-
During the course of industrialization, particulate matter has become a common pollutant. The International Agency for Research on Cancer has classified the particulate matter in outdoor air pollution as a group 1 carcinogen [1]. In urban areas, motor vehicle emissions and fuel burning are major sources of particulate matter air pollution [2, 3]. Fine particulate matter (PM2.5), whose aerodynamic diameter is smaller than 2.5 μm, can adsorb a wide variety of toxic compounds (such as organic compounds and poisonous metals). PM2.5 can penetrate deeply into the lungs through breathing, cross the blood-air barrier and cause lesions in multiple organs, particularly the lungs [4, 5].
According to the World Cancer Report published by the International Agency for Research on Cancer, lung cancer was the leading cause of cancer-associated death worldwide in 2020 [6]. In addition, lung cancer is the most common cause of cancer disability-adjusted life years [7]. Metastasis is a major cause of the high mortality in lung cancer; however, the mechanisms of lung cancer metastasis have not been fully elucidated [8, 9]. Many studies have reported adverse effects of PM2.5 on lung cancer occurrence and development. The Harvard Six Cities study has shown that PM2.5 exposure increases the risk of lung cancer death (RR, 1.27; 95% CI, 0.96–1.69), and decreasing PM2.5 concentrations is critical to diminish lung cancer mortality [10]. Chao et al. have found that PM2.5 exposure contributed to proliferation, migration and invasion in lung cancer cell lines, and promoted tumor growth and metastasis in vivo [11].
Exosomes are extracellular vesicles with a diameter of approximately 40–160 nm, containing many constituents of cells, such as DNA, RNA, lipids and metabolites [12]. Exosomes are secreted by most cells and have been detected in a variety of body fluids including blood, urine, saliva, lymph and bronchoalveolar fluids [13-15]. Exosomes secreted by cells can be taken up by adjacent cells or can enter the circulation and affect distant cells [15, 16]. Numerous studies have focused on the biology and function of exosomes in cancers. Exosomal miR-21-5p derived from gastric cancer cells has been found to promote peritoneal metastasis [17]. The prometastatic secretome can be delivered by exosomes and lead to breast cancer multi-organ metastasis [18]. Moreover, exosomal miRNAs have been found to boost lung metastasis via STAT3-induced epithelial-to-mesenchymal transition (EMT) in vitro and in vivo[19]. In a review on the function of exosomes in tumors, Zhang et al. have noted that exosomes played crucial roles in tumor pathogenesis and progression [20].
Bronchial epithelial cells are located at the lung-environment interface and directly contact PM2.5. We hypothesized that signals of PM2.5 exposure might be transduced to other lung cells through exosomes derived from human bronchial epithelial (HBE) cells. In the present study, HBE cells were treated with PM2.5 originating from traffic. The exosomes were isolated, and their roles in the malignant transformation of lung cancer were investigated.
-
HBE cells (human bronchial epithelial cells), and A549 and H1975 cells (human lung adenocarcinoma cells) were obtained from the Shanghai Institute for Biological Sciences, Chinese Academy of Sciences (Shanghai, China). A549-luciferase (A549-luc) cells were purchased from Shanghai Zhong Qiao Xin Zhou Biotechnology (Shanghai, China). HBE cells were cultured in MEM (BasalMedia Technologies, China) supplemented with 10% fetal bovine serum (FBS, Sigma-Aldrich, USA). A549, A549-luc and H1975 cells were cultured in DMEM (BasalMedia Technologies, China) supplemented with 10% FBS (Sigma-Aldrich, USA). Cells were grown at 37 °C under 5% CO2. SP600125 (Sigma-Aldrich, USA) was dissolved in DMSO (Sigma-Aldrich, USA). Antibodies against E-cadherin (#3195), N-cadherin (#D4R1H), ZEB1 (#3396), JNK (#9252), P-JNK (#4668), ERK (#4370), P-ERK (#4695) and P38 (#8690) were purchased from Cell Signaling Technology, USA. Antibodies against GAPDH (AF1186) and P-P38 (AM063) were purchased from Beyotime Institute of Biotechnology, China. Antibodies against CD9 (EXOAB-CD9A-1) and CD81 (EXOAB-CD81A-1) were purchased from System Biosciences, USA.
-
A large-volume PM2.5 sampler (Intelligent 2031, Qingdao Laoying Inc., China) was set on top of a five-story building in the vicinity Chongqing South Road, a two-lane, two-way road with heavy traffic in Shanghai, China. Before collection, the quartz filter membrane was dried in a muffle furnace for 2 h at a temperature of 500 °C to remove any organic matter attached to the filter membrane. Sampling was performed for 3–5 days per week, 24 h per day. After the filter membrane was saturated with adsorption, it was folded in half, wrapped in aluminum foil and stored at −20 °C. During fall/winter 2018 (from 1st Nov. to 31st Dec. 2018), the main components of PM2.5 were sampled. The method of sample preparation has been described in our previous study [21]. Briefly, ultrasonic treatment of the sampling filter membranes was performed to obtain PM2.5 suspensions. Then the suspensions were stored at −80 °C. When the suspension had solidified, the samples were freeze-dried under vacuum to obtain PM2.5 particles (Labconco, USA). Finally, the PM2.5 samples were weighed and stored at −20 °C. The particles were resuspended in sterilized ultrapure water before experiments. The sampling conditions (including location, equipment and season), collection methods and extraction methods were as previously reported; therefore, the composition of PM2.5 samples was expected to be the same as that in the previous study, including several toxic heavy metals (such as lead and nickel) and a variety of polycyclic aromatic hydrocarbons [21].
-
Cells were seeded in 96-well plates (3,000 cells per well). After treatment with PM2.5 or exosomes, cell viability was assessed with Cell Counting Kit-8 assays (Beyotime Institute of Biotechnology, Shanghai, China) according to the manufacturer's protocol. The absorbance at 450 nm was measured with a microplate reader (BioTek, Winooski, VT).
-
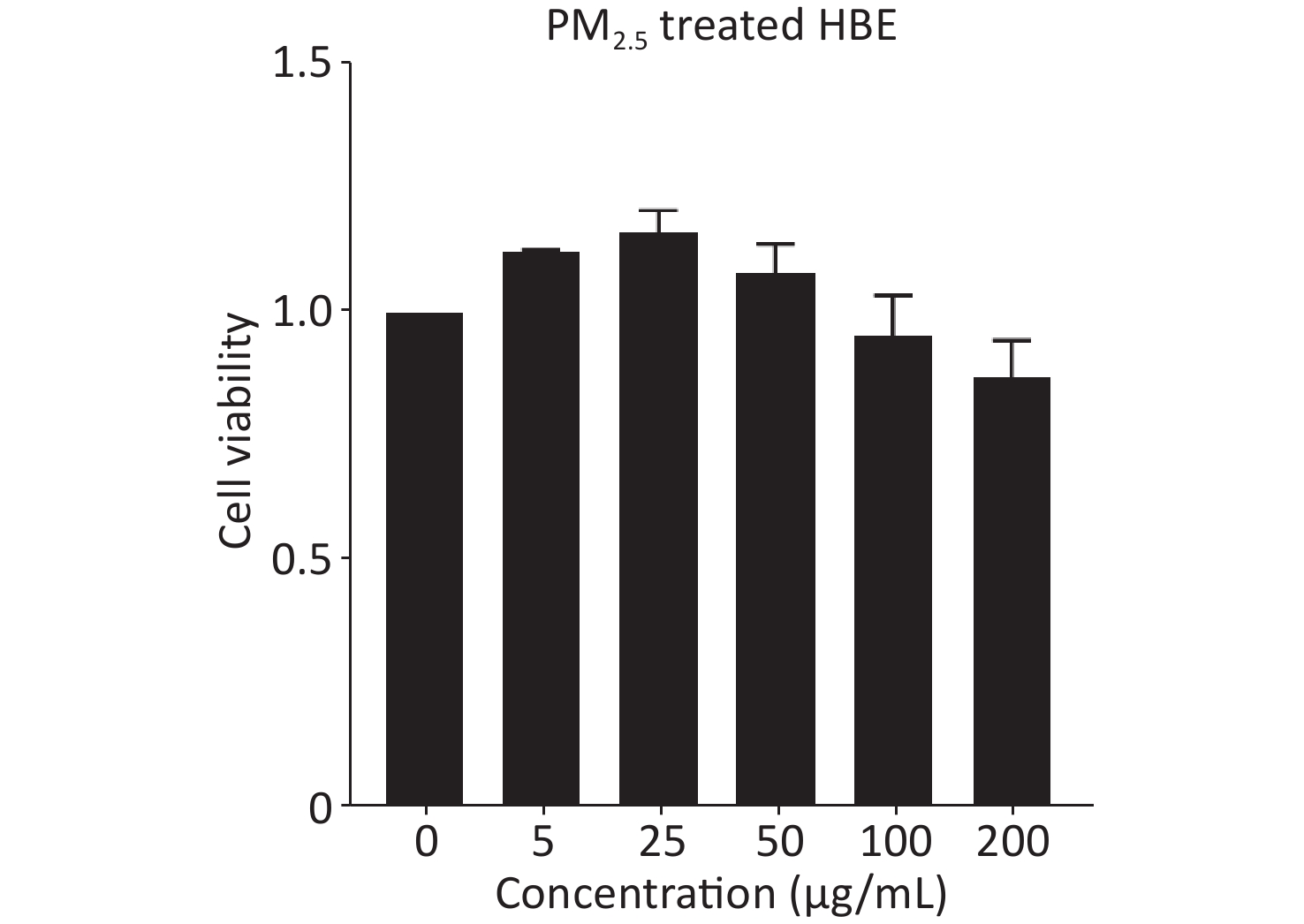
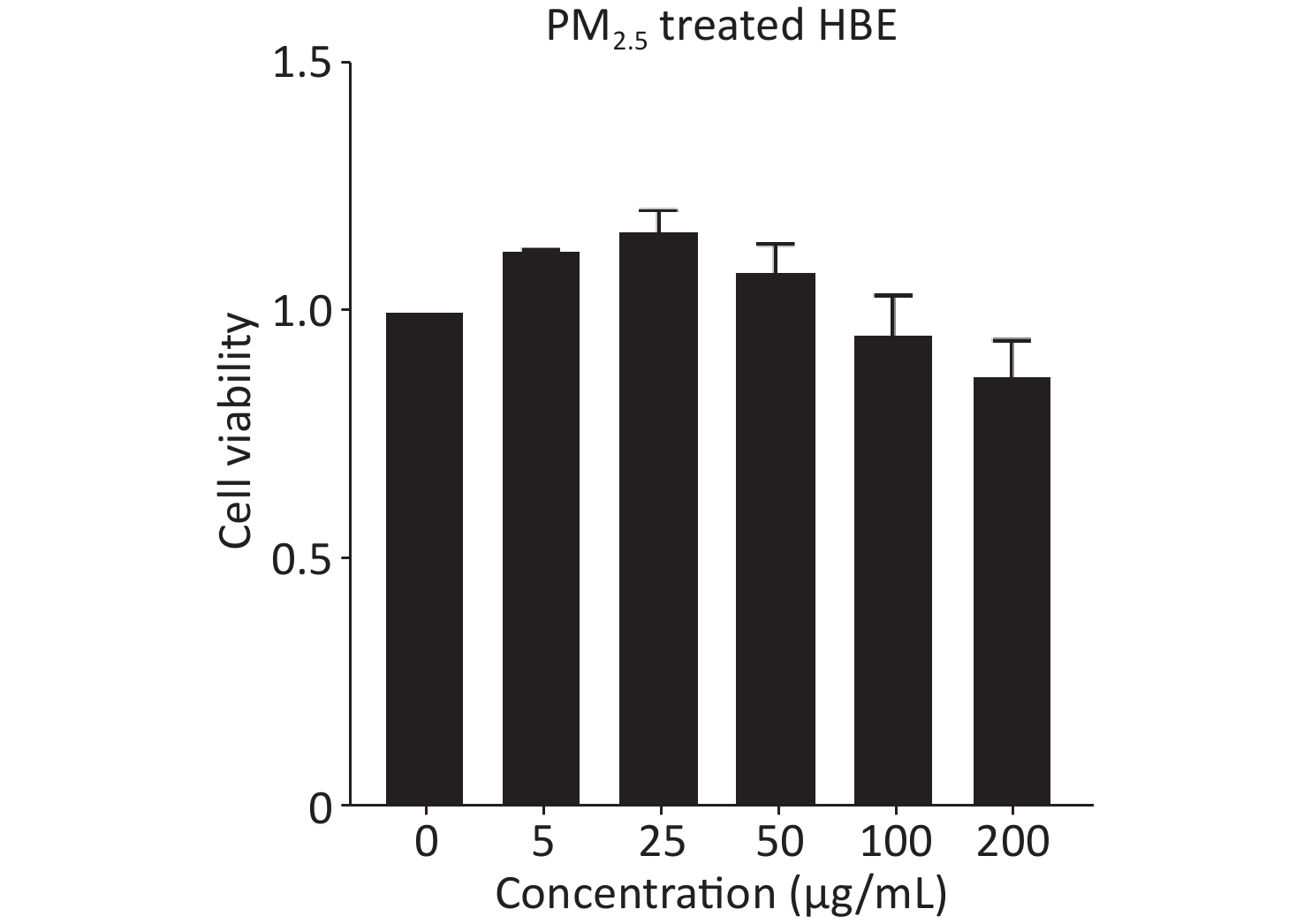
Because PM2.5 exposure in real life is long-term and chronic, we treated HBE cells with PM2.5 for 20 passages in this study. Considering the possible adverse effects of chronic PM2.5 exposure on cell viability, we used 25 μg/mL of PM2.5 as the experimental dose for treating HBE cells, which showed no overt toxic effects and was more beneficial than other dose of PM2.5 for cells (Supplementary Figure S1, available in www.besjournal.com). First, HBE cells were treated for 19 passages with PM2.5 at a dose of 25 μg/mL. Then cells were cultured in MEM with PM2.5 and 5% exosome-depleted FBS (by ultracentrifugation at 100,000 ×g at 4 °C for 12 h) for 48 h to collect exosomes. The cell supernatant was centrifuged at 3,000 ×g for 15 min and passed through a 0.22-μm filter (Millipore, USA) to remove floating cells and cellular debris; this was followed by concentration with 100 k ultrafiltration tubes (Millipore, USA). Finally, Exoquick-TC reagent (System Biosciences) was added to the concentrated solution of the supernatant according to the protocol. The mixture was incubated at 4 °C overnight and then centrifuged at 1,500 ×g for 30 min. The precipitate at the bottom, containing exosomes, was resuspended in phosphate buffered saline (PBS) and stored at −80 °C. The exosome samples were quantified with bicinchoninic acid (BCA) assays.
-
The exosomes were diluted in PBS and fixed with 4% paraformaldehyde and 4% glutaraldehyde in 0.1 mol/L phosphate buffer (pH 7.4). Then 10 μL of exosome sample was placed on a carbon-coated copper grid and negatively stained with phosphotungstic acid (pH 7.0) for 30 s. Finally, the exosomes were visualized under a transmission electron microscope (JEM-1200EX; JEOL Ltd., Tokyo, Japan). Nanoparticle tracking analysis was performed to measure the size distribution and concentration of exosomes with a ZetaView particle tracker (Particle Metrix, Munich, Germany) according to the instruction manual.
-
Proteins were extracted from cells with cell lysis buffer for western blotting containing protease inhibitors (Beyotime Institute of Biotechnology). A BCA protein assay kit (Beyotime Institute of Biotechnology) was used for quantification. The standard protocol for western blotting was as previously described[22]. The proteins (35 μg per well) were separated and transferred to polyvinylidene difluoride membranes (Millipore, USA). Primary antibodies (diluted at 1:1,000) were incubated with membranes at 4 °C overnight. After washing and incubation of the membrane with secondary antibodies (diluted at 1:1,000) for 1 h at room temperature, the protein band density was detected with a ChemiDoc MP system (Bio-Rad, USA), and quantified in Image J Software.
-
Transwell assays were used to assess the migration or invasion of A549 and H1975 cells. For migration assays, 2 × 104 A549 cells or 5 × 104 H1975 cells were suspended in 200 μL serum-free medium and seeded into the upper chambers, and 600 μL medium containing 10% FBS was added to the lower chamber. The cells were incubated at 37 °C. After 24 h, the non-migrating cells were removed with cotton swabs. The cells that migrated to the bottom of the membrane were fixed with 4% paraformaldehyde and stained with crystal violet. Migrated cells were visualized under a microscope (Olympus, Japan) and quantified by counting of the average number of cells in four random fields. The invasion assay protocol was the same as that of the migration assay except that A549 and H1975 cells were seeded into the upper chambers, which had been coated with Matrigel (Millipore, USA).
-
Seven-week-old female BALB/c nude mice were purchased from Shanghai Jihui Laboratory Animal Care Co., Ltd. (Shanghai, China). The mice were raised according to the institutional guidelines in the SPF level animal experimental center of the Ninth People's Hospital of Shanghai. The mice were divided into four groups randomly: a, control group; b, A549 group; c, A549+HBE-exo group; and d, A549+PHBE-exo group. The mice in the control group did not receive any treatment. The mice in the other three groups were injected with 5 × 106 A549-luc cells via the tail vein to establish a mouse model of lung metastasis. Additionally, the mice in the A549+HBE-exo group and A549+PHBE-exo group were injected with 200 μg exosomes derived from HBE cells or PM2.5-treated HBE cells via the tail vein twice per week. At four weeks after inoculation of A549-luc cells, the luciferase activity in the mice was evaluated through bioluminescence imaging (IVIS Lumina II; PerkinElmer, USA). The mice were then sacrificed, and the lung tissues were collected for histological analysis with hematoxylin and eosin staining. This experiment was approved by the Animal Care and Use Committee of Shanghai Jiao Tong University School of Medicine, China.
-
The statistical analyses were performed in SPSS 19.0 software (IBM, USA), and the graphs were generated in GraphPad Prism 8.0 software (Graphpad, USA). The data for cell experiments are shown as mean ± SD of three independent experiments, whereas the metastatic characteristics of experimental mice were assessed according to the median in each group. Statistics were analyzed with one-way ANOVA with the LSD post hoc test. The significance level was defined as ≤ 0.05.
-
Exosomes were isolated with Exoquick-TC reagent, and qualitative analyses were then performed. As indicated by the electron micrographs, the cup-shaped nano-vesicles had a diameter of approximately 50–150 nm (Figure 1A). Nanoparticle tracking analysis indicated no clear difference in the size distribution between exosomes derived from HBE cells (HBE-exo) and PM2.5-treated HBE cells (PHBE-exo) (Figure 1B). Furthermore, exosomal markers, including protein CD9 and CD81, were positively expressed in exosome samples (Figure 1C). Collectively, these results demonstrated that the exosomes were extracted successfully.
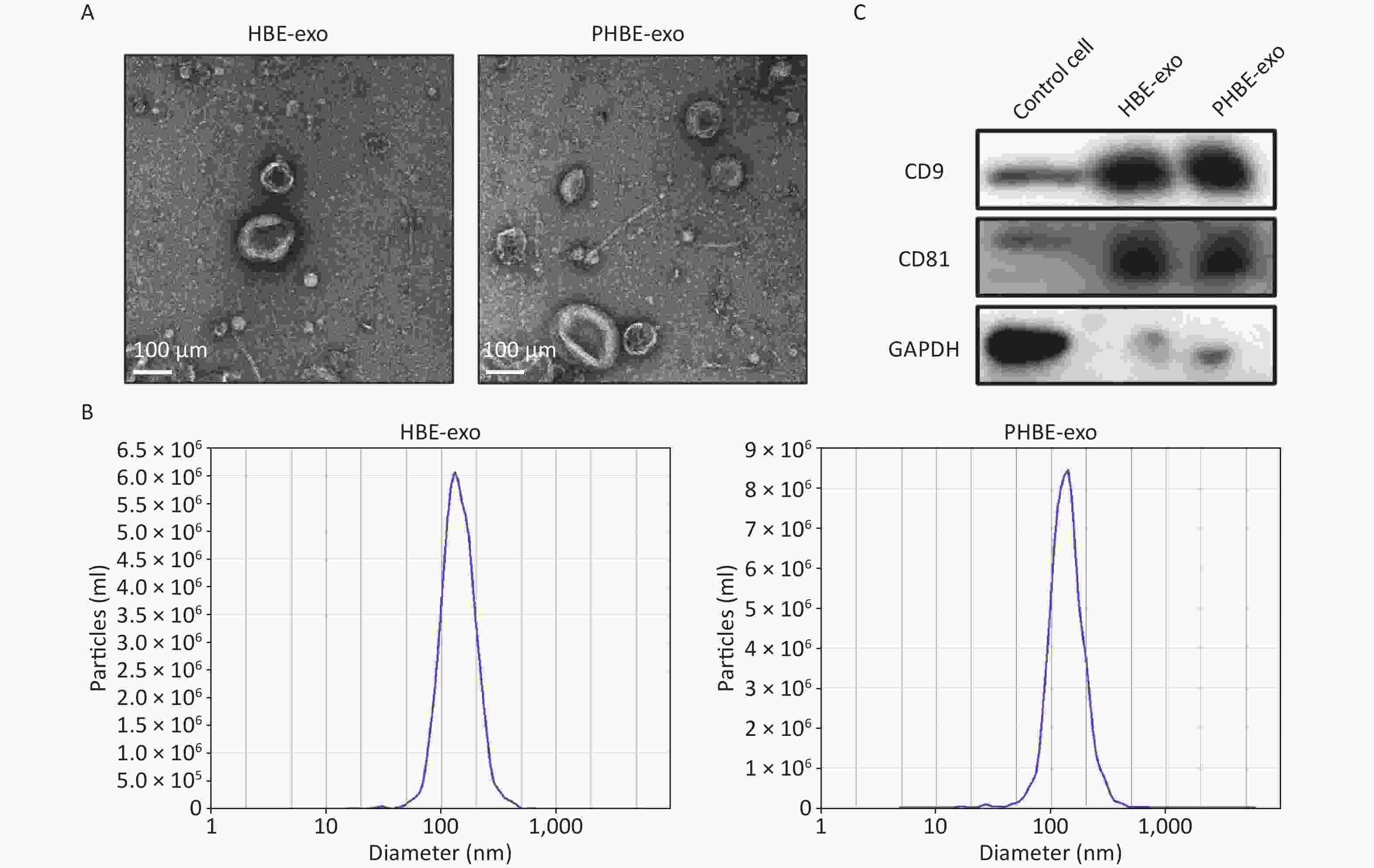
Figure 1. Identification of exosomes derived from HBE cells (HBE-exo) and exosomes derived from PM2.5-treated HBE cells (PHBE-exo). (A) Transmission electron microscopy (TEM) pictures of HBE-exo and PHBE-exo. Scale bar: 100 μm. (B) Nanoparticle tracking analysis (NTA) tested the size distribution of HBE-exo and PHBE-exo. (C) Western blotting results of exosomal markers CD9 and CD81 of control cells, HBE-exo and PHBE-exo.
-
To explore the role of PHBE-exo in metastasis of lung cancer, we used an animal model of lung metastasis. Figure 2A shows the schedule of the animal experiment. The body weight of each mouse was measured before every treatment (injection of cells or exosomes). As shown in Supplementary Figure S2 (available in www.besjournal.com), no clear difference in mouse body weight was observed among the four groups. Moreover, the lung weights and organ coefficients were also similar among the four groups (Supplementary Figure S3, available in www.besjournal.com).
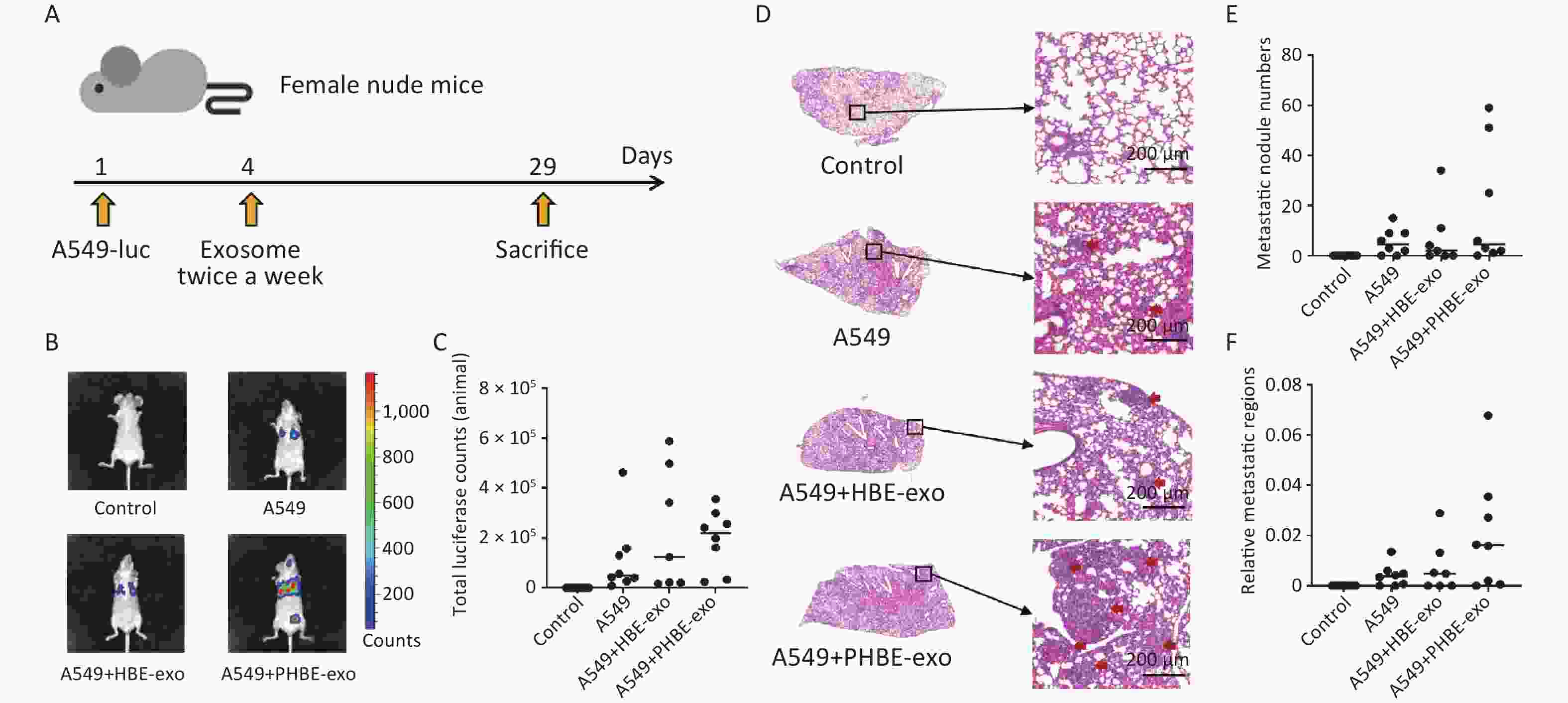
Figure 2. Exosomes derived from PM2.5-treated HBE cells promoted metastasis of lung cancer. (A) The procedure of animal experiment. (B&C) The representative images of the mice in each group (B) and total luciferase counts of each mouse (C) were assessed by biofluorescence image system. (D–F) Hematoxylin and Eosin (H&E) staining analysis of lung tissues. The representative histopathological pictures of each mice group (D): left panel, magnification ×10; right panel, magnification ×100. Red arrows: metastatic nodules. Scale bars: 200 μm. Metastatic nodule numbers (E) and relative metastatic regions (tumor region / tissue region) (F) of each mouse were calculated and shown as scatter graph with median, n ≥ 7.
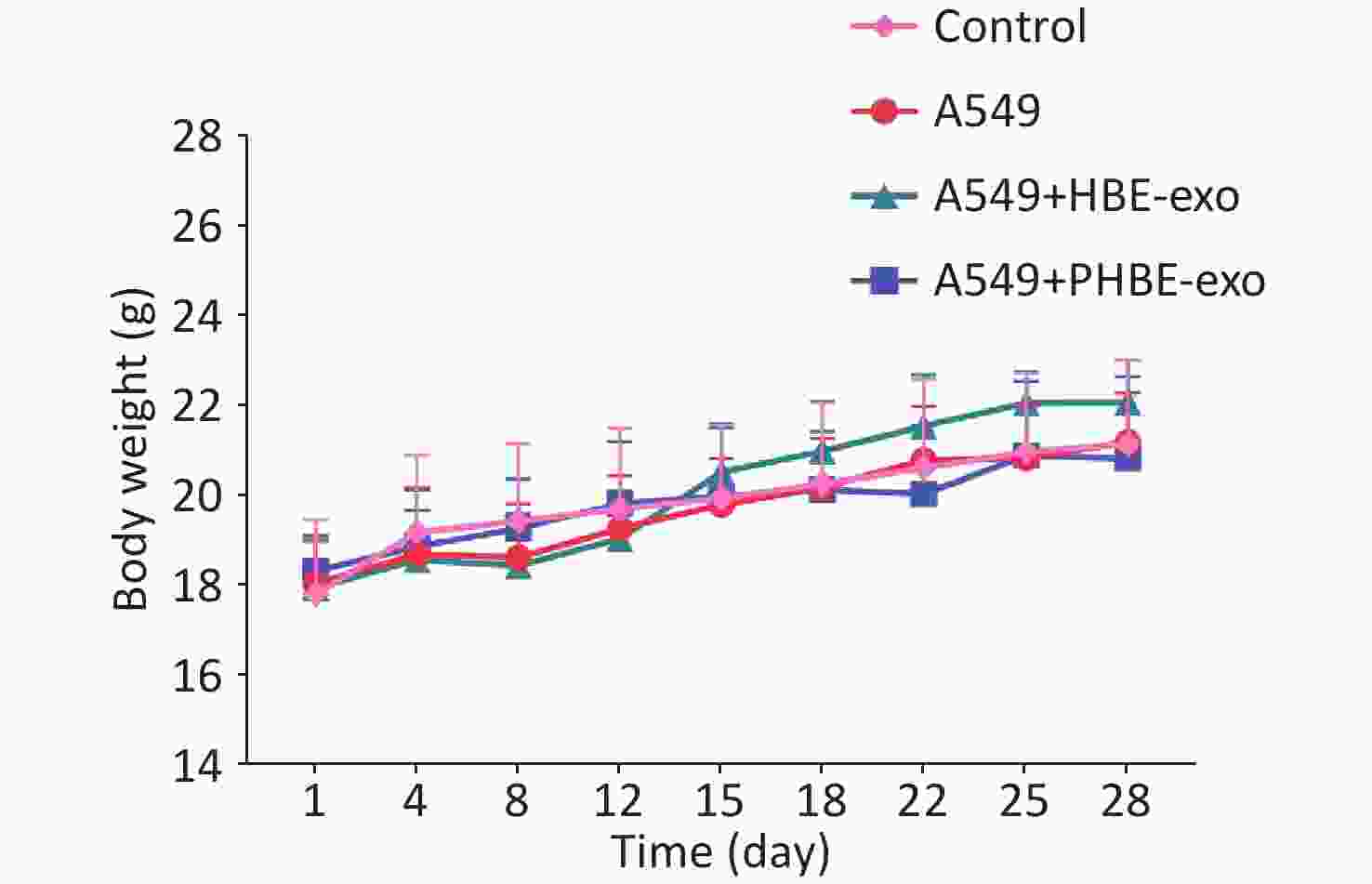
Figure S2. The body weight of mice.
Female BALB/c nude mice were divided in four groups randomly. Body weight of each mouse was measured before each injection (mean ± SD, n ≥ 7). 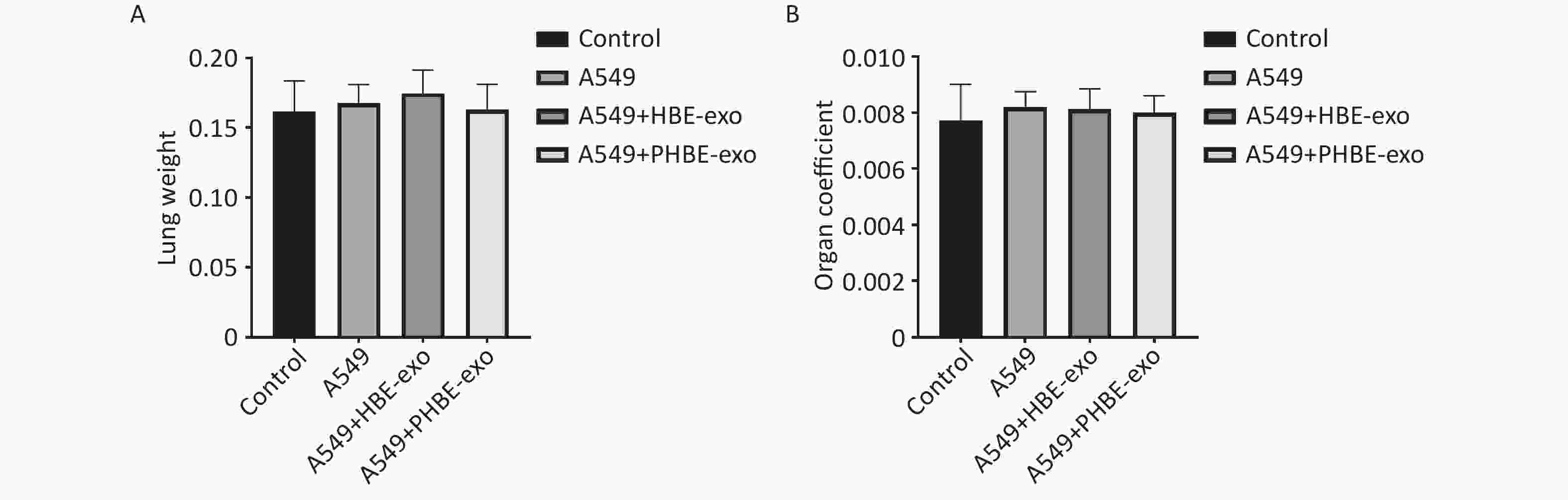
Figure S3. The lung weight and organ coefficient of mice.
Female BALB/c nude were divided in four groups randomly. Body weight were measured before the mice were sacrificed and the lung weight of each mouse was measured immediately after the anatomy (mean ± SD, n ≥ 7). The characteristics of lung metastasis were assessed with two methods: in vivo imaging and histopathological analysis. Considering the individual differences in mice, we assessed the metastatic characteristics of experimental mice according to the median in each group. As shown in Figure 2, higher luciferase activity was observed in mice in the A549+PHBE-exo group (total luciferase: 2.20 × 105) than the A549 group (total luciferase: 0.50 × 105) and A549+HBE-exo group (total luciferase: 1.24 × 105) (Figure 2B and 2C). Histopathological analysis showed no clear difference in metastatic nodule numbers among the mice with lung cancer (A549 group: 4.5; A549+HBE-exo group: 2; A549+PHBE-exo group: 4.5) (Figure 2D and 2E). Notably, more large metastatic nodules were observed in the lungs of mice treated with PHBE-exo than in the mice in the other groups. The calculated ratio of tumor to lung tissue (relative metastatic region) was higher in the A549 + PHBE-exo group than the other groups (A549 group: 0.004; A549+HBE-exo group: 0.005; A549+PHBE-exo group: 0.016) (Figure 2F). These results implied that PHBE-exo might promote metastasis of lung cancer in vivo.
-
To assess the mechanisms through which PHBE-exo promoted the lung metastatic potential, we treated A549 and H1975 cells with HBE-exo or PHBE-exo at a dose of 200 μg/mL, at which no clear change in cell viability was observed (Supplementary Figure S4, available in www.besjournal.com).
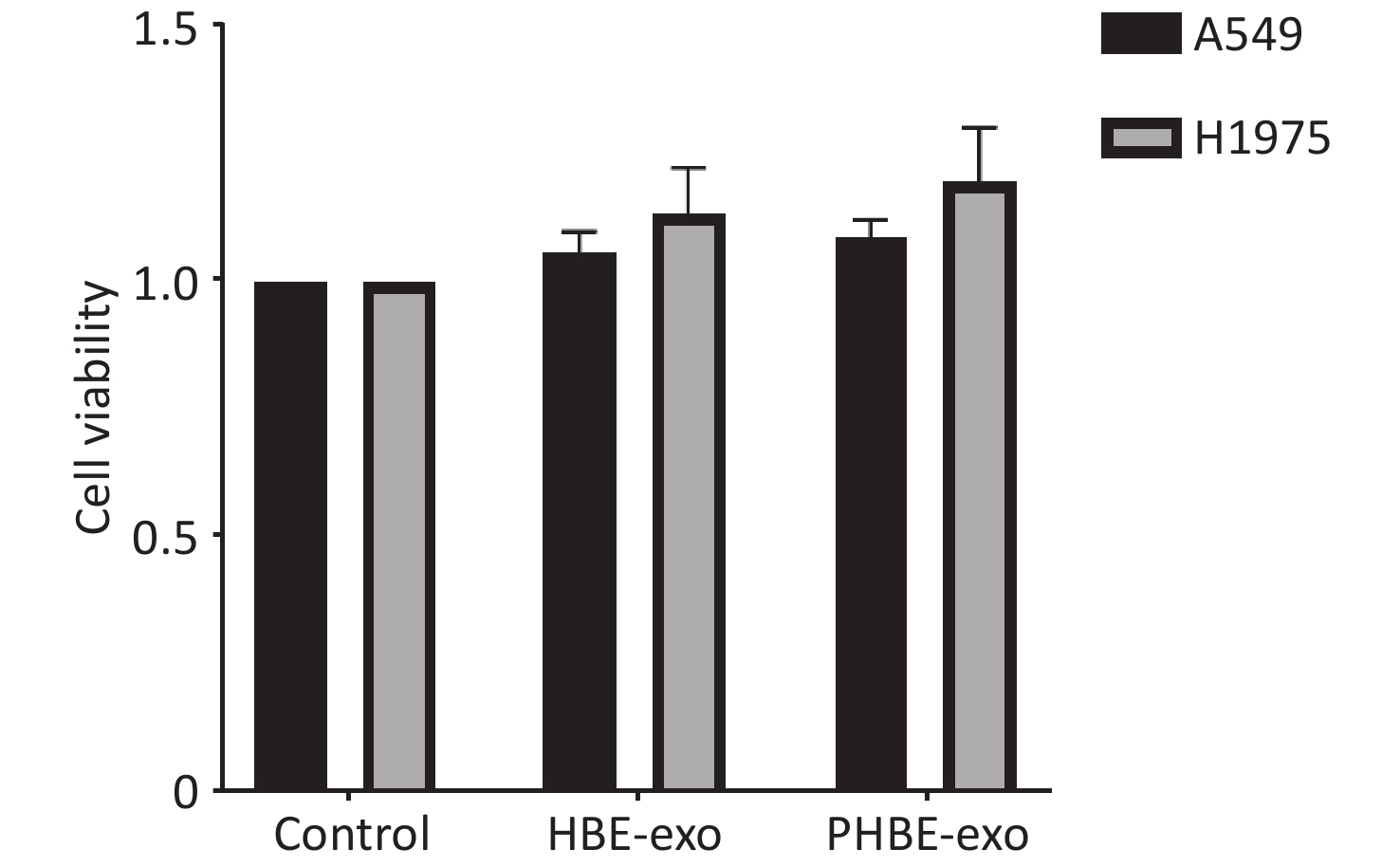
Figure S4. The effects of exosomes derived from PM2.5-treated HBE cells on the viability of lung cancer cells. Lung cancer cells were treated with PBS, exosomes derived from human bronchial epithelial cell (HBE-exo) and exosomes derived from human bronchial epithelial cells treated with PM2.5 (PHBE-exo). After 24 h, cell viabilities of lung cancer cells were assessed by cck8 kit (mean ± SD, n = 3).
Exosomes Derived from PM2.5-treated HBE Cells Promote the Migration and Invasion of A549 and H1975 Cells Through Transwell assays, we evaluated cell migration and invasion to determine the effects of PHBE-exo on lung cancer cell lines. As shown in Figure 3, more A549 and H1975 cells migrated to the bottom of the membrane in the PHBE-exo group than the PBS or HBE-exo group (P < 0.05). Moreover, the effects of exosomes on the invasive ability of A549 and H1975 cells were examined. More cells treated with PHBE-exo passed through the Matrigel (P < 0.05) (Figure 3), thus indicating that PHBE-exo enhanced the invasive potential of A549 and H1975 cells.
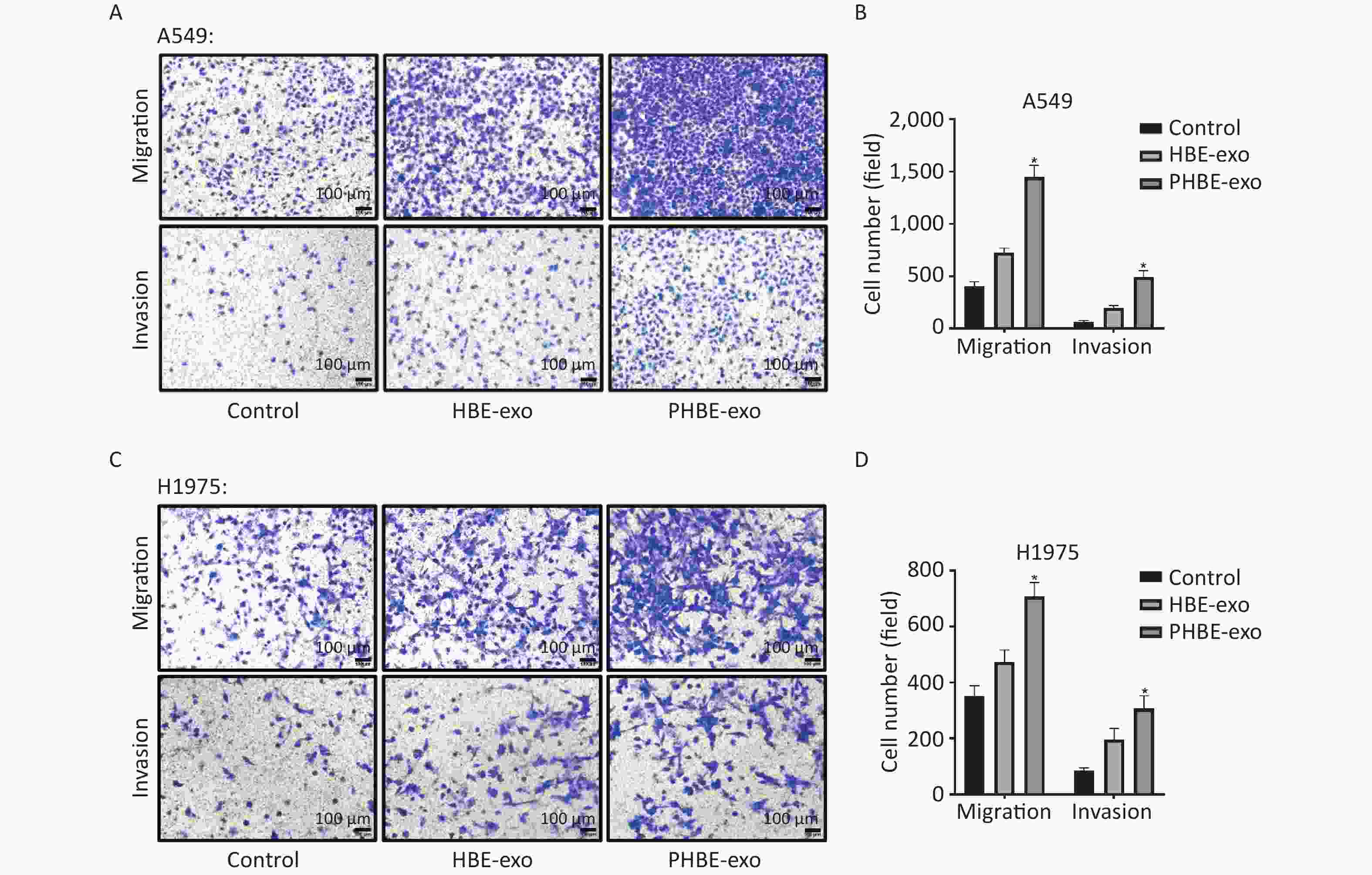
Figure 3. Exosomes derived from PM2.5-treated HBE cells promoted the migration and invasion capacity of lung cancer cells. Lung cancer cells were treated with PBS, exosomes derived from HBE cells (HBE-exo) or exosomes derived from PM2.5-treated HBE cells (PHBE-exo). Representative figures of A549 (A) and H1975 (C) cells in transwell assays. Scale bar: 100 μm. A549 (B) and H1975 (D) cell numbers were quantified by image J (mean ± SD, n = 3). *P < 0.05, different from HBE-exo-treated group.
Exosomes Derived from PM2.5-treated HBE Cells Regulate the Protein Expression of EMT Markers in A549 and H1975 Cells EMT is considered an important process in tumor metastasis [23]. In the present study, western blotting results indicated that (Figure 4) the protein levels of the epithelial marker E-cadherin were downregulated (P < 0.05), whereas the protein levels of the mesenchymal markers N-cadherin and ZEB1 were up-regulated (P < 0.05) after treatment with PHBE-exo, compared with PBS or HBE-exo, thus indicating that EMT was induced by PHBE-exo.
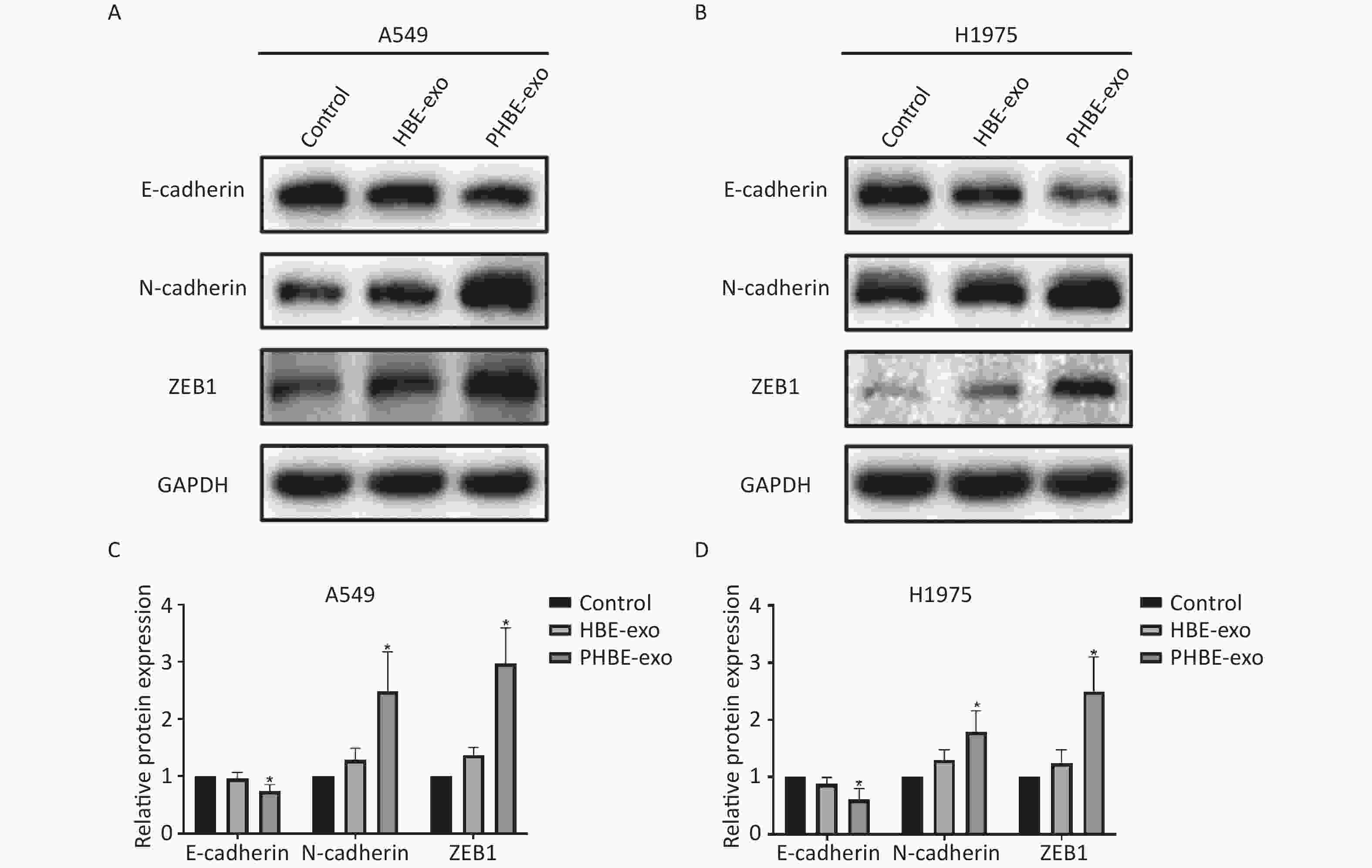
Figure 4. Exosomes derived from PM2.5-treated HBE cells changed the protein expression of epithelial-to-mesenchymal transition (EMT) markers of lung cancer cells. Lung cancer cells were treated with PBS, exosomes derived from HBE cells (HBE-exo) or exosomes derived from PM2.5-treated HBE cells (PHBE-exo). Protein expression of EMT markers in A549 (A&C) and H1975 (B&D) cells were quantified by western blotting (mean ± SD, n = 3). The intensities of protein bands were analyzed and normalized to GAPDH. *P < 0.05, different from HBE-exo-treated group.
Exosomes Derived from PM2.5-treated HBE Cells Affect the Protein Levels of Phosphorylated c-Jun N-terminal Kinase (P-JNK) Mitogen-activated protein kinase (MAPK) signaling pathways regulate a variety of cellular functions including gene transcription and protein biosynthesis [24]. To determine whether PHBE-exo might affect MAPK activation in lung cancer cell lines, we assessed the JNK, extracellular regulated protein kinase (ERK) and P38 mitogen-activated protein kinase (P38) activity. As shown in Figure 5, treatment with PHBE-exo significantly increased the protein level of P-JNK (P < 0.05) but not those of P-ERK (P > 0.05) and P-P38 (P > 0.05) in A549 cells. The increase in the protein level of P-JNK was also observed in H1975 cells (Supplementary Figure S5, available in www.besjournal.com). These results indicated that PHBE-exo activated JNK in lung cancer cells.
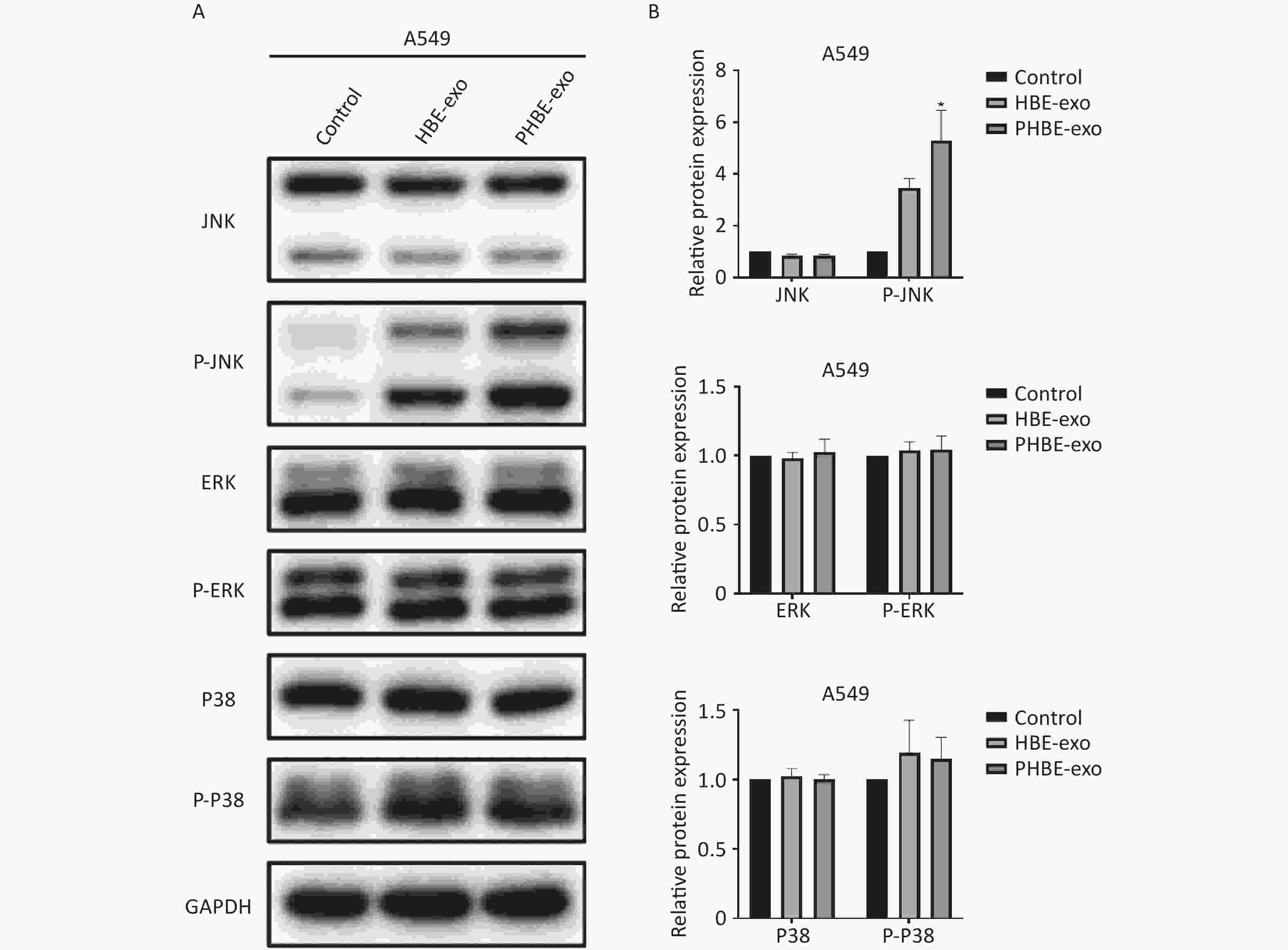
Figure 5. Exosomes derived from PM2.5-treated HBE cells affected the activation of MAPKs. A549 cells were treated with PBS, exosomes derived from HBE cells (HBE-exo) or exosomes derived from PM2.5-treated HBE cells (PHBE-exo). Representative figures of western blotting (A) and the protein expression of JNK, P-JNK, ERK, P-ERK, P-38, and P-P38 were quantified by image J (B) (mean ± SD, n = 3). The intensities of protein bands were analyzed and normalized to GAPDH. *P < 0.05, different from HBE-exo treated group.
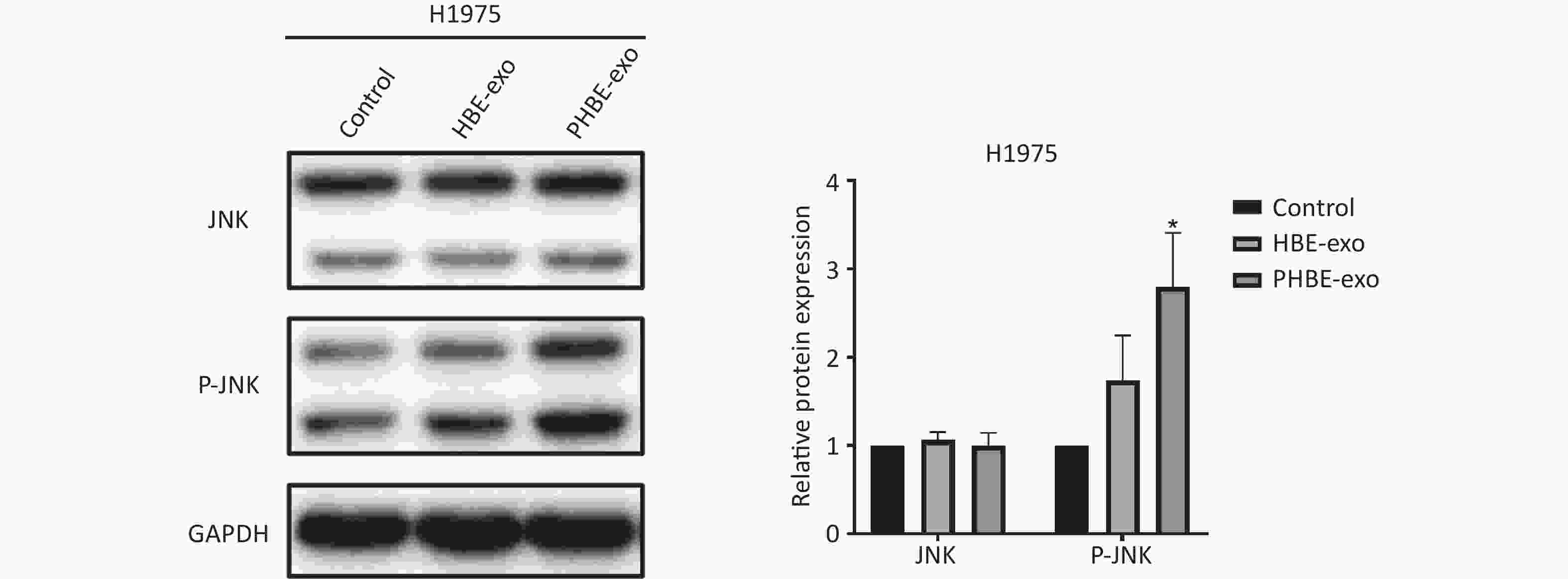
Figure S5. Exosomes derived from PM2.5-treated HBE cells affected JNK signaling pathway in H1975 cells. Lung cancer cells H1975 were treated with PBS, exosomes derived from human bronchial epithelial cell (HBE-exo) and exosomes derived from human bronchial epithelial cell treated with PM2.5 (PHBE-exo). Protein expression of JNK and phosphorylated JNK in H1975 cells were quantified by western blotting (mean ± SD, n = 3). *P < 0.05, different from HBE-exo treated group.
-
We further explored the role of JNK in promoting lung cancer induced by PHBE-exo. A549 and H1975 cells were treated with DMSO or 20 μmol/L SP600125, a selective inhibitor of JNK phosphorylation, then treated with PHBE-exo. As shown in Supplementary Figure S6 ( available in www.besjournal.com), the increase in protein levels of P-JNK mediated by PHBE-exo was effectively inhibited by SP600125 (P < 0.05).
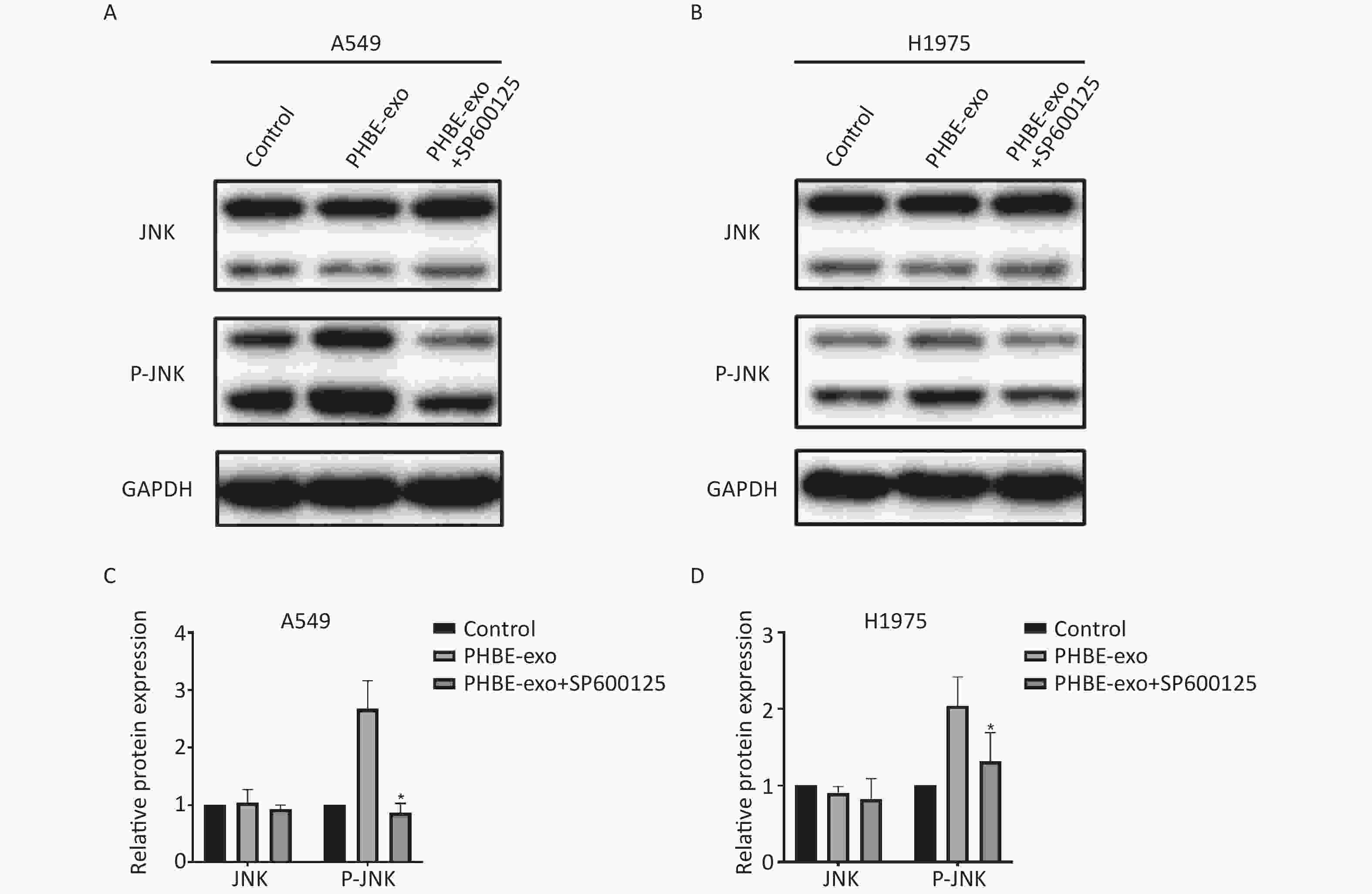
Figure S6. The JNK pathway activated by exosomes was effectively inhibited by SP600125. Lung cancer cells were treated with DMSO, exosomes derived from human bronchial epithelial cell treated with PM2.5 (PHBE-exo), PHBE-exo+SP600125. Protein expression of JNK and phosphorylated JNK in A549 (A&C) and H1975 (B&D) were quantified by western blotting (mean ± SD, n = 3). *P < 0.05, different from PHBE-exo treated group.
Enhancement of Migration and Invasion by Exosomes Derived from PM2.5-treated HBE Cells is Suppressed by SP600125 To elucidate whether PHBE-exo might promote lung cancer migration and invasion through activating JNK, we exposed A549 and H1975 cells with or without SP600125 treatment to exosomes. As shown in Figure 6, SP600125 diminished the cell migration and invasion promoted by PHBE-exo (P < 0.05), thus indicating that JNK plays a key role in these processes.
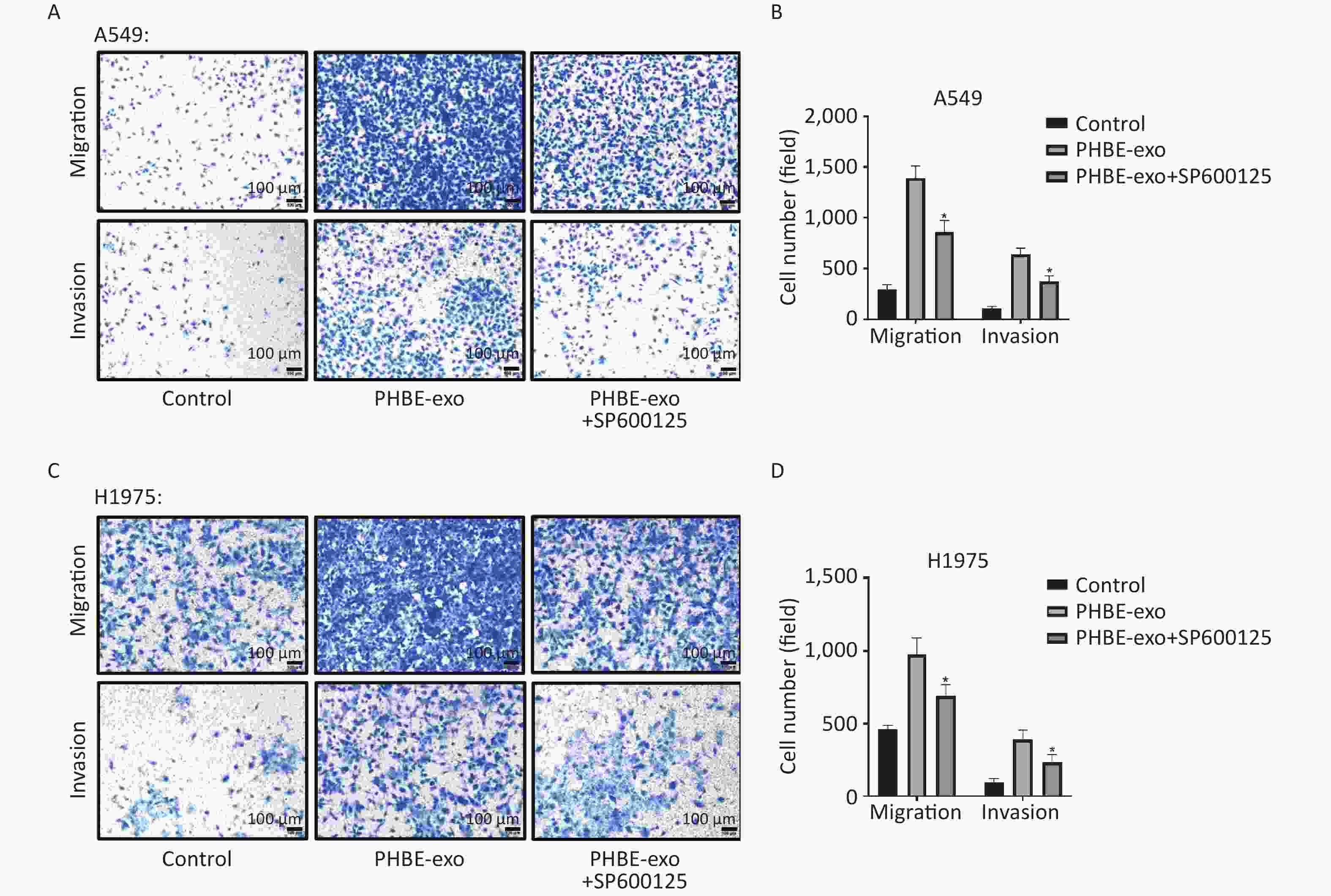
Figure 6. Inhibition of the JNK signaling pathway suppressed the migration and invasion capacity enhanced by exosomes derived from PM2.5-treated HBE cells. Lung cancer cells were treated with DMSO, exosomes derived from PM2.5-treated HBE cells (PHBE-exo), or SP600125+PHBE-exo. Representative figures of A549 (A) and H1975 (C) cells in transwell assays. Scale bar: 100 μm. Cell numbers of A549 (B) and H1975 (D) were quantified by image J (mean ± SD, n = 3). *P < 0.05, different from PHBE-exo-treated group.
EMT Induced by Exosomes Derived from PM2.5-treated HBE Cells Is Suppressed by SP600125 Next, we investigated the effect of JNK on PHBE-exo-induced EMT. Western blotting analysis indicated that the levels of EMT-associated proteins in PHBE-exo treated A549 cells were modulated by SP600125. As shown in Figure 7, the protein levels of the epithelial marker E-cadherin were up-regulated (P < 0.05) and those of the mesenchymal marker N-cadherin were down-regulated (P < 0.05) in cells initially treated with SP600125, compared with A549 cells cultured with only PHBE-exo. The same results were observed in H1975 cells (Figure 7). These results indicated that inhibition of JNK restrained the PHBE-exo-induced EMT.
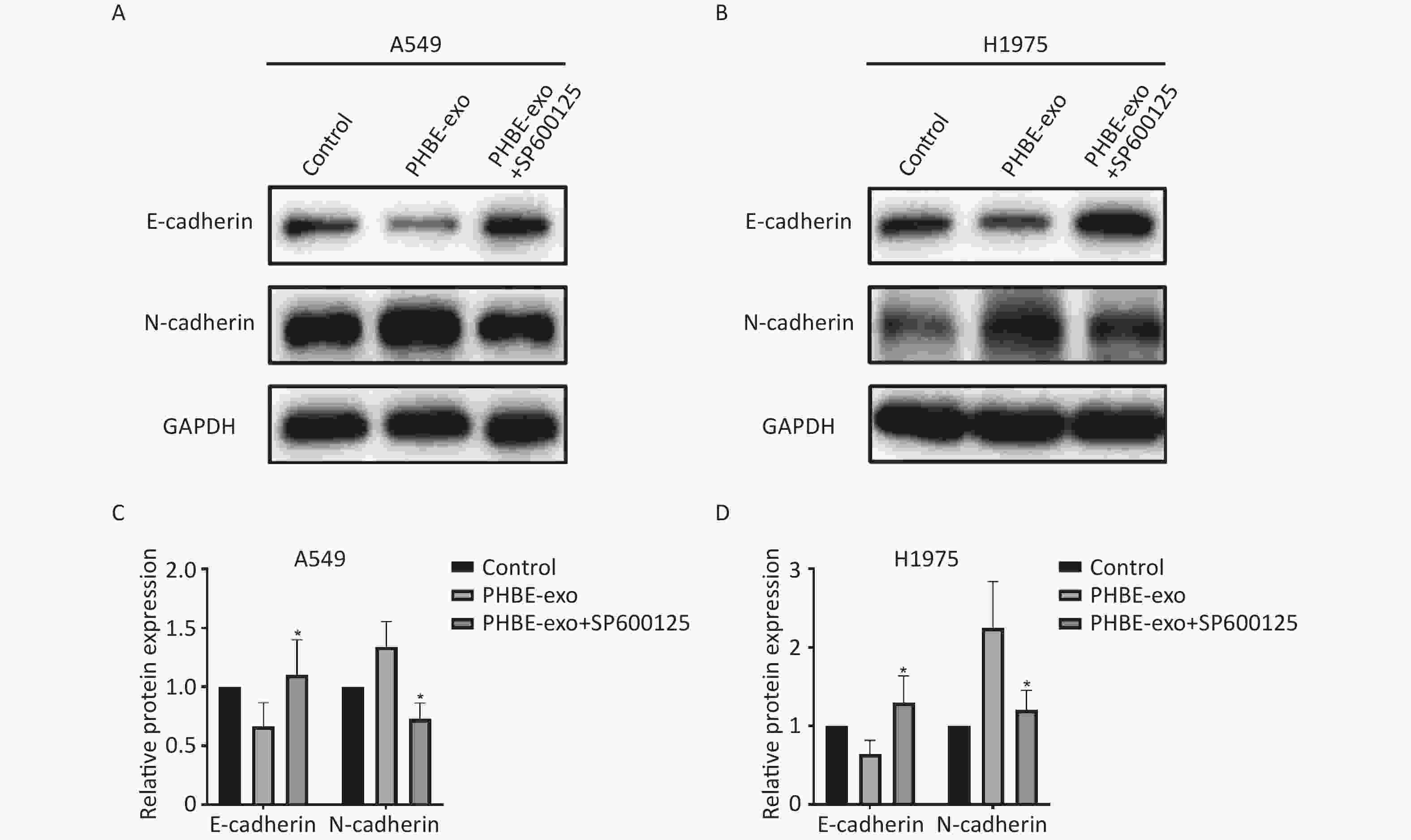
Figure 7. Inhibition of the JNK signaling pathway suppressed EMT induced by exosomes derived from PM2.5-treated HBE. Lung cancer cells were treated with DMSO, exosomes derived from PM2.5-treated HBE cells (PHBE-exo), PHBE-exo+SP600125. A549 (A&C) and H1975 (B&D) cell protein expression of EMT markers were quantified by western blotting (mean ± SD, n = 3). The intensities of protein bands were analyzed and normalized to GAPDH. *P < 0.05, different from PHBE-exo-treated group.
-
In the past century, the harm of PM2.5 has been widely reported [25]. Hundreds of millions of people live in environments with excessive PM2.5 [26]. Numerous studies have revealed that PM2.5 contributes to lung cancer [10, 27, 28]. To date, the lung cancer promotion mechanism of PM2.5 has not yet been fully elucidated. In the current study, the animal experiment results indicated that exosomes derived from PM2.5-treated human bronchial epithelial cells (PHBE-exo) might promote lung cancer metastasis in vivo. Furthermore, PHBE-exo promoted the migration, invasion and EMT of lung cancer cells, but these effects were partially suppressed by inhibition of JNK activation. These findings broadened understanding of the cancer-promoting mechanisms of environmental pollutants.
Although the functions of exosomes have been extensively studied, the roles of exosomes secreted by cells exposed to environmental pollutants remain to be investigated. Bronchial epithelial cells are a major source of exosomes in the lungs [29]; consequently, bronchial epithelial cell-derived exosomes may be involved in the pathogenesis of a wide range of lung diseases. Some studies have concluded that exosomes derived from HBE cells (HBE-exo) played important roles in several lung diseases including tumor and inflammatory diseases. Xu et al. have reported that, through intercellular communication, exosomal miRNAs derived from arsenite-transformed HBE cells participated in carcinogenesis induced by environmental chemicals [30]. Exosomal miRNA-21 derived from cigarette smoke extract treated HBE cells has been reported to be involved in cigarette smoking induced chronic obstructive pulmonary disease, pulmonary fibrosis and carcinogenesis [31-33]. Environmental pollutants can induce or promote lung diseases via exosomes from HBE cells. Therefore, we wondered whether exosomes from HBE cells might be involved in PM2.5-mediated promotion of lung cancer.
Using a lung cancer animal model, we observed that the exosomes from PM2.5-treated HBE cells increased the luciferase activity and the relative metastatic region in the lungs of mice. These results indicated that PHBE-exo promote lung metastasis. Xu et al. have reported that exosomes derived from PM2.5-treated A549 cells increase proliferation of A549 cells in vitro and increase tumor growth in vivo, thus demonstrating that exosomes derived from PM2.5 treated cells might be involved in lung cancer development [34]. Several studies have reported that, after PM2.5 exposure, the composition of exosomes derived from lung cells is altered [35, 36]: on the basis of bioinformatics analysis of exosomal miRNAs from A549 and BEAS-2B cells, the authors have speculated that exosomes might be involved in the tumorigenesis and development of PM2.5-associated cancer. In the present study, exosomes from bronchial epithelial cells (HBE-exo) also promoted lung cancer cell migration and invasion, and exosomes from PM2.5-treated cells (PHBE-exo) enhanced these effects (Figure 3). In addition, EMT was induced by PHBE-exo. Therefore, we propose that PM2.5 might promote lung cancer metastasis through exosomes derived from human bronchial epithelial cells.
MAPK signaling pathways are widely known to mediate cellular responses to various stimuli. Activation of MAPK signaling pathways often occurs in PM2.5-treated cells, and MAPK signaling pathways are associated with stress and inflammation [37-39]. In the present study, we observed the upregulation of P-JNK protein expression induced by PHBE-exo. Xu et al. have also reported that exosomal miRNAs of PM2.5-treated BEAS-2B cells might target MAPK signaling pathways [35].
JNK is a member of the MAPK family that has been reported to be involved in carcinogenesis in many studies. Exosomes can accelerate cancer development, and the JNK signaling pathway is involved in this process [40-42]. Wagner et al. have reported hyperactivation of JNK proteins in biological samples from patients with cancer, thus indicating that the JNK signaling pathway is associated with cancers in humans [43]. JNK plays multiple roles in many types of cancer progression including proliferation, migration and invasion[44-47]. Moreover, the JNK signaling pathway is involved in cancer cell growth, tumor formation and metastasis in the lungs [24, 48]. Furthermore, the EMT and cell migration ability of mouse transformed keratinocytes caused by TGF-β1 rely on activation of the JNK signaling pathway [49]. In the present study, we further explored the effects of JNK in promoting lung cancer induced by PHBE-exo. The results showed that exosomes from PM2.5-treated HBE cells promoted migration, invasion and EMT of lung cancer cells, which were accompanied by activation of JNK in the cells. SP600125, a specific inhibitor of JNK phosphorylation, diminished the migration, invasion and EMT induced by PHBE-exo. Thus, we speculate that PHBE-exo might facilitate the malignant transformation of lung cancer cells, partially through JNK.
-
All authors declare that they have no competing interests.
-
YU Heng Yi: conceptualization, methodology, formal analysis, investigation, writing the original draft. GUO Hua Qi: conceptualization, ethodology, investigation. FENG Yan: methodology, resources. CHENG Wei: resources. Wang Yan: supervision, funding acquisition, writing the review & editing.
doi: 10.3967/bes2022.066
Exosomes from PM2.5-treated Human Bronchial Epithelial Cells Increase Lung Cancer Metastatic Potential
-
Abstract:
Objective Fine particulate matter (PM2.5) is an air pollutant that has become of great concern in recent years. Numerous studies have found that PM2.5 may contribute to lung cancer, but the pathogenesis has not yet been fully elucidated. In this study, we explored the roles of exosomes from bronchial epithelial cells in PM2.5-promoted lung cancer metastasis. Methods Exosomes were isolated from cell supernatants. An animal model of lung metastasis (established by tail vein injection of A549-luc) and in vitro studies with lung cancer cell lines were used to investigate the effects of exosomes derived from PM2.5-treated human bronchial epithelial cells (PHBE-exo). Results The animal experiments revealed that PHBE-exo-treated mice showed stronger luciferase activity and a larger relative metastatic region in the lungs, thus indicating that PHBE-exo promoted the metastatic potential of lung cancer. Additionally, PHBE-exo promoted the migration, invasion and epithelial-to-mesenchymal transition of lung cancer cells, in a manner mediated by activation of c-Jun N-terminal kinase. Conclusion These results implied that PM2.5 may promote the development of lung cancer through exosomes derived from bronchial epithelial cells, thus providing a potential interventional target for lung cancer. These findings broadened our understanding of cancer-promoting mechanisms of environmental pollutants. -
Key words:
- Fine particulate matter /
- Exosome /
- Lung cancer /
- C-Jun N-terminal kinase /
- Bronchial epithelial cell
-
Figure 1. Identification of exosomes derived from HBE cells (HBE-exo) and exosomes derived from PM2.5-treated HBE cells (PHBE-exo). (A) Transmission electron microscopy (TEM) pictures of HBE-exo and PHBE-exo. Scale bar: 100 μm. (B) Nanoparticle tracking analysis (NTA) tested the size distribution of HBE-exo and PHBE-exo. (C) Western blotting results of exosomal markers CD9 and CD81 of control cells, HBE-exo and PHBE-exo.
Figure 2. Exosomes derived from PM2.5-treated HBE cells promoted metastasis of lung cancer. (A) The procedure of animal experiment. (B&C) The representative images of the mice in each group (B) and total luciferase counts of each mouse (C) were assessed by biofluorescence image system. (D–F) Hematoxylin and Eosin (H&E) staining analysis of lung tissues. The representative histopathological pictures of each mice group (D): left panel, magnification ×10; right panel, magnification ×100. Red arrows: metastatic nodules. Scale bars: 200 μm. Metastatic nodule numbers (E) and relative metastatic regions (tumor region / tissue region) (F) of each mouse were calculated and shown as scatter graph with median, n ≥ 7.
S4. The effects of exosomes derived from PM2.5-treated HBE cells on the viability of lung cancer cells. Lung cancer cells were treated with PBS, exosomes derived from human bronchial epithelial cell (HBE-exo) and exosomes derived from human bronchial epithelial cells treated with PM2.5 (PHBE-exo). After 24 h, cell viabilities of lung cancer cells were assessed by cck8 kit (mean ± SD, n = 3).
Figure 3. Exosomes derived from PM2.5-treated HBE cells promoted the migration and invasion capacity of lung cancer cells. Lung cancer cells were treated with PBS, exosomes derived from HBE cells (HBE-exo) or exosomes derived from PM2.5-treated HBE cells (PHBE-exo). Representative figures of A549 (A) and H1975 (C) cells in transwell assays. Scale bar: 100 μm. A549 (B) and H1975 (D) cell numbers were quantified by image J (mean ± SD, n = 3). *P < 0.05, different from HBE-exo-treated group.
Figure 4. Exosomes derived from PM2.5-treated HBE cells changed the protein expression of epithelial-to-mesenchymal transition (EMT) markers of lung cancer cells. Lung cancer cells were treated with PBS, exosomes derived from HBE cells (HBE-exo) or exosomes derived from PM2.5-treated HBE cells (PHBE-exo). Protein expression of EMT markers in A549 (A&C) and H1975 (B&D) cells were quantified by western blotting (mean ± SD, n = 3). The intensities of protein bands were analyzed and normalized to GAPDH. *P < 0.05, different from HBE-exo-treated group.
Figure 5. Exosomes derived from PM2.5-treated HBE cells affected the activation of MAPKs. A549 cells were treated with PBS, exosomes derived from HBE cells (HBE-exo) or exosomes derived from PM2.5-treated HBE cells (PHBE-exo). Representative figures of western blotting (A) and the protein expression of JNK, P-JNK, ERK, P-ERK, P-38, and P-P38 were quantified by image J (B) (mean ± SD, n = 3). The intensities of protein bands were analyzed and normalized to GAPDH. *P < 0.05, different from HBE-exo treated group.
S5. Exosomes derived from PM2.5-treated HBE cells affected JNK signaling pathway in H1975 cells. Lung cancer cells H1975 were treated with PBS, exosomes derived from human bronchial epithelial cell (HBE-exo) and exosomes derived from human bronchial epithelial cell treated with PM2.5 (PHBE-exo). Protein expression of JNK and phosphorylated JNK in H1975 cells were quantified by western blotting (mean ± SD, n = 3). *P < 0.05, different from HBE-exo treated group.
S6. The JNK pathway activated by exosomes was effectively inhibited by SP600125. Lung cancer cells were treated with DMSO, exosomes derived from human bronchial epithelial cell treated with PM2.5 (PHBE-exo), PHBE-exo+SP600125. Protein expression of JNK and phosphorylated JNK in A549 (A&C) and H1975 (B&D) were quantified by western blotting (mean ± SD, n = 3). *P < 0.05, different from PHBE-exo treated group.
Figure 6. Inhibition of the JNK signaling pathway suppressed the migration and invasion capacity enhanced by exosomes derived from PM2.5-treated HBE cells. Lung cancer cells were treated with DMSO, exosomes derived from PM2.5-treated HBE cells (PHBE-exo), or SP600125+PHBE-exo. Representative figures of A549 (A) and H1975 (C) cells in transwell assays. Scale bar: 100 μm. Cell numbers of A549 (B) and H1975 (D) were quantified by image J (mean ± SD, n = 3). *P < 0.05, different from PHBE-exo-treated group.
Figure 7. Inhibition of the JNK signaling pathway suppressed EMT induced by exosomes derived from PM2.5-treated HBE. Lung cancer cells were treated with DMSO, exosomes derived from PM2.5-treated HBE cells (PHBE-exo), PHBE-exo+SP600125. A549 (A&C) and H1975 (B&D) cell protein expression of EMT markers were quantified by western blotting (mean ± SD, n = 3). The intensities of protein bands were analyzed and normalized to GAPDH. *P < 0.05, different from PHBE-exo-treated group.
-
[1] IARC. Outdoor air pollution. IARC. 2013, 9-444. [2] Vattanasit U, Navasumrit P, Khadka MB, et al. Oxidative DNA damage and inflammatory responses in cultured human cells and in humans exposed to traffic-related particles. Int J Hyg Environ Health, 2014; 217, 23−33. doi: 10.1016/j.ijheh.2013.03.002 [3] Harrison RM. Airborne particulate matter. Philos Trans A Math Phys Eng Sci, 2020; 378, 20190319. [4] Ghio AJ, Carraway MS, Madden MC. Composition of air pollution particles and oxidative stress in cells, tissues, and living systems. J Toxicol Environ Health B Crit Rev, 2012; 15, 1−21. doi: 10.1080/10937404.2012.632359 [5] Feng S L, Gao D, Liao F, et al. The health effects of ambient PM2.5 and potential mechanisms. Ecotoxicol Environ Saf, 2016; 128, 67−74. doi: 10.1016/j.ecoenv.2016.01.030 [6] IARC. World cancer report: cancer research for cancer prevention. 2020, 15-572. [7] Global Burden of Disease Cancer Collaboration. Global, regional, and national cancer incidence, mortality, years of life lost, years lived with disability, and disability-adjusted life-years for 29 cancer groups, 1990 to 2017: a systematic analysis for the global burden of disease study. JAMA Oncol, 2019; 5, 1749−68. doi: 10.1001/jamaoncol.2019.2996 [8] Lambert AW, Pattabiraman DR, Weinberg RA. Emerging biological principles of metastasis. Cell, 2017; 168, 670−91. doi: 10.1016/j.cell.2016.11.037 [9] Yin L, Liu XT, Shao XJ, et al. The role of exosomes in lung cancer metastasis and clinical applications: an updated review. J Transl Med, 2021; 19, 312. doi: 10.1186/s12967-021-02985-1 [10] Laden F, Schwartz J, Speizer FE, et al. Reduction in fine particulate air pollution and mortality: Extended follow-up of the Harvard Six Cities study. Am J Respir Crit Care Med, 2006; 173, 667−72. doi: 10.1164/rccm.200503-443OC [11] Chao X, Yi L, Lan LL, et al. Long-term PM2.5 exposure increases the risk of non-small cell lung cancer (NSCLC) progression by enhancing interleukin-17a (IL-17a)-regulated proliferation and metastasis. Aging, 2020; 12, 11579−602. doi: 10.18632/aging.103319 [12] Kalluri R, LeBleu VS. The biology, function, and biomedical applications of exosomes. Science, 2020; 367, eaau6977. doi: 10.1126/science.aau6977 [13] Keller S, Ridinger J, Rupp AK, et al. Body fluid derived exosomes as a novel template for clinical diagnostics. J Transl Med, 2011; 9, 86. doi: 10.1186/1479-5876-9-86 [14] Kim JE, Eom JS, Kim WY, et al. Diagnostic value of microRNAs derived from exosomes in bronchoalveolar lavage fluid of early-stage lung adenocarcinoma: a pilot study. Thorac Cancer, 2018; 9, 911−5. doi: 10.1111/1759-7714.12756 [15] Kojima M, Gimenes-Junior JA, Langness S, et al. Exosomes, not protein or lipids, in mesenteric lymph activate inflammation: unlocking the mystery of post-shock multiple organ failure. J Trauma Acute Care Surg, 2017; 82, 42−50. doi: 10.1097/TA.0000000000001296 [16] Javeed N, Mukhopadhyay D. Exosomes and their role in the micro-/macro-environment: a comprehensive review. J Biomed Res, 2017; 31, 386−94. doi: 10.7555/JBR.30.20150162 [17] Li Q, Li BW, Li Q, et al. Exosomal miR-21-5p derived from gastric cancer promotes peritoneal metastasis via mesothelial-to-mesenchymal transition. Cell Death Dis, 2018; 9, 854. doi: 10.1038/s41419-018-0928-8 [18] Lin QS, Chen XS, Meng FZ, et al. ASPH-notch axis guided exosomal delivery of prometastatic secretome renders breast cancer multi-organ metastasis. Mol Cancer, 2019; 18, 156. doi: 10.1186/s12943-019-1077-0 [19] Zhang XN, Sai BQ, Wang F, et al. Hypoxic BMSC-derived exosomal miRNAs promote metastasis of lung cancer cells via STAT3-induced EMT. Mol Cancer, 2019; 18, 40. doi: 10.1186/s12943-019-0959-5 [20] Zhang L, Yu DH. Exosomes in cancer development, metastasis, and immunity. Biochim Biophys Acta Rev Cancer, 2019; 1871, 455−68. doi: 10.1016/j.bbcan.2019.04.004 [21] Wei HY, Liang F, Meng G, et al. Redox/methylation mediated abnormal DNA methylation as regulators of ambient fine particulate matter-induced neurodevelopment related impairment in human neuronal cells. Sci Rep, 2016; 6, 33402. doi: 10.1038/srep33402 [22] Guo HQ, Feng Y, Yu HY, et al. A novel lncRNA, loc107985872, promotes lung adenocarcinoma progression via the notch1 signaling pathway with exposure to traffic-originated PM2.5 organic extract. Environ Pollut, 2020; 266, 115307. doi: 10.1016/j.envpol.2020.115307 [23] Thiery JP. Epithelial-mesenchymal transitions in tumour progression. Nat Rev Cancer, 2002; 2, 442−54. doi: 10.1038/nrc822 [24] Kyriakis JM, Avruch J. Mammalian MAPK signal transduction pathways activated by stress and inflammation: a 10-year update. Physiol Rev, 2012; 92, 689−737. doi: 10.1152/physrev.00028.2011 [25] Dockery DW, Speizer FE, Stram DO, et al. Effects of inhalable particles on respiratory health of children. Am Rev Respir Dis, 1989; 139, 587−94. doi: 10.1164/ajrccm/139.3.587 [26] Zhang LL, Wilson JP, MacDonald B, et al. The changing PM2.5 dynamics of global megacities based on long-term remotely sensed observations. Environ Int, 2020; 142, 105862. doi: 10.1016/j.envint.2020.105862 [27] Yue HF, Yun Y, Gao R, et al. Winter polycyclic aromatic hydrocarbon-bound particulate matter from peri-urban north china promotes lung cancer cell metastasis. Environ Sci Technol, 2015; 49, 14484−93. doi: 10.1021/es506280c [28] Hamra GB, Guha N, Cohen A, et al. Outdoor particulate matter exposure and lung cancer: a systematic review and meta-analysis. Environ Health Perspect, 2014; 122, 906−11. doi: 10.1289/ehp/1408092 [29] Kulshreshtha A, Ahmad T, Agrawal A, et al. Proinflammatory role of epithelial cell-derived exosomes in allergic airway inflammation. J Allergy Clin Immunol, 2013; 131, 1194−1203.e14. doi: 10.1016/j.jaci.2012.12.1565 [30] Xu Y, Luo F, Liu Y, et al. Exosomal miR-21 derived from arsenite-transformed human bronchial epithelial cells promotes cell proliferation associated with arsenite carcinogenesis. Arch Toxicol, 2015; 89, 1071−82. doi: 10.1007/s00204-014-1291-x [31] Xu H, Ling M, Xue JC, et al. Exosomal microRNA-21 derived from bronchial epithelial cells is involved in aberrant epithelium-fibroblast cross-talk in COPD induced by cigarette smoking. Theranostics, 2018; 8, 5419−33. doi: 10.7150/thno.27876 [32] Liu Y, Luo F, Wang BR, et al. STAT3-regulated exosomal miR-21 promotes angiogenesis and is involved in neoplastic processes of transformed human bronchial epithelial cells. Cancer Lett, 2016; 370, 125−35. doi: 10.1016/j.canlet.2015.10.011 [33] Bai J, Deng JJ, Han ZX, et al. CircRNA_0026344 via exosomal miR-21 regulation of Smad7 is involved in aberrant cross-talk of epithelium-fibroblasts during cigarette smoke-induced pulmonary fibrosis. Toxicol Lett, 2021; 347, 58−66. doi: 10.1016/j.toxlet.2021.04.017 [34] Xu H, Jiao XA, Wu YL, et al. Exosomes derived from PM2.5-treated lung cancer cells promote the growth of lung cancer via the Wnt3a/β-catenin pathway. Oncol Rep, 2019; 41, 1180−8. [35] Xu ZH, Wang N, Xu Y, et al. Effects of chronic PM2.5 exposure on pulmonary epithelia: transcriptome analysis of mRNA-exosomal miRNA interactions. Toxicol Lett, 2019; 316, 49−59. doi: 10.1016/j.toxlet.2019.09.010 [36] Wang YX, Zhong YJ, Sun KY, et al. Identification of exosome miRNAs in bronchial epithelial cells after PM2.5 chronic exposure. Ecotoxicol Environ Saf, 2021; 215, 112127. doi: 10.1016/j.ecoenv.2021.112127 [37] Kyriakis JM, Avruch J. Mammalian mitogen-activated protein kinase signal transduction pathways activated by stress and inflammation. Physiol Rev, 2001; 81, 807−69. doi: 10.1152/physrev.2001.81.2.807 [38] Zhao C, Wang Y, Su ZL, et al. Respiratory exposure to PM2.5 soluble extract disrupts mucosal barrier function and promotes the development of experimental asthma. Sci Total Environ, 2020; 730, 139145. doi: 10.1016/j.scitotenv.2020.139145 [39] Zhou ZX, Liu YH, Duan FK, et al. Transcriptomic analyses of the biological effects of airborne PM2.5 exposure on human bronchial epithelial cells. PLoS One, 2015; 10, e0138267. doi: 10.1371/journal.pone.0138267 [40] Shao C, Huang YY, Fu BJ, et al. Targeting c-Jun in A549 cancer cells exhibits antiangiogenic activity in vitro and in vivo through exosome/miRNA-494-3p/PTEN signal pathway. Front Oncol, 2021; 11, 663183. doi: 10.3389/fonc.2021.663183 [41] Xiao M, Zhang JJ, Chen WJ, et al. M1-like tumor-associated macrophages activated by exosome-transferred THBS1 promote malignant migration in oral squamous cell carcinoma. J Exp Clin Cancer Res, 2018; 37, 143. doi: 10.1186/s13046-018-0815-2 [42] Liao J, Liu R, Shi YJ, et al. Exosome-shuttling microRNA-21 promotes cell migration and invasion-targeting PDCD4 in esophageal cancer. Int J Oncol, 2016; 48, 2567−79. doi: 10.3892/ijo.2016.3453 [43] Wagner EF, Nebreda ÁR. Signal integration by JNK and p38 MAPK pathways in cancer development. Nat Rev Cancer, 2009; 9, 537−49. doi: 10.1038/nrc2694 [44] Xu RY, Hu JP. The role of JNK in prostate cancer progression and therapeutic strategies. Biomed Pharmacother, 2020; 121, 109679. doi: 10.1016/j.biopha.2019.109679 [45] Yang YM, Kim SY, Seki E. Inflammation and Liver Cancer: Molecular Mechanisms and Therapeutic Targets. Semin Liver Dis, 2019; 39, 26−42. doi: 10.1055/s-0038-1676806 [46] Antonyak MA, Kenyon LC, Godwin AK, et al. Elevated JNK activation contributes to the pathogenesis of human brain tumors. Oncogene, 2002; 21, 5038−46. doi: 10.1038/sj.onc.1205593 [47] Wu QH, Wu WD, Jacevic V, et al. Selective inhibitors for JNK signalling: a potential targeted therapy in cancer. J Enzyme Inhib Med Chem, 2020; 35, 574−83. doi: 10.1080/14756366.2020.1720013 [48] Zhang DY, Jiang QW, Ge XW, et al. RHOV promotes lung adenocarcinoma cell growth and metastasis through JNK/c-Jun pathway. Int J Biol Sci, 2021; 17, 2622−32. doi: 10.7150/ijbs.59939 [49] Santibañez JF. JNK mediates TGF-β1-induced epithelial mesenchymal transdifferentiation of mouse transformed keratinocytes. FEBS Lett, 2006; 580, 5385−91. doi: 10.1016/j.febslet.2006.09.003 -
 22070Supplementary Materials.pdf
22070Supplementary Materials.pdf

-




 下载:
下载:





 Quick Links
Quick Links