-
Acute promyelocytic leukemia (APL) is a subtype of acute myeloid leukemia (AML) that has helped establish key paradigms in cancer[1,2]. The vast majority of APL specimens contain recurrent t (15;17) chromosomal translocation that results in PML-RARa fusion oncogene[3]. Therapeutic advances in the use of all-trans retinoic acid and arsenic trioxide have revolutionized the treatment of APL, with the vast majority of patients being cured with this therapeutic combination[4]. Despite that treatment advances have established APL therapy as an exemplary success of modern medicine, the pathogenesis and pathophysiology of PML-RARa+ APL are incompletely understood[5]. PML-RARA is a relatively weak oncogene. Mouse models with human PML-RARA require long latency before leukemia development[6-8]. A study further showed that the physiological level of PML-RARa is incapable of initiating leukemia in mice[9]. These data argue for the need for cooperating oncogenic events for leukemogenesis. In addition, sporadic reports—often case studies—have noted that APL patients often have low white blood cell (WBC) counts in peripheral blood, which differs from many other AMLs. However, molecular insights into why many APL patients have low WBC counts are nonexistent. MicroRNAs (miRNAs) are short non-coding RNAs that play key roles in cancer. Many miRNAs have been reported to be dysregulated in AML, and some have been demonstrated to functionally contribute to leukemogenesis[10-13]. This condition raised the question of the possible role of deregulated miRNAs in APL regulating APL pathogenesis in vivo.
Previously, we demonstrated a cooperative and dependent association between miR-125b and the MLL oncogene in AML leukemogenesis and showed that the miR-125b-TET2-VEGFA pathway mediates non-cell-intrinsic leukemia-promoting effects via an oncogenic miRNA[14]. Furthermore, we observed that miR-125b is relatively highly expressed in AML with MLL and PML-RARa translocation on the basis of our previous miRNA profiling data of human AML patients[14]. However, the underlying mechanism for this phenomenon is not understood. Thus, we can hypothesize that PML-RARa might promote leukemia formation through the upregulation of miR-125b and that miR-125 expression level is important for both the initiation and maintenance of AML in cooperation with not only MLL-AF9 but also PML-RARa. The other speculation is that miR-125b can play a parallel role as PML-RARa in the process of leukemia generation. To test these hypotheses, we generated mouse AML by crossing inducible miR-125b mouse with mouse carrying PML-RARa fusion gene from Dr. Timothy Ley’s group and explored whether miR-125b can promote the initiation of PML-RARa gene and induce APL.
-
This study was performed under a protocol approved by the Institutional Animal Care and Use Committee of Yale University. All mice were maintained at the Yale Animal Resource Center. Ctsg-PML-RARA (PR) mice on a C57BL/6 background were gifted by Dr. Timothy Ley’s group, as published[6]. C57BL/6 mice and the Rosa26-rtTA-M2 mice2 were from Jackson Laboratory[15]. The doxycycline-inducible miR-125b allele (i125b) was recently generated by our group. Briefly, mouse A2Lox.cre embryonic stem cells were used with an amiR-125b-containing targeting vector to generate the knock-in line 818-7, which generated knock-in mice through blastocyst injection performed by Yale Animal Genomics Services. Germline transmitted mice were crossed with Rosa26-rtTAm2 mice and backcrossed for six generations onto the C57BL/6 background (National Cancer Institute strain no. 01B96) to generate Ri125b mice[12,14]. For simplicity, the resultant mice are referred to as Ri125b mice, with male Ri125b mice having a genotype of rtTA/rtTA, i125b/i125y and female Ri125b mice having a genotype rtTA/rtTA, i125b/i125b. Genotyping was performed as published[6,12,14].
-
A total of 366 patients with AML data who were admitted between June 2000 and May 2020 were recruited following a protocol approved by the Chinese PLA General Hospital. A total of 264 males and 102 females were included, aged 18 to 64 years. AML was diagnosed by morphology, immunology, cytogenetics, and molecular biology according to the French-American-British (FAB) cooperative group subtype and World Health Organization classification system at the first visit. Fifty cases of normal data were from healthy volunteers aged above 18 years. WBC counts and bone marrow blast percentage were obtained from routine clinical tests. The patient characteristics are summarized in Table 1.
Table 1. Clinical characteristics of 366 clinical patients with AML
FAB subtype Patients (n) Gender (M/F) Median age at
diagnosis (Years)WBC
(1 x 103/μL)BM Blast% Ratio of
WBC/Blast%M1 30 23/7 41 75.89 ± 4.06 88.9 ± 4.56 0.83 ± 1.12 M2 57 38/19 41 28.43 ± 44.85 51.48 ± 20.93 0.56 ± 0.84 M3 (APL) 109 66/43 37 9.24 ± 15.60 79.03 ± 14.56 0.11 ± 0.18 M4 61 37/24 50 44.57 ± 60.24 52.79 ± 20.42 0.80 ± 1.06 M5 59 33/26 41 50.47 ± 79.37 68.80 ± 19.57 0.66 ± 0.88 Normal 50 35/15 55 6.15 ± 1.46 N/A N/A Note. FAB: French-American-Britiish1cooperative1group; WBC: White blood cells; BM: Bone marrow; AML, acute myeloid leukemia; APL, Acute promyelocytic leukemia. M1: Acute myeloid leukemia of FAB subtype M1; M2: Acute myeloid leukemia of FAB subtype M2; M3: Acute promyelocytic leukemia; M4: Acute myelomonocytic leukemia; M5: Acute monocytic leukemia. The study design and protocols were approved by the Ethics Committee of the Chinese PLA General Hospital. Written informed consent was obtained from each enrolled subject.
-
PR mice were crossed with Rosa26-rtTA-M2 (rtTA allele) and i125b alleles to generate the PR+rtTA, Ri125b, and PR+Ri125b genotypes. Mice were administered with 2 mg/mL Dox water supplemented with 1% sucrose (+Dox) or with regular water (−Dox) from 6 weeks of age. The survival of the mice was monitored with moribund events documented for the Kaplan–Meier curve.
-
Recipient wild-type C57/Bl6 mice were sublethally irradiated at 6 Gy on the day or the day before the transplantation. Leukemia cells for secondary transplantation were harvested from the bone marrow of primary moribund mice, which contained –60% to > 90% leukemia blasts. Cells from primary PR+Ri125b+Dox leukemia mice were used, with 125,000 cells transplanted into each recipient through tail vein injection. On the same day of transplantation, recipient mice were administered with 2 mg/mL Dox water supplemented with 1% sucrose or with regular water.
-
Peripheral blood (PB) was collected from the tail vein of mice. Complete blood count (CBC) count was performed with the PB samples on a Hemavet machine (Drew Scientific). Red blood cells were lysed using ACK lysis buffer (Thermo Fisher) before the PB samples were analyzed by performing flow cytometry[16]. Bone marrow cells from moribund mice were collected from the femur and tibia. Bone marrow cells were brushed with fetal bovine serum onto slides for bone marrow smears. Slides were stained with May–Grünwald–Giemsa (Sigma MG500) following the manufacturer’s protocol.
To isolate Mac1+ cells, cells were stained with PE-CD11b antibody (Biolegend) and sorted on a FACS Aria machine (BD).
For flow cytometry, all data were acquired on LSRII (BD) and analyzed with FlowJo 8.6.
-
Total RNA was extracted from cells using TRIzol (Invitrogen) according to the manufacturer’s instructions. For qRT-PCR of miR-125a and miR-125b, cDNA synthesis was performed with 500 ng of total RNA with miRScript II RT KIT (Qiagen) and subjected to SYBR green (Applied Biosystems) qPCR following the manufacturer’s protocol. Real-time PCR was performed using CFX96 Real-Time System (Bio-Rad). Primers for miR-125a and miR-125b were purchased from Qiagen. U6 small RNA was used as a control in qPCR. Relative expressions were calculated by 2−ΔCT.
-
Continuous variables underlying normal distribution were presented as mean ± standard deviations, and the between-group comparisons were tested by using two independent samples t-tests. If a variable was skewed, then it was described using the median (interquartile range) and compared using a nonparametric approach. Survival analyses were identified with bivariate analyses using Kaplan–Meier curves and log-rank testing for comparison. All the tests were two-tailed at a significance level of 0.05 unless otherwise stated. All the analyses were performed by using SPSS 22.0 (IBM Corp., Armonk, NY, USA).
-
We previously noted that miR-125 family miRNAs were often overexpressed in adult AML samples[14]. Particularly, we noticed that the majority of PML-RARα+ APL samples showed miR-125b and/or miR-125a overexpression as compared with normal controls or other AML subtypes. This result was further confirmed by examining the publicly available TCGA AML dataset (Figure 1A), in which the majority of PML-RARα+ APL cases generally showed higher levels of miR-125b overexpression compared with other AML subtypes.
