal regeneration; thus, a more effective therapy is necessary[4, 5].
Periodontal ligament stem cells (PDLSCs) are important cell sources for periodontal regeneration as a result of their self-renewal and multidirectional differentiation abilities[6,7]. Osteogenic differentiation of PDLSCs is vital for the reconstruction of lost alveolar bone caused by periodontitis[1, 6]. However, the osteogenic differentiation ability of PDLSCs is inhibited under high glucose (HG) conditions[8-10]. Hence, the possibility of alleviating the inhibitory effect of HG on the osteogenic differentiation function of PDLSCs has become the key to the treatment of diabetic periodontitis.
Exosomes are extracellular vesicles secreted by cells with a diameter ranging from 50–200 nm[11, 12]. Exosomes can mediate intercellular communication by delivering miRNAs, mRNAs, and proteins to target cells. Studies have shown that exosomes possess functional properties similar to those of their parent cells and can be used as a substitute for mesenchymal stem cells (MSCs) in tissue repair, thereby avoiding the shortcomings of MSCs treatment, such as the low efficiency of cells homing, the possibility of immune rejection, and tumorigenic effects[13, 14]. Recent evidences have revealed that exosomes not only take part in tissue repair and regeneration but also play roles in disease diagnosis, tumor invasion and metastasis, immune regulation, and targeted drug delivery[15, 16]. Human umbilical cord mesenchymal stem cells (hUCMSCs) are derived from neonatal umbilical cord tissue and possess strong self-renewal and multidirectional differentiation capabilities[17]. hUCMSCs have become an ideal seed cell for tissue regeneration as a result of their rich sources, minimal immune rejection, fewer ethical issues, and noninvasive procedure used for harvest[18]. In terms of periodontal regeneration, studies have found that hUCMSCs effectively promote the regeneration of periodontal hard tissues and help form new functional periodontal attachments[19, 20]. For hUCMSC-derived exosomes (hUCMSC-exo), Yang et al.[21] found that hUCMSC-exo can accelerate the proliferation and migration of endothelial cells to promote angiogenesis and subsequent diabetic wound healing. Moreover, our previous study revealed that hUCMSC-exo could promote bone regeneration by enhancing the proliferation, migration, and osteogenic differentiation of preosteoblast cells[22]. Therefore, we speculated that hUCMSC-exo may be able to promote the osteogenic differentiation of PDLSCs under HG conditions and may be a promising candidate for promoting periodontal wound healing and bone regeneration in individuals with diabetes.
Herein, the current study aimed to explore the effect of hUCMSC-exo on the osteogenic differentiation of hPDLSCs under HG conditions and the related mechanism underlying this effect.
-
The study was approved by the Ethics Committee of Chinese PLA General Hospital. hUCMSCs were purchased from ScienCell Company (San Diego, CA, USA) and expanded in α-MEM (Hyclone, USA) containing 10% fetal bovine serum (FBS, Gibco, USA) and 10% penicillin/streptomycin (Invitrogen, USA) at 37 °C with 5% CO2. The hPDLSCs were cultured as previously described[7]. Periodontal ligaments were obtained from healthy premolars extracted due to orthodontics placement. Periodontal ligament tissues in the middle of the root were cut into 1 mm3 pieces, treated with type І collagenase (Sigma, USA) for 1 h, and then cultured in α-MEM containing 10% FBS and 1% penicillin/streptomycin. In osteogenic differentiation assays, the cells were cultured in osteogenic medium (α-MEM containing 10% FBS, 10 nmol/L dexamethasone, 50 μg/mL vitamin C, and 10 mmol/L β-glycerophosphate). According to previous studies, a 30 mmol/L D-glucose concentration was used to simulate HG conditions in vitro, and a 5.6 mmol/L D-glucose concentration was used as a control[23, 24].
-
The surface markers of hPDLSCs were detected using flow cytometry (BD Biosciences, USA). Antibodies against CD105-FITC, CD90-PE, CD73-FITC, CD44-PE, CD45-PE, CD34-FITC, CD31-FITC, and CD11b-PE were used in the analysis.
-
Exosomes were extracted using ultracentrifugation as described in our previous study[22]. Briefly, the culture medium was replaced with exosome-free FBS medium when the hUCMSCs reached 80%−90% confluence. After culturing for 48 h, the culture supernatant was harvested and centrifuged to remove dead cells. Subsequently, the supernatant was filtered through a 0.22 µm pore membrane and ultracentrifuged at 100,000 ×g for 90 min to obtain exosomes. Exosomes were identified using transmission electron microscopy (TEM, JEOL, Japan), Nanosight (Malvern, UK), and Western blot.
-
The exosomes were labeled with the red red fluorescent dye Dil (Beyotime, China) as described in our previous study and then cocultured with PDLSCs for 24 h. Subsequently, the cells were washed with phosphate-buffered saline (PBS), fixed with 4% paraformaldehyde and stained with DAPI (Solarbio, China). The uptake of exosomes by hPDLSCs was observed using laser scanning confocal microscopy.
-
The cells were divided into four groups for cell proliferation and differentiation assays: (i) control group, cells treated with 5.6 mmol/L D-glucose; (ii) HG group, cells treated with 30 mmol/L D-glucose; (iii) HG + 25 µg/mL exo group, cells treated with 30 mmol/L D-glucose and 25 µg/mL exosomes; (iiii) HG + 50 µg/mL exo group, cells treated with 30 mmol/L D-glucose and 50 µg/mL exosomes. The Cell Counting Kit-8 (CCK-8) assay was conducted to detect the proliferation of hPDLSCs on Days 1, 3, and 5 following the manufacturer’s instructions.
-
ALP staining and ALP activity were assessed as reported previously[22]. Briefly, after osteoinduction for 14 days, hPDLSCs were fixed with paraformaldehyde and stained with an ALP staining kit (Solarbio, China) for 30 min. ALP activity was detected with an ALP kit (Nanjing Jiancheng Biotech, Nanjing, China) following the manufacturer’s instructions.
-
ARS staining was conducted to assess mineral deposition in hPDLSCs after osteoinduction for 14 days. Cells were washed with PBS, fixed with paraformaldehyde and stained with an ARS solution (Solarbio, China) for 20 min following the manufacturer’s instructions.
-
qRT-PCR was performed as reported in our previous study[22]. Briefly, total RNA was extracted using TRIzol (Invitrogen, USA), and cDNAs were synthesized using a cDNA synthesis kit (Takara, Japan). Then, qRT-PCR was performed using an ABI Prism 7900 system with SYBR Master Mix (Takara, Japan). The primers used for qRT-PCR were referred to previous studies[6, 7, 25], and the sequences are shown in Table 1. β-actin was used as a reference gene for internal normalization.
Gene Forward (5’–3’) Reverse (5’–3’) ALP GTGAACCGCAACTGGTACTC GAGCTGCGTAGCGATGTCC OCN AGCAAAGGTGCAGCCTTTGT GCGCCTGGGTCTCTTCACT Runx2 CACTGGCGCTGCAACAAGA CATTCCGGAGCTCA
GCAGAATAβ-actin ATGCCAACACAGTGTTGTCTGG TACTCCTGCTTGCT
GATCCACATTable 1. Primers used for qRT-PCR
-
Western blot analyses were performed as reported in our previous study[22]. Antibodies used were: CD9 (Affinity, USA), CD63 (NOVUS, USA), TSG101 (Affinity, USA), ALP (Affinity, USA), Runx2 (Affinity, USA), OCN (Affinity, USA), OPN (Proteintech Group, USA), p-AKT (Cell Signaling Technology, USA), AKT (Affinity, USA), p-PI3K (Cell Signaling Technology, USA), PI3K (Cell Signaling Technology, USA), and β-actin (Boster Bioengineering Co., Wuhan, China). The results were normalized to β-actin.