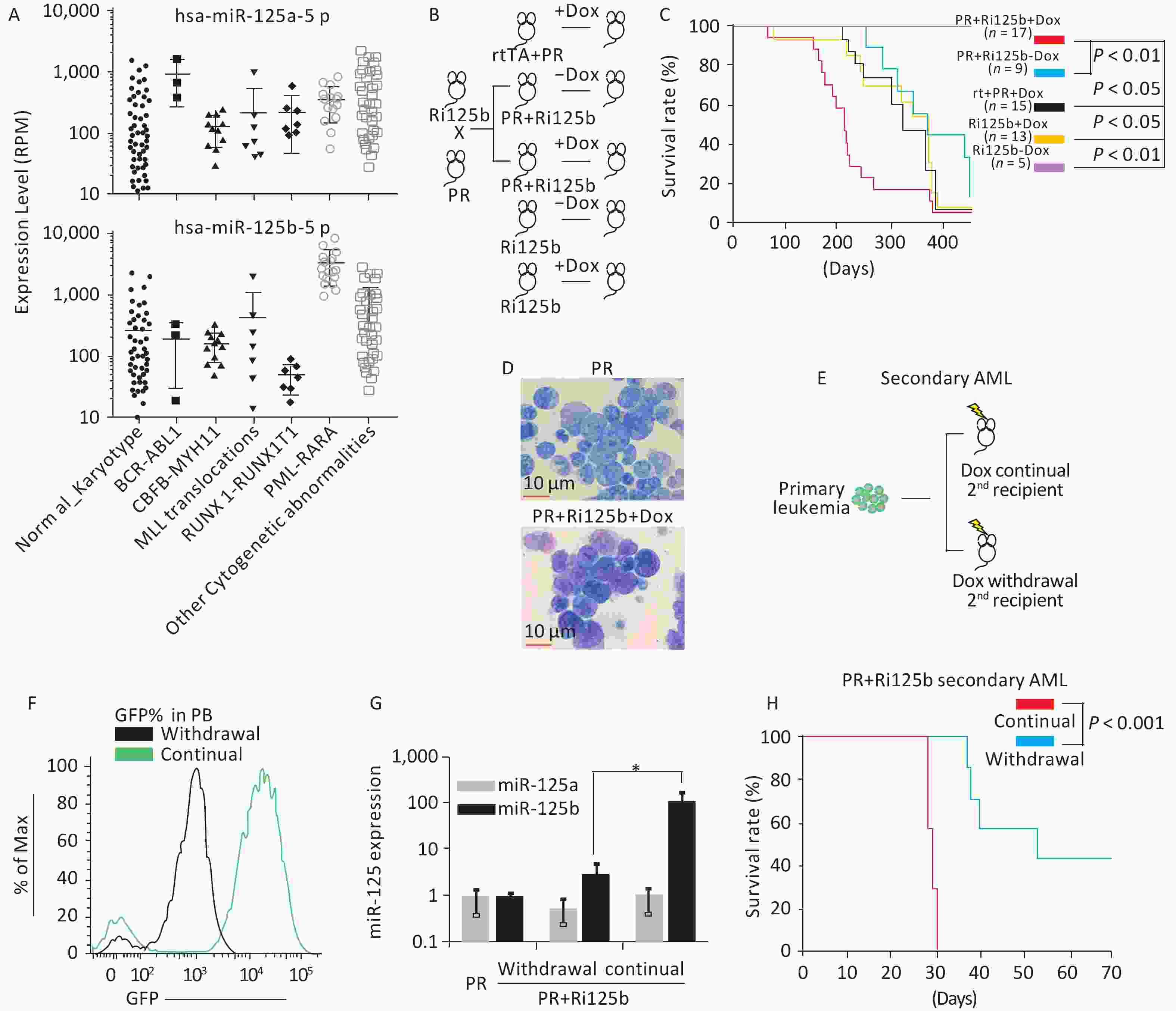
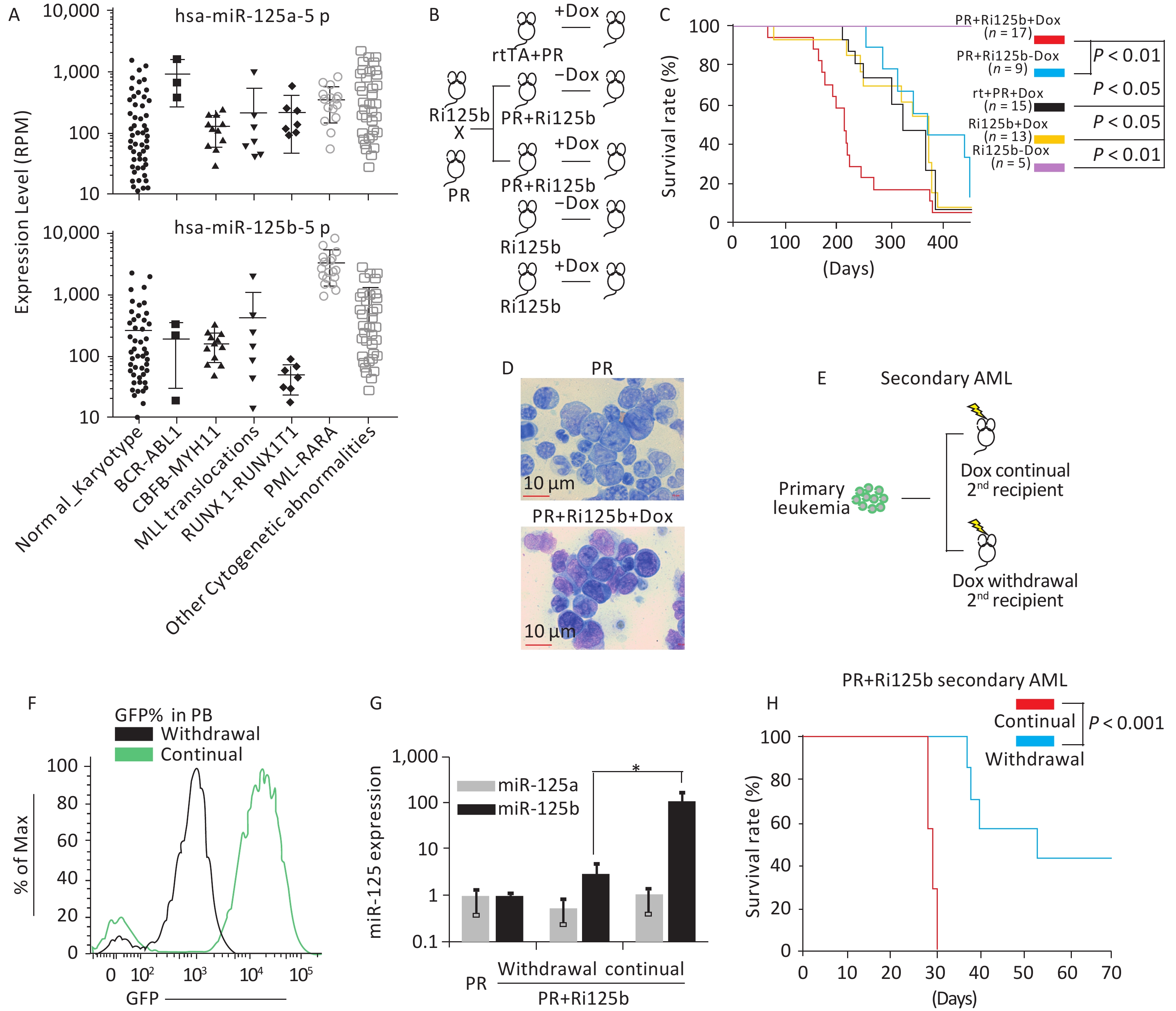
Figure 1. miR-125b accelerates the leukemogenesis induced by PR. (A) Expression levels of miR125b and miR125a in the TCGA dataset plotted across multiple AML subtypes, in reads per million. (B) Schematics of the experiment of PML-RARA (PR) and miR-125b-induced leukemogenesis, including control groups. (C) Kaplan–Meier survival curves of the indicated cohorts of mice. P values are indicated. (D) Representative Giemsa staining of bone marrow cells from moribund PR+Ri125b+Dox or PR mice. (E) Schematics of the experiment to analyze leukemia dependence on miR-125b overexpression. Primary AML cells from PR+Ri125b+Dox mice were transplanted into lethally irradiated wild-type mice. Secondary recipients were randomized into cohorts for continued Dox or withdrawal of Dox. (F) Representative flow cytometry plots of PB four weeks after Dox withdrawal in secondary PR+Ri125b recipients. (G) Relative miR-125a and miR-125b expression levels in bone marrow cells of PR mice, PR+Ri125b mice with continued Dox or Dox withdrawal. U6 was used as a control. *P < 0.05. (H) Kaplan–Meier survival curves of PR+Ri125b mice with continued Dox (n = 7) or Dox withdrawal (n = 7), P < 0.001. AML, acute myeloid leukemia; GFP, green fluorescent protein.
The overexpression of miR-125b in APL led us to ask whether miR-125b cooperates with PR in leukemogenesis. We previously published a knock-in inducible mouse model Ri125b, in which a human miR-125b-GFP transgene cassette was under the control of a tetracycline-responsive promoter (i125b allele) and was crossed with a rtTA allele in Rosa26 locus to provide doxycycline (Dox)-inducible expression of miR-125b (referred to as Ri125b mice)[14]. We further showed that this design resulted in 25-fold overexpression of miR-125b in a murine leukemia model[14]. We thus crossed the Ri125b mice with a PML-RARA knock-in allele (abbreviated as the “PR” allele)[6]. Adult compound knock-in mice (referred to as PR+Ri125b) were induced with doxycycline or treated with regular water from 6 weeks of age (Figure 1B). The only abnormality that we observed in these mice was the development of leukemia despite miR-125b being induced ubiquitously. Consistent with the PR allele being a weak oncogene, PR+Ri125b-Dox mice (without miR-125b induction) and PR+rtTA+Dox mice (without i125b allele) had a long latency before developing lethal leukemia. These cohorts showed a median survival of 404 and 345 days, respectively (Figure 1C), which are comparable to the published PR-only mice[6]. In contrast, Ri125b+Dox mice (without PR allele) developed myeloproliferative neoplasm-like symptoms, similar to our previous observations[14]. In addition, we did not notice the development of full-blown AML in this cohort. Unlike multiple control groups, PR+Ri125b+Dox mice exhibited accelerated development of lethal AML, with a median survival of 212 days (Figure 1C). An examination of the bone marrow blast cell morphology indicated APL, similar to PR-only mice (Figure 1D). Figure 1D shows the morphology of bone marrow (BM) cells in both Ri125b+Dox mice and Ri125b-only mice (Giemsa staining showed abnormal promyelocytes). These data support the idea that miR-125b accelerates PML-RARα-driven APL.
We next examined whether the development of APL in the presence of miR-125b overexpression is dependent on the continued expression of this miRNA. We took advantage of the doxycycline inducibility of the Ri125b allele and transplanted primary APL cells from PR+Ri125b+Dox mice into wild-type recipients (Figure 1E). This transplantation effectively limited the inducibility of miR-125b within the hematopoietic system, as host cells were wild-type. Transplant recipient mice were randomly divided, with half of the mice receiving continued Dox treatment and the other half being subjected to Dox withdrawal. Withdrawal of Dox lowered the GFP mean fluorescent intensity in donor-derived PB cells, as expected for effective cessation of Dox induction (Figure 1F). Consistently, miR-125b expression decreased by > 40-fold (Figure 1G), indicating that transgene expression was reduced successfully. We observed that the removal of Ri125b induction (secondary PR+Ri125b, Dox withdrawal) led to significantly prolonged survival of mice compared with mice with continued miR-125b induction (secondary PR+Ri125b, continued Dox) (Figure 1H). Overall, the results indicate that miR-125b-overexpressing APL is partially dependent on the continued overexpression of this miRNA.
-
The association of miR-125b overexpression in APL specimens may also be contributed by PR regulating miR-125a/b expression. To test this possibility, we examined bone marrow cells from wild-type mice, from PR mice before the onset of APL, or from leukemia-bearing PR mice. The expression levels of miR-125a and miR-125b were determined by quantitative RT-PCR, and we did not observe increases in the expression of these two miRNAs (Figure 2A). Purification of Mac-1+ myeloid cells from wild-type mice or PR mice before APL onset further confirmed that wild-type and PR cells had similar miR-125a and miR-125b expression levels (Figure 2B). These results suggest that PR alone cannot account for the upregulation of miR-125a and miR-125b. These data also suggest that the deregulation of miR-125b and miR-125a in human APL cases is likely due to other genetic or epigenetic changes.
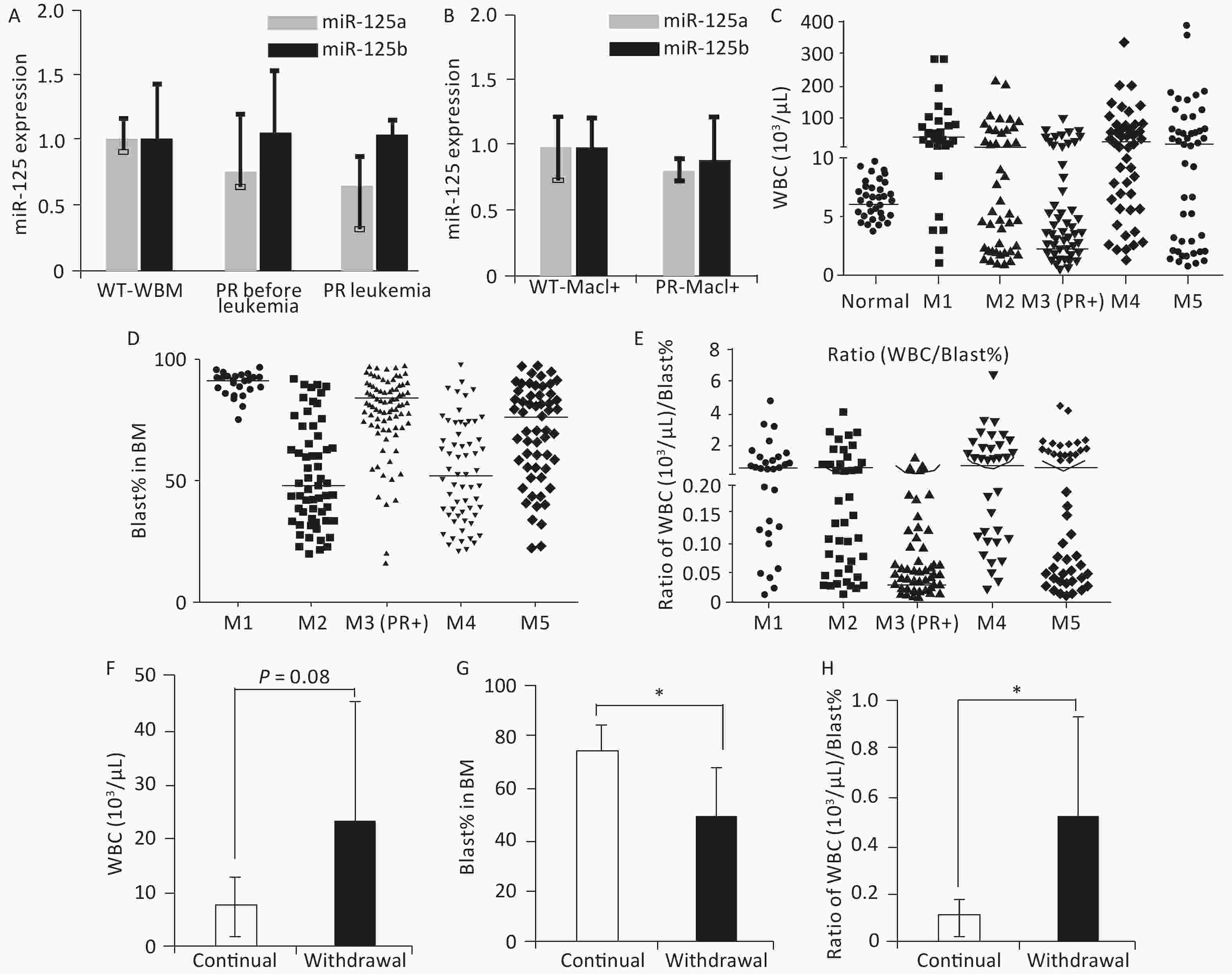
Figure 2. Features associated with peripheral WBC counts in human and mouse APL. (A) Relative miR-125a-5p and miR-125b-5p expression in bone marrow cells in wild-type mice, PR mice before APL onset, and PR mice with APL was determined by qRT-PCR, n = 3. (B) Mac1+ myeloid cells sorted from the bone marrow of wild-type and PR mice before APL onset. Relative miR-125a-5p and miR-125b-5p expression was determined by qRT-PCR. (C) WBC counts from a cohort of human AMLs at disease presentation plotted according to their FAB subtypes. For the M3 subtype, only PML-RARA+ (PR+) cases are shown. WBC data from individuals without hematological disorders were used as normal (n = 50). The horizontal lines represent the medians of the corresponding groups. FAB-M1 (n = 30); FAB-M2 (n = 57); FAB-M3 (PR+) (n = 109); FAB-M4 (n = 61); FAB-M5 (n = 59). (D) Bone marrow blast percentage data from the same patient cohort as in (C). (E) Ratios of WBC (103/μL) to bone marrow blast percentage for data in (C, D). (F) WBC from secondary PR+Ri125b AML mice treated with Dox (continued, n = 5) or without Dox (withdrawal, n = 9). WBC was evaluated only in moribund mice with APL. (G) Bone marrow blast percentage data from the same mice described in (F) are shown. *P < 0.05. (H) Ratios of WBC (103/uL) to bone marrow blast percentage. *P < 0.05. Error bars represent standard deviations. WBC, white blood cell. M1: Acute myeloid leukemia of FAB subtype M1; M2: Acute myeloid leukemia of FAB subtype M2; M3: Acute promyelocytic leukemia; M4: Acute myelomonocytic leukemia; M5: Acute monocytic leukemia
PR patients have been sporadically reported to exhibit low peripheral WBC counts[3,4]. We systematically examined this notion in a cohort of 316 human AMLs (Table 1), among which 109 were PML-RARα+ M3 AMLs. Fifty human subjects who were not diagnosed with AML were also examined. A substantial fraction of PML-RARα+ patients had low WBC counts (Figure 2C). This result was not due to these patients being at an early stage of AML because most PML-RARα+ AMLs showed a high bone marrow blast percentage (Figure 2D). Findings show that 90.8% of PR+ AML patients exhibited a low WBC/BM blast% ratio (< 0.2 103/μL/%), which occurred more frequently than the other AML subtypes examined (Figure 2E).
To determine if miR-125b overexpression alters peripheral WBC, bone marrow blast percentage, or WBC/BM blast% ratio in murine PML-RARA AMLs, we examined secondary PR+Ri125b mice with continued Dox or Dox withdrawal. We observed that mice with continued Dox induction showed a trend of decreased peripheral WBC counts, unlike mice without Dox induction (Figure 2F). The Dox induction cohort also had a significantly elevated bone marrow blast percentage (Figure 2G). Overall, PR+Ri125b mice with continued Dox induction showed a significant decrease in WBC/bone marrow blast percentage ratios compared with Dox withdrawal mice (Figure 2H), reaching a level similar to the majority of human PR+ AMLs (< 0.2 103/μL percentage). Overall, the above data support the idea that miR-125b overexpression not only accelerates PML-RARα-driven APL but also modifies disease pathophysiology. These data also suggest that miR-125b plays a role in controlling leukemia cell distribution between different hematopoietic compartments.
-
In this study, we modeled the synergy and dependence of miR-125b overexpression in PR APL leukemogenesis in vivo. miR-125band/or miR-125a is overexpressed in most of the examined adult PML-RARα+ APL specimens. A recent study reported increased miR-125b expression in pediatric APL cases, indicating that this dysregulation is not limited to adults. Moreover, miR-125b-1 expression is likely due to PML-RARα-mediated NRF2 activity[17], thus explaining the elevated miR-125b-1 in primary APL samples, which might indicate that NRF2 activation induced by PML-RARα promotes microRNA 125b-1 expression in vitro and confers resistance to chemotherapy in acute promyelocytic leukemia.
We observed that miR-125b withdrawal had an effect on cell survival in the bulk leukemia population, suggesting that miR-125b overexpression selectively rewires and impacts the addiction of more mature leukemia cells to its overexpression. The effect of miR-125b on cellular apoptosis in the bulk leukemia cells is likely a consequence of miR-125b targeting multiple downstream targets simultaneously. Several groups, including ours, previously demonstrated that miR-125 family miRNAs are capable of inhibiting multiple proapoptotic genes[17-23]. For example, BAK1, PUMA, BMF, KLF13, and TP53INP1 are known targets of miR-125 family miRNAs[24-26]. However, none of these targets have been reported to individually rescue the effect of miR-125 overexpression, consistent with miR-125 exerting an effect on the proapoptotic network. The role of miR-125b in the signaling pathways that underlie cancer development and progression has been explored comprehensively, and miR-125b is known to act largely through its multiple molecular targets, which are involved in signaling cascades, such as the canonical Wnt/β-catenin, PI3K/Akt, Hexokinase 2, VEGR, and p53 pathways[27-33]. miR-125b is expressed in bone marrow multipotent progenitors and myeloid cells but is shut down in the B-cell lineage, and the gene encoding miR-125b lacks transcriptional activation markers in B-cells[34,35]. In addition to impairing B-cell egress, miR-125b dysregulation initially reduced pre-B-cell output but later induced pre-B-cell lymphoma/leukemia in mice[35].
Our data show that overexpression of miR-125b accelerates leukemogenesis by significantly shortening the disease latency, thus supporting the model in which miR-125b overexpression cooperates with PR in leukemogenesis in vivo[36,37]. In addition, we found that although miR-125b overexpression did not change the cellular morphology of the resultant APL cells, it significantly decreased the PB WBC count to bone marrow blast percentage ratio. As the majority of human PR+ APL cases also have low WBC count to bone marrow blast percentage ratios, our data suggest that dysregulation of miR-125b likely plays an important role in modifying the pathophysiology of this disease. WBC count in AML, although often correlated with survival, does not have a clear molecular explanation[38-41]. The molecular mechanisms downstream of miR-125b were previously studied in other AML settings. For example, we showed that miR-125b suppresses Tet2 and Tet2-downstream pathways in MLL-AF9-induced leukemogenesis[14]. However, the difficulty of culturing primary mouse APL cells significantly limits our ability to dissect the molecular mechanisms. Nevertheless, in vivo characterization of miR-125b mechanisms can be speculated to possibly assist in elucidating the pathways that govern leukemia cell trafficking in the future.
Computational models for potential miRNA-disease association inference have been developed, thus saving time and effort on experimental studies and providing new methods to explore molecular mechanisms and develop new drugs for patients with cancer[42-45]. More recently, some miRNA biomarkers, such as miR-382-5p[46], miR-139-5p[47], and miR-638[48], have been identified and have expanded our understanding of APL. Future studies should be undertaken to determine the clinical significance of the combination of miRNA biomarkers in APL through computational models and provide a useful model for the onset or outcome of the disease.
We observed that the PML-RARA-driven APL mouse model does not upregulate mir-125a or mir-125. Importantly, we found that the overexpression of mir-125b in PML-RARA-induced leukemia caused higher blast percentages compared with other AMLs and led to prolonged survival and reduced blast percentages after withdrawal of mir-125b overexpression. This result indicates that PML-RARA-induced leukemia is partially addicted to mir-125b expression. As such, mir-125b addition on top of PR might be a more faithful model for APL.
-
Using a recently developed doxycycline-inducible knock-in mouse model, this study demonstrated that miR-125b overexpression accelerates PML-RARA-induced leukemogenesis in vivo, with the resultant leukemia cells being partially addicted to miR-125b overexpression. miR-125b overexpression leads to low WBC/BM blast ratios in mice to a level similar to the majority of human APL cases. Overall, these findings indicate that targeting miR-125b may be a new therapeutic option for APL. The precise molecular mechanisms underlying in vivo characterization of miR-125b should be investigated in the future.
-
We thank Dr. Timothy Ley for kindly providing the PML-RARA knock-in mouse model.
-
The authors declare no conflict of interest.
-
GUO Bo designed and supervised the project; GUO Bo, QIN Ran, and CHEN Ji Jun designed the experiments; GUO Bo performed the experiments; all authors analyzed the data; GUO Bo prepared the figures; GUO Bo and QIN Ran wrote the first draft of the manuscript. All authors discussed the results of the experiments and edited and approved the final version of the manuscript.
doi: 10.3967/bes2022.067
MicroRNA-125b Accelerates and Promotes PML-RARa-driven Murine Acute Promyelocytic Leukemia
-
Abstract:
Objective Most acute promyelocytic leukemia cases are characterized by the PML-RARa fusion oncogene and low white cell counts in peripheral blood. Methods Based on the frequent overexpression of miR-125-family miRNAs in acute promyelocytic leukemia, we examined the consequence of this phenomenon by using an inducible mouse model overexpressing human miR-125b. Results MiR-125b expression significantly accelerates PML-RARa-induced leukemogenesis, with the resultant induced leukemia being partially dependent on continued miR-125b overexpression. Interestingly, miR-125b expression led to low peripheral white cell counts to bone marrow blast percentage ratio, confirming the clinical observation in acute promyelocytic leukemia patients. Conclusion This study suggests that dysregulated miR-125b expression is actively involved in disease progression and pathophysiology of acute promyelocytic leukemia, indicating that targeting miR-125b may represent a new therapeutic option for acute promyelocytic leukemia. -
Key words:
- miR-125b /
- PML-RARA /
- White blood cell /
- Bone marrow blast
-
Figure 1. miR-125b accelerates the leukemogenesis induced by PR. (A) Expression levels of miR125b and miR125a in the TCGA dataset plotted across multiple AML subtypes, in reads per million. (B) Schematics of the experiment of PML-RARA (PR) and miR-125b-induced leukemogenesis, including control groups. (C) Kaplan–Meier survival curves of the indicated cohorts of mice. P values are indicated. (D) Representative Giemsa staining of bone marrow cells from moribund PR+Ri125b+Dox or PR mice. (E) Schematics of the experiment to analyze leukemia dependence on miR-125b overexpression. Primary AML cells from PR+Ri125b+Dox mice were transplanted into lethally irradiated wild-type mice. Secondary recipients were randomized into cohorts for continued Dox or withdrawal of Dox. (F) Representative flow cytometry plots of PB four weeks after Dox withdrawal in secondary PR+Ri125b recipients. (G) Relative miR-125a and miR-125b expression levels in bone marrow cells of PR mice, PR+Ri125b mice with continued Dox or Dox withdrawal. U6 was used as a control. *P < 0.05. (H) Kaplan–Meier survival curves of PR+Ri125b mice with continued