-
The PI3K/AKT signaling inhibitor LY294002 and AKT inhibitor MK2206 was were used to investigate the role of the PI3K/AKT signaling pathway in the exosome-mediated osteogenic differentiation of hPDLSCs. The cells were divided into 5 groups: the control group (cells treated with 5.6 mmol/L D-glucose), HG group (cells treated with 30 mmol/L D-glucose), HG + exo group (cells treated with 30 mmol/L D-glucose and 50 µg/mL exosomes), and HG + exo + LY294002 group (cells treated with 30 mmol/L D-glucose, 50 µg/mL exosomes, and 10 µmol/L LY294002), and HG + exo + MK2206 group (cells treated with 30 mmol/L D-glucose, 50 µg/mL exosomes, and 1 µmol/L MK2206). CCK-8 assay, ALP staining, ARS staining, and Western blot analysis were conducted as described above.
-
All data are presented as the means ± standard deviations of at least 3 experiments per group. SPSS 19.0 software (Chicago, IL, USA) was used to analyze the data. One-way ANOVA, followed by Tukey’s multiple comparison test was used to analyze differences between groups, and P < 0.05 was considered statistically significant.
-
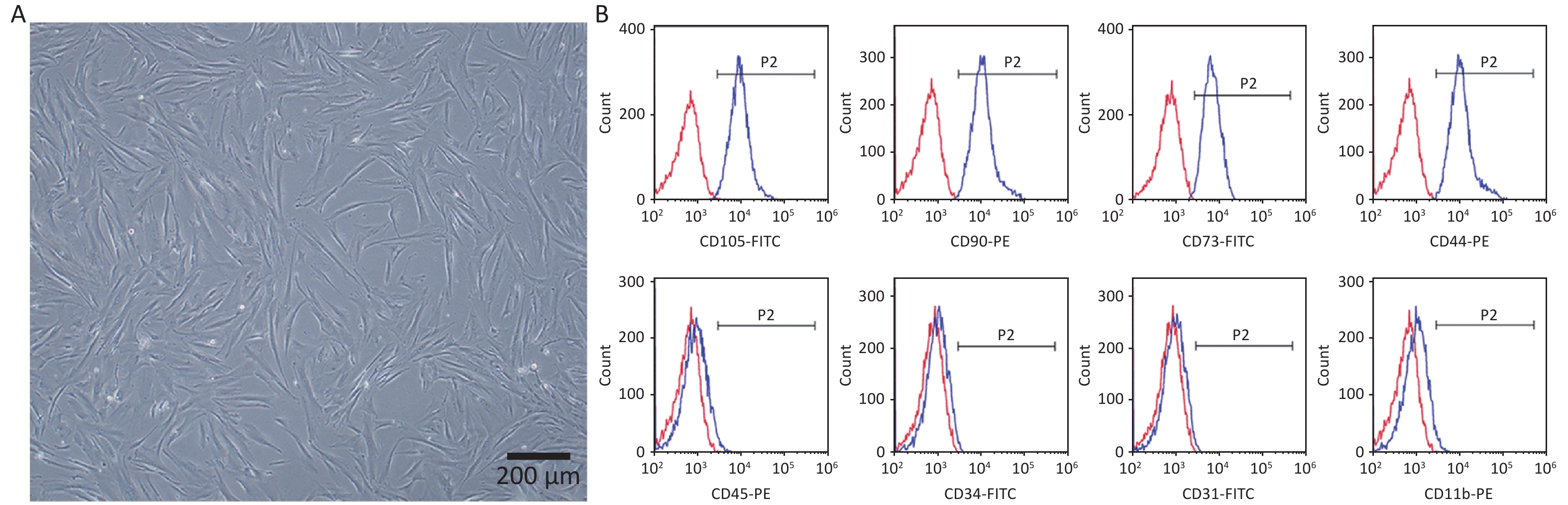
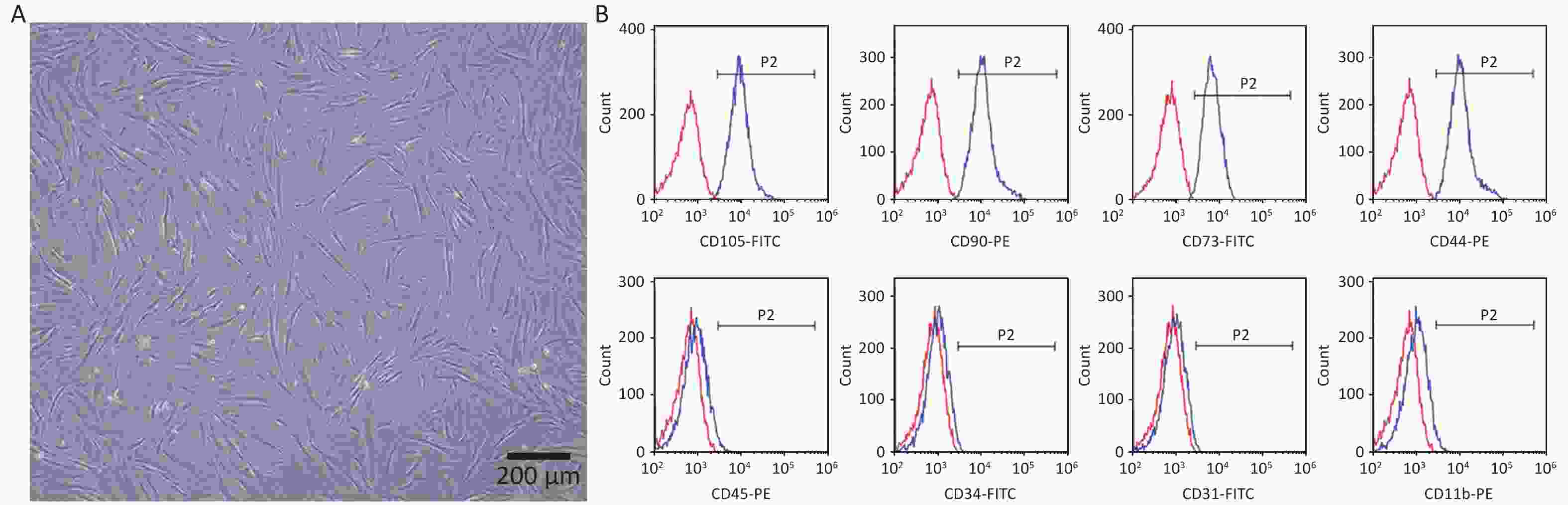
The hPDLSCs were successfully extracted from periodontal tissues and presented a long spindle shape (Figure 1A). The flow cytometry results revealed that the cells positively expressed the MSCs surface markers CD105-FITC (99.01% ± 0.30%), CD90-PE (98.92% ± 0.29%), CD73-FITC (98.03% ± 0.56%), CD44-PE (98.83% ± 0.61%), negatively expressed the hematopoietic surface markers CD45-PE (2.17% ± 0.49%), CD34-FITC (0.98% ± 0.25%), CD31-FITC (2.28% ± 1.15 %), and CD11b-PE (3.03% ± 0.60%) (Figure 1B). All the results revealed that we successfully obtained hPDLSCs.
-
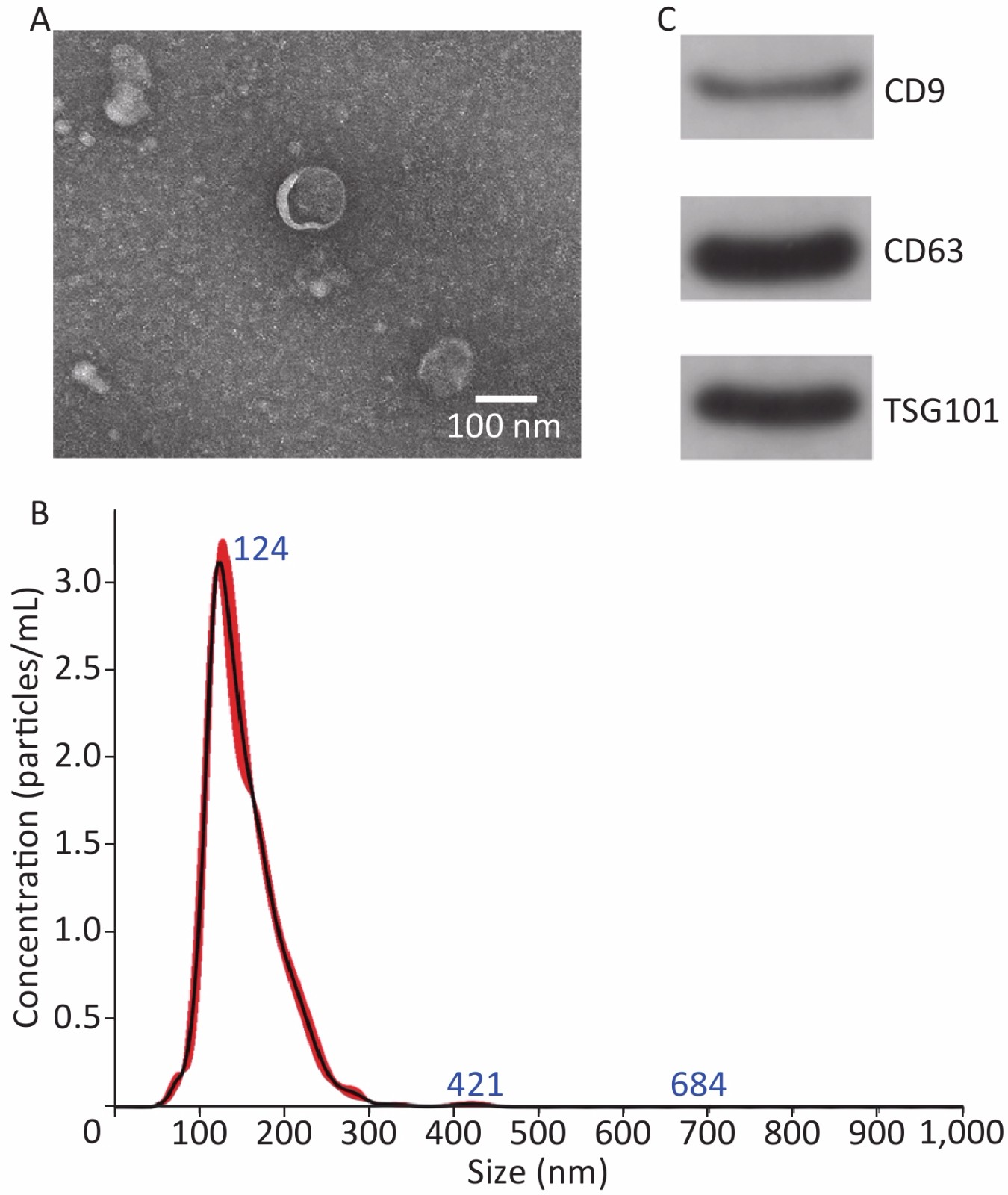
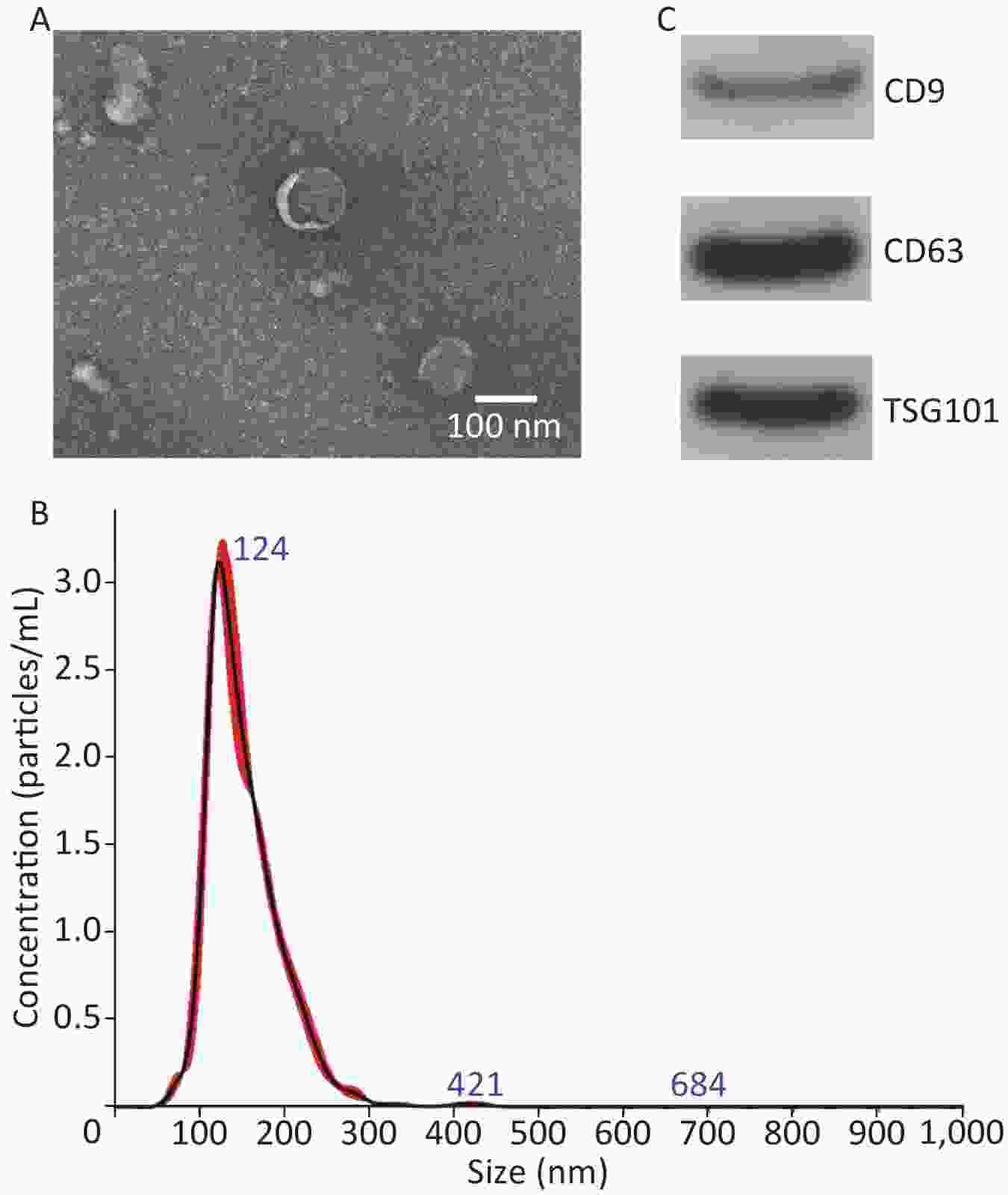
TEM, NanoSight, and Western blot were performed to identify exosomes. The TEM results revealed that the exosomes mainly exhibited a cup-shaped morphology, with a diameter of approximately 100 nm (Figure 2A). NanoSight results revealed that the exosomes size distribution ranged from 50 to 200 nm, with a peak at 124 nm (Figure 2B). Western blot results revealed that exosomes expressed CD9, CD63 and TSG101 (Figure 2C). These results are consistent with the relevant characteristics of exosomes, indicating that we successfully extracted exosomes.
-
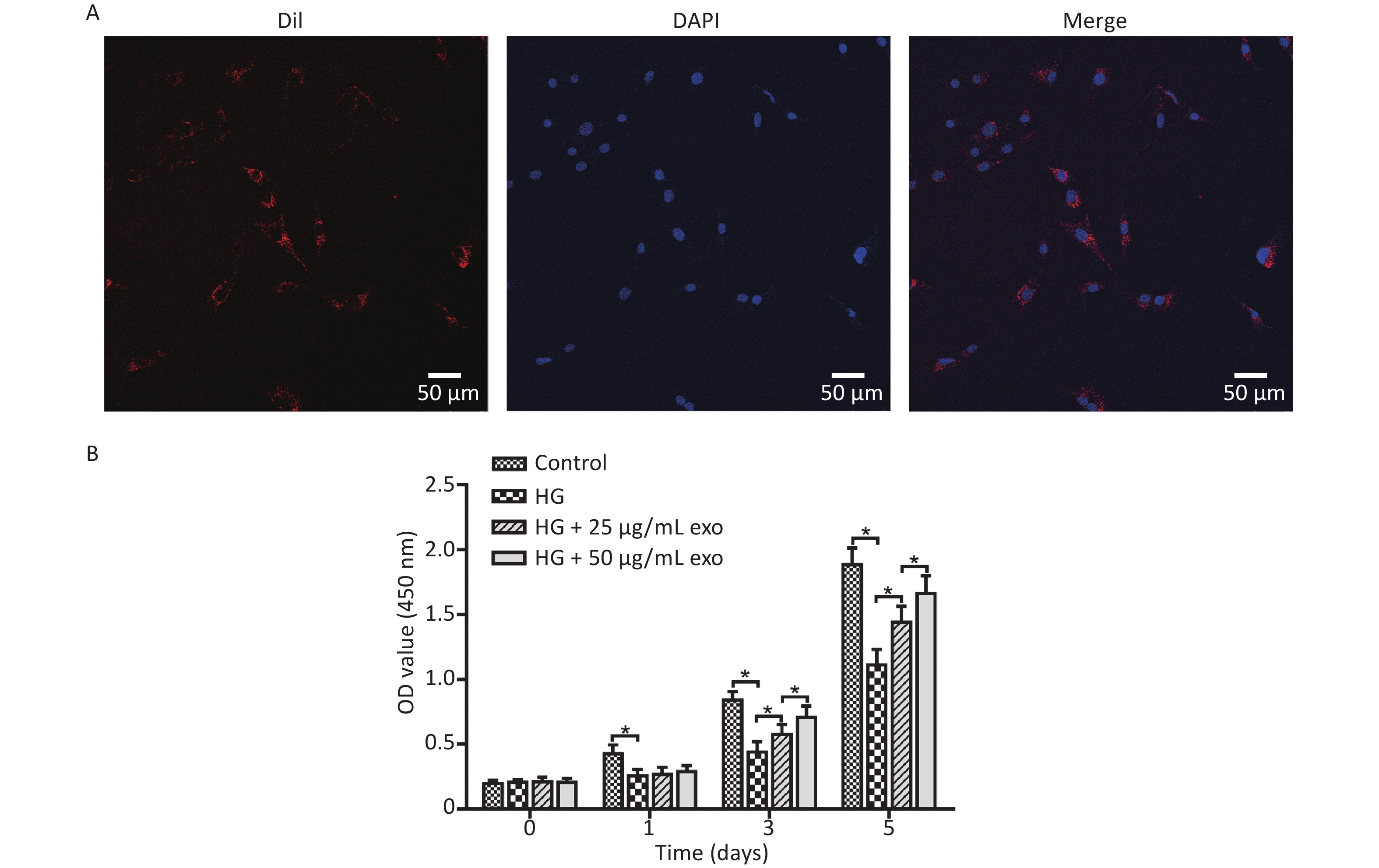
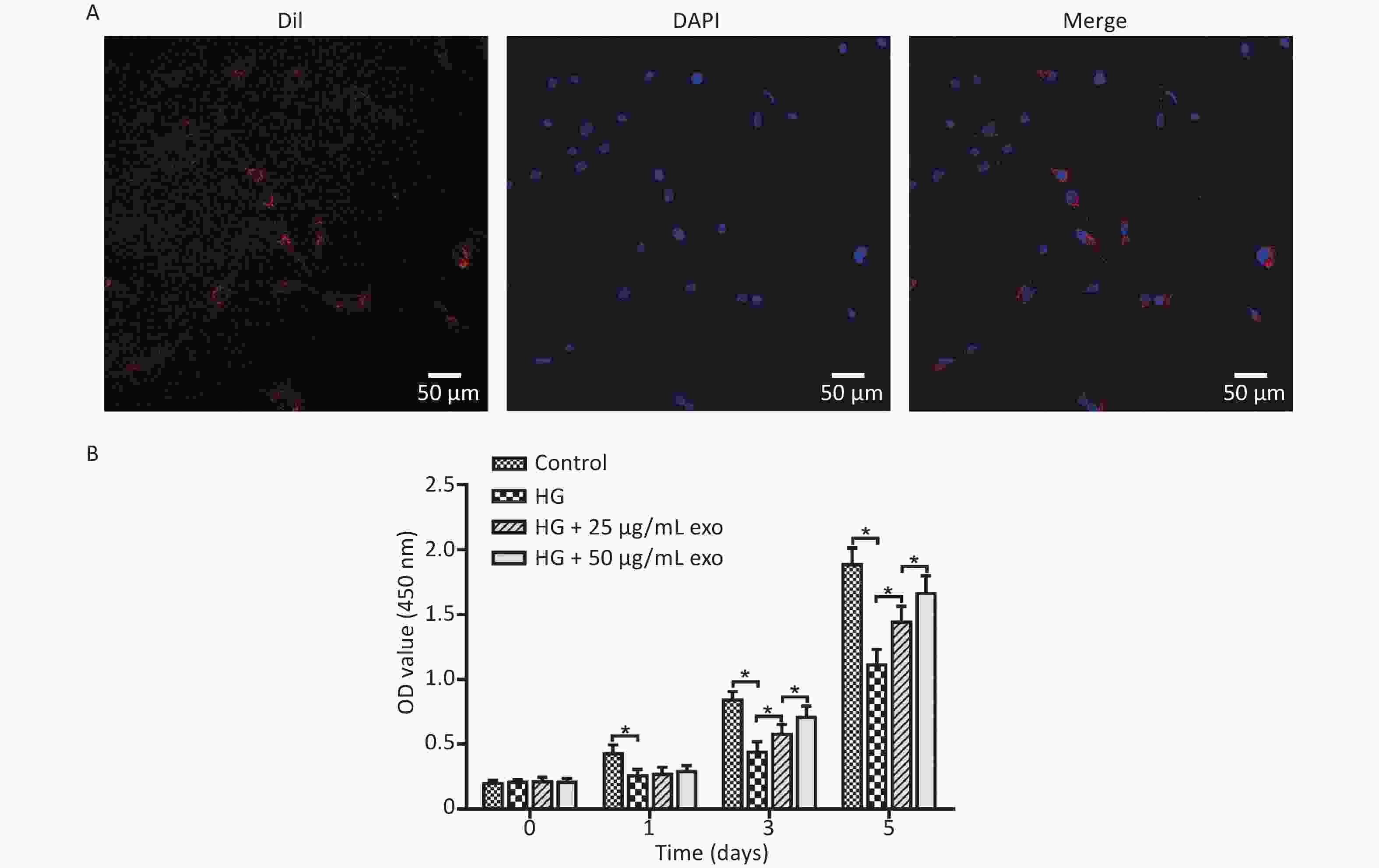
After the cocultivation of exosomes and hPDLSCs for 24 hours, laser confocal microscopy showed that a large amount of red fluorescence appeared near the blue-labeled nucleus of hPDLSCs, indicating that the exosomes were successfully taken up by hPDLSCs (Figure 3A). The effect of different concentrations of exosomes on the proliferation of hPDLSCs cultured under HG conditions was detected using the CCK-8 assay. The results revealed that HG conditions significantly inhibited the proliferation of hPDLSCs on Days 1, 3, and 5 (Figure 3B). Moreover, the addition of exosomes significantly promoted cell proliferation on Days 3 and 5 in a dose-dependent manner.
-
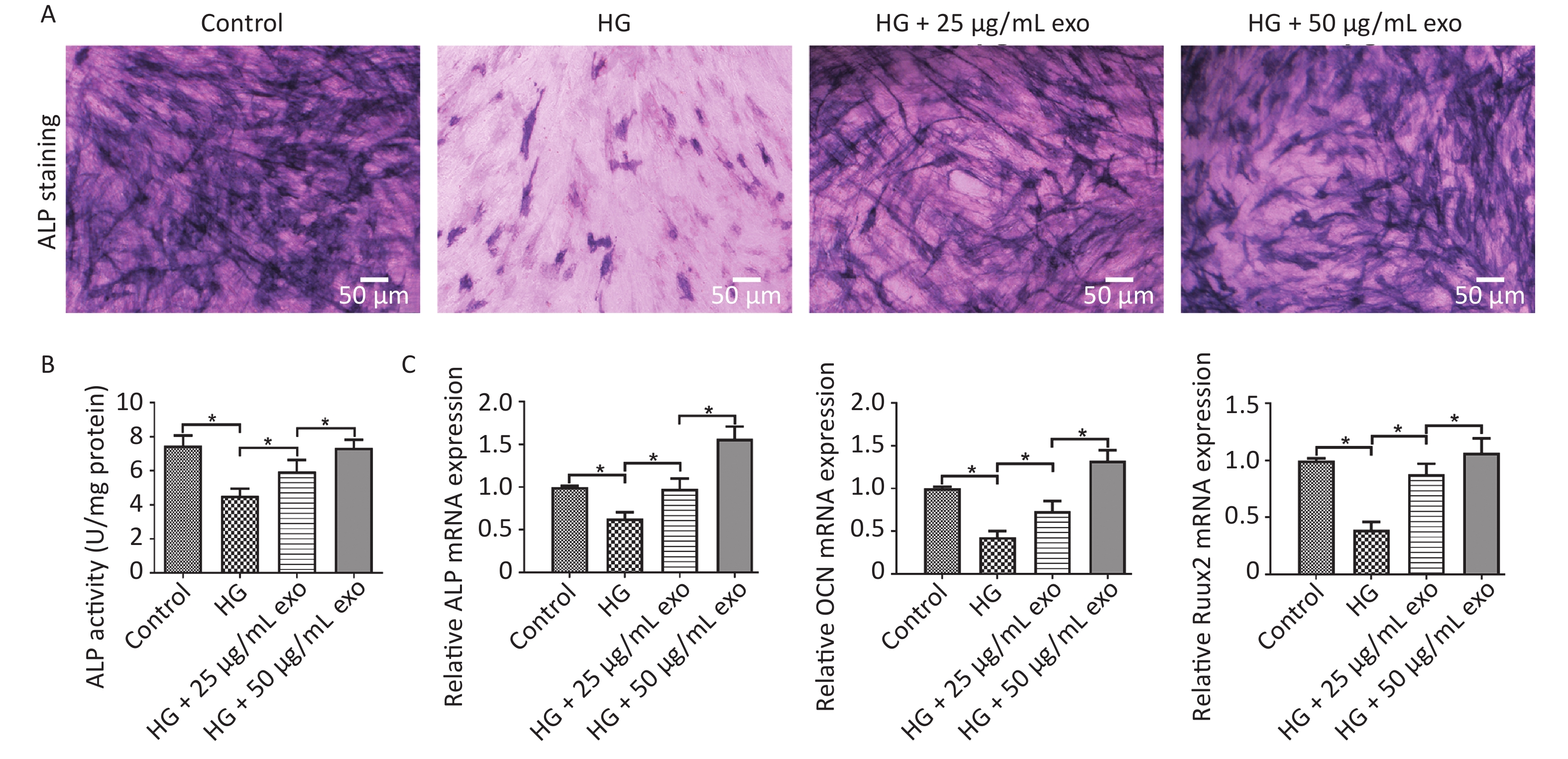
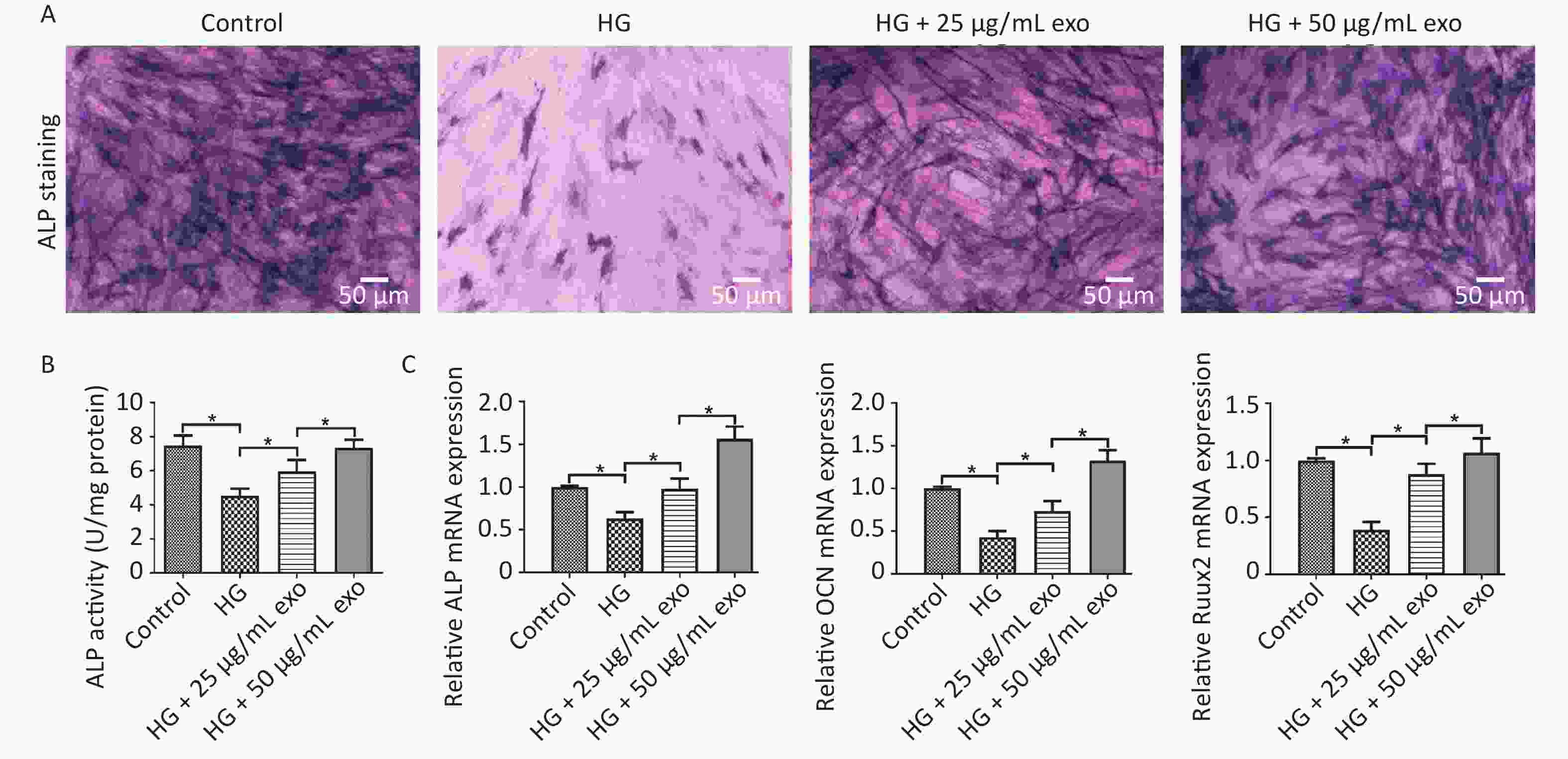
ALP staining, ALP activity and qRT-PCR were performed to evaluate the pro-osteogenic effect of hUCMSC-exo on hPDLSCs under HG conditions. The results showed that HG significantly inhibited the osteogenic differentiation of hPDLSCs (Figure 4). However, similar to the effect of HG on cell proliferation, exosomes reversed the inhibitory effect of HG on the osteogenic differentiation of hPDLSCs in a dose-dependent manner (Figure 4). Since 50 µg/mL exosomes exerted a better pro-osteogenic effect on hPDLSCs, we chose this concentration for subsequent experiments.
-
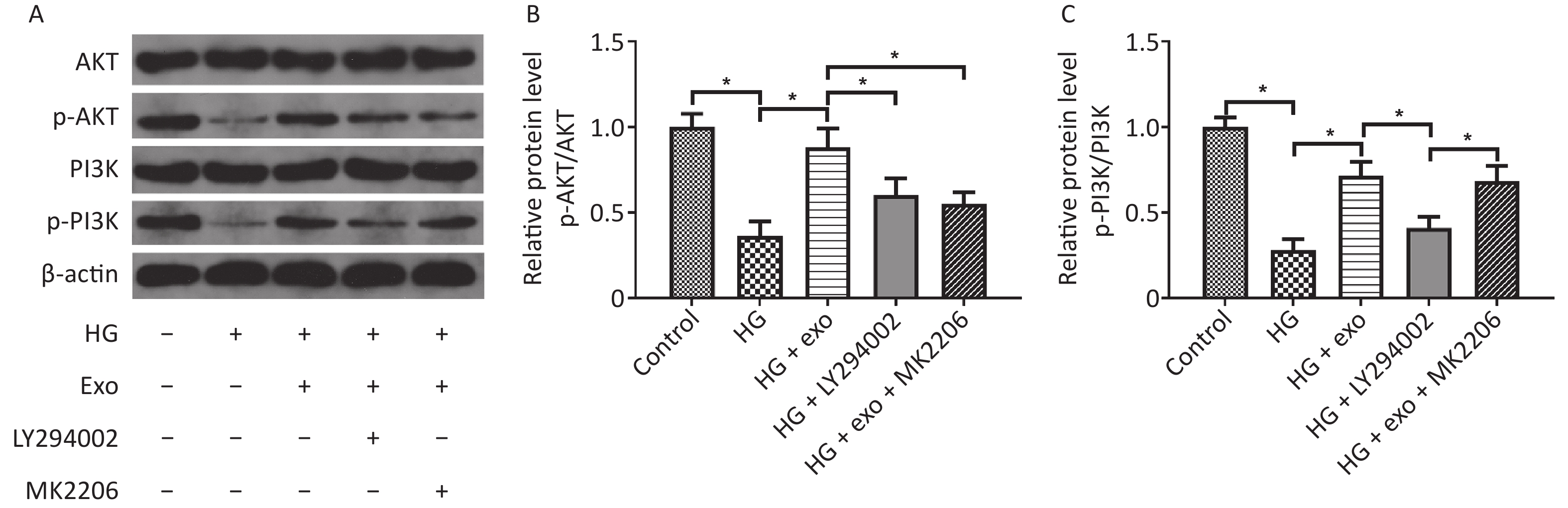
We used western blot analysis to assess the levels of the PI3K/AKT signaling pathway-related proteins AKT, p-AKT, PI3K, and p-PI3K and to explore whether the PI3K/AKT signaling pathway was activated during the process by which exosomes promoted the osteogenic differentiation of PDLSCs under HG conditions. The results showed that HG decreased the levels of p-AKT and p-PI3K, while exosomes increased the levels of these proteins, suggesting that exosomes activated the PI3K/AKT signaling pathway during the process (Figure 5). However, the increases in p-AKT and p-PI3K levels were suppressed following the addition of the PI3K inhibitor LY294002 (Figure 5). What’s more, the addition of AKT inhibitor MK2206 decreased the elevated levels of p-Akt, but not p-PI3K.
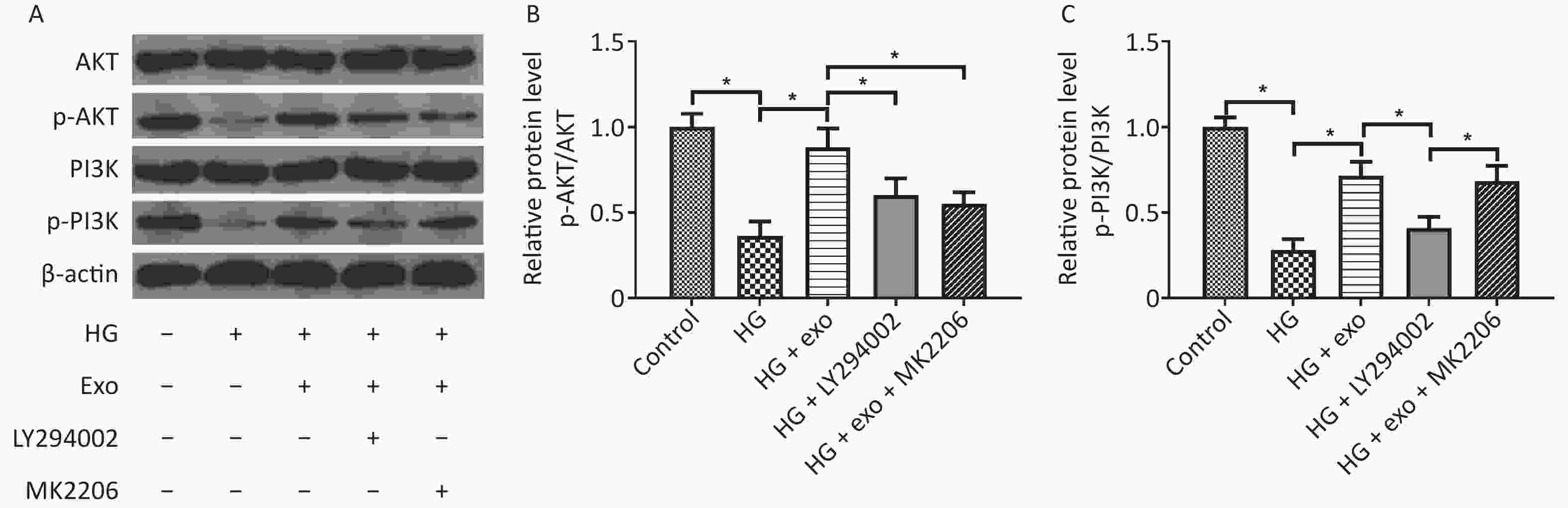
Figure 5. hUCMSC-exo promoted the osteogenic differentiation of hPDLSCs under HG conditions by activating the PI3K/AKT signaling pathway. (A) Levels of the AKT, p-AKT, PI3K, and p-PI3K proteins. (B) Quantitative analysis of p-AKT/AKT. (C) Quantitative analysis of p-PI3K/PI3K. HG, high glucose. *P < 0.05.
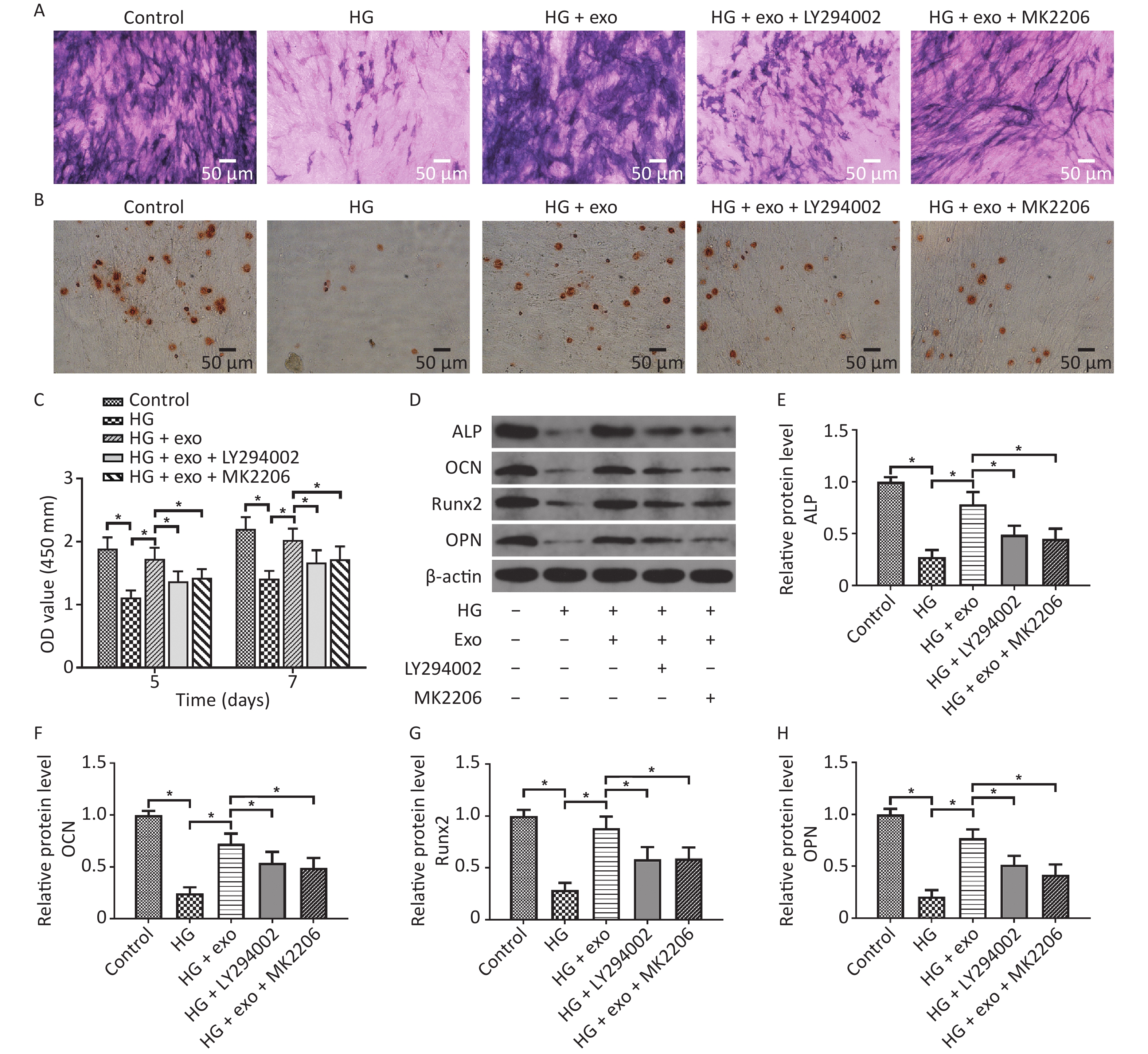
LY294002 and MK2206 were then utilized. We performed ALP staining, ARS staining, and Western blot assays after osteoinduction for 14 days to further explore the role of the PI3K/AKT signaling pathway in the pro-osteogenic effect of exosomes under HG conditions. The CCK-8 results showed that hUCMSC-exo improved the reduced proliferation ability of hPDLSCs on Days 5 and 7. Similarly, the ALP staining, ARS staining, and Western blot assays results revealed that hUCMSC-exo reversed the inhibitory effect of HG on the osteogenic differentiation of hPDLSCs (Figure 6). However, this change was these changes were significantly suppressed by LY294002 and MK2206 (Figure 6). Taken together, these results indicated that the activation of the PI3K/AKT signaling pathway may be the underlying mechanism by which hUCMSC-exo promote the osteogenic differentiation of hPDLSCs under HG conditions.
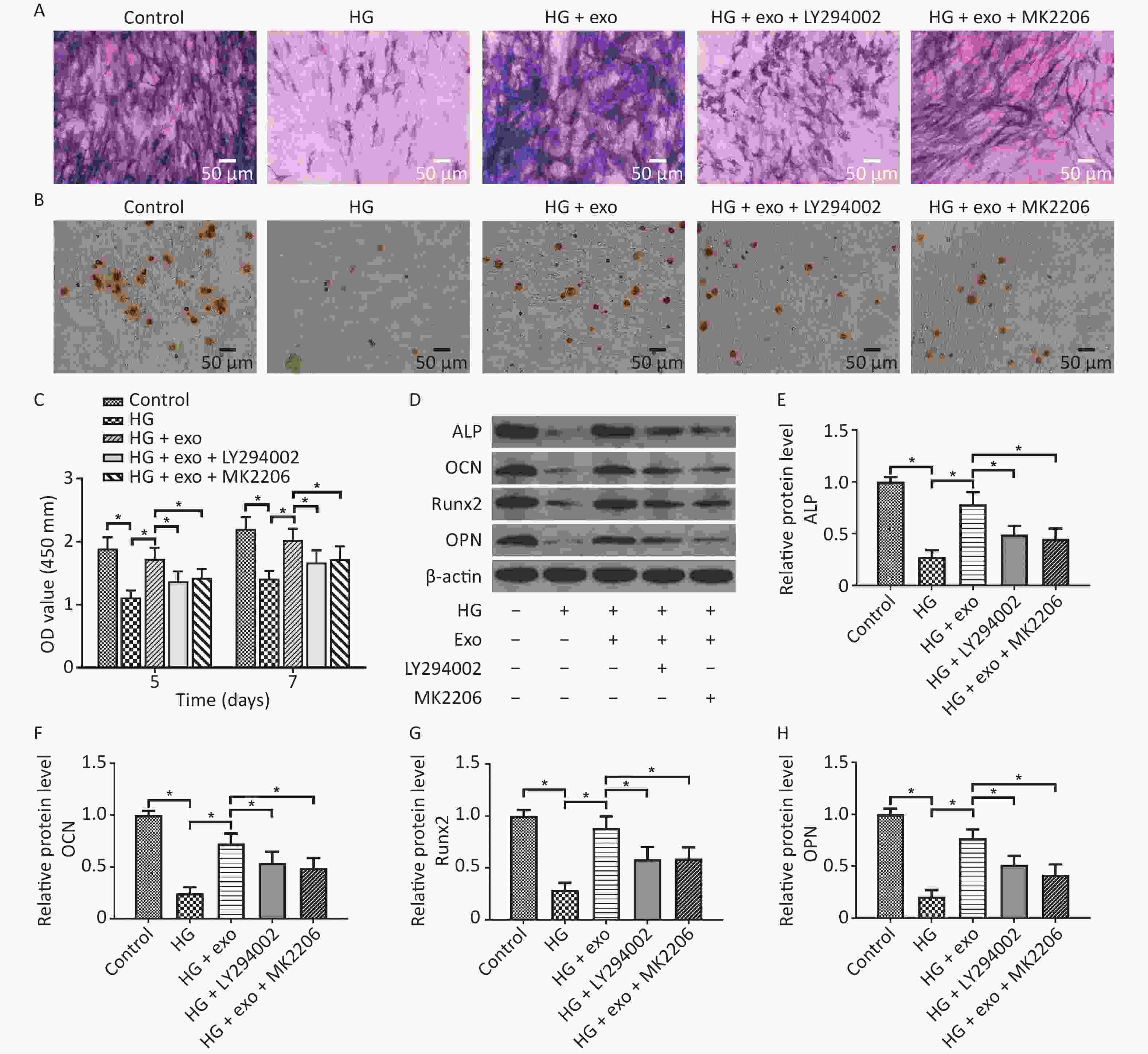
Figure 6. LY294002 and MK2206 inhibited the pro-osteogenic effect of exosomes on hPDLSCs cultured under HG conditions. (A) ALP staining. (B) ARS staining. (C) CCK-8 results on Days 5 and 7. (D) The ALP, OCN, OPN, and Runx2 proteins expression. (E)–(H) Quantitative analysis of ALP, OCN, Runx2, and OPN levels. HG, high glucose. *P < 0.05.
-
In recent years, the prevalence of diabetes mellitus has increased rapidly worldwide, and diabetes mellitus has become a serious public health concern[3]. High blood glucose levels in patients with diabetes can cause many complications, such as poor wound healing, bone disease, cardiovascular disease, stroke, kidney disease, and retinopathy[26, 27]. Diabetes is also a risk factor for periodontitis[28]. High blood glucose levels may exacerbate bone loss in patients with periodontitis and ultimately cause tooth loss[29]. hPDLSCs have been considered the key cells to promote periodontal regeneration. Numerous studies have indicated that the function of PDLSCs is inhibited in an HG environment, especially the function of osteogenic differentiation, which is an important cause of limited periodontal regeneration in diabetes patients[8, 9, 24].
In recent years, MSCs have shown good repair effects on periodontal regeneration and have attracted increasing attention[19]. According to recent studies, the role of MSCs in tissue repair mainly depends on their paracrine functions[30, 31]. Exosomes are extracellular vesicles secreted by cells and are the main paracrine factors[31]. They influence the function of target cells by transferring miRNAs, mRNAs, and proteins and show great potential in tissue repair and disease treatment[11]. At present, vari ous exosomes from different cell sources have been used in periodontal regeneration. Mohammed et al.[32] injected exosomes derived from adipose-derived stem cells (ADSCs) into the periodontal pockets of rats with ligature-induced periodontitis. The results of histological staining showed that the defect area formed a structure similar to that of healthy periodontal tissue. Additionally, both ADSCs and their derived exosomes exhibited certain anti-inflammatory and immunomodulatory activities and induced tissue regeneration by promoting the migration, differentiation, proliferation, and angiogenesis of different cell types. Chew et al.[33] loaded exosomes secreted by human embryonic stem cells on a collagen sponge, placed them on rat periodontal defects, and observed the regeneration of alveolar bone and functional periodontal ligament fibers. Further studies confirmed that exosomes promote the migration and proliferation of PDLSCs and promote periodontal regeneration through the CD73-mediated AKT/ERK signaling pathway.
hUCMSCs have become an ideal source of exosomes due to their advantages, such as noninvasive procedures used for harvest, convenient in vitro isolation and culture, and extensive sources[34]. Various studies have shown that hUCMSC-exo have broad application prospects in the treatment of cardiovascular diseases, bone diseases, wound healing, kidney diseases and other diseases[31, 34]. These findings suggest the potential application of hUCMSC-exo in periodontal regeneration. Recently, many studies have explored the effect of exosomes on hPDLSCs, but studies examining the effect of exosomes on hPDLSCs under HG conditions are still lacking. Whether hUCMSCs-exo could promote the osteogenic differentiation of hPDLSCs under HG condition is uncertain. In the present study, we successfully extracted and identified hPDLSCs and hUCMSC-exo. We labeled exosomes with the red fluorescent dye Dil and then cocultured them with hPDLSCs to observe the direct effect of exosomes on PDLSCs. After 24 hours, a large amount of red fluorescence appeared around the nucleus of the cells, indicating that the exosomes were internalized by the cells. Then, we administered a 30 mmol/L glucose to simulate HG conditions for in vitro experiments, and this glucose concentration has been widely used to study the effect of HG on cells in vitro[23, 24]. We first observed the effect of exosomes on cell proliferation, as this process may affect the osteogenic differentiation of cells. The CCK-8 results showed that exosomes effectively alleviate the inhibitory effect of HG on the proliferation of hPDLSCs, and this effect becomes more obvious as the concentration of exosomes increases, which were consistent with previous study[17]. Subsequently, we performed ALP staining and ALP activity assays to explore the pro-osteogenic effect of exosomes on hPDLSCs, as ALP is an early osteogenic differentiation marker of cells. The results revealed that hUCMSC-exo inhibited the HG-induced decrease in ALP activity. The qRT-PCR results showed that exosomes promoted the expression of the osteogenic-related genes ALP, OCN, and Runx2. These results further confirmed that hUCMSC-exo effectively promoted the osteogenic differentiation of PDLSCs under HG conditions.
The PI3K/AKT signaling pathway is involved in the regulation of cell proliferation, differentiation, and metabolism[1]. Additionally, the PI3K/AKT signaling pathway plays an important role in bone development and bone remodeling. Zhai et al.[35] demonstrated that human MSC-derived exosomes induce osteogenic differentiation by activating the PI3K/AKT and MAPK pathways. Xiong et al.[7] revealed that curcumin promotes the osteogenic differentiation of hPDLSCs through the PI3K/AKT/Nrf2 signaling pathway. Thus, in the present study, we explored whether hUCMSC-exo promoted the osteogenic differentiation of hPDLSCs under HG conditions through the PI3K/AKT signaling pathway. Western blot results showed that hUCMSC-exo significantly increased the levels of p-PI3K and p-AKT in cells cultured under HG conditions, and the effect was inhibited by LY294002, an inhibitor of the PI3K/AKT signaling pathway. Moreover, the increased levels of osteogenic indicators, such as ALP staining, ARS staining, and osteogenic-related proteins, induced by hUCMSC-exo were significantly suppressed by LY294002 and MK2206. These results suggested that hUCMSC-exo promoted the osteogenic differentiation of hPDLSCs cultured under HG condition through the PI3K/AKT signaling pathway. It is worth noting that either LY294002 or MK2206 led to the suppression of the increase in cell proliferation, whose results may affect the osteogenic differentiation of hPDLSCs to some extent.
This study has some limitations. First, the effect of hUCMSC-exo on the osteogenic differentiation of hPDLSCs under HG conditions was studied in vitro, and further in vivo studies are needed. Second, intracellular signal transduction is a complex regulatory network. Although the current study confirmed that the PI3K/AKT signaling pathway is involved in the process by which exosomes promote the osteogenic differentiation of hPDLSCs under HG conditions, we did not clearly determine whether other signaling pathways, such as the p38/MAPK, ERK, JNK, and Wnt/β-catenin pathways, are involved. For example, Wnt/β-catenin is the classic signaling pathway that mediates the osteogenic differentiation of hPDLSCs. Further studies are needed to determine whether Wnt/β-catenin signal transduction is involved in this process and its their relationship with the PI3K/AKT signaling pathway. Third, the exosomal contents play important roles in exosome regulation, and thus further studies are required to reveal the exosomal contents for in-depth evaluation of the mechanism of hUCMSC-exo in diabetic periodontitis.
In conclusion, hUCMSC-exo promoted the osteogenic differentiation of hPDLSCs under HG conditions through the PI3K/AKT signaling pathway, and thus the application of hUCMSC-exo may be a potential treatment for diabetic periodontitis.
-
The authors have no conflicts of interest to declare.
Exosomes Derived from Human Umbilical Cord Mesenchymal Stem Cells Enhance the Osteoblastic Differentiation of Periodontal Ligament Stem Cells Under High Glucose Conditions Through the PI3K/AKT Signaling Pathway
doi: 10.3967/bes2022.105
- Received Date: 2021-10-08
- Accepted Date: 2022-04-04
-
Key words:
- Exosomes /
- Human umbilical cord mesenchymal stem cell /
- Periodontal ligament stem cell /
- Osteogenic differentiation /
- High glucose /
- PI3K/AKT
Abstract:
| Citation: | YANG Shuo, ZHU Biao, TIAN Xiao Yu, YU Han Ying, QIAO Bo, ZHAO Li Sheng, ZHANG Bin. Exosomes Derived from Human Umbilical Cord Mesenchymal Stem Cells Enhance the Osteoblastic Differentiation of Periodontal Ligament Stem Cells Under High Glucose Conditions Through the PI3K/AKT Signaling Pathway[J]. Biomedical and Environmental Sciences, 2022, 35(9): 811-820. doi: 10.3967/bes2022.105 |







 Quick Links
Quick Links
 DownLoad:
DownLoad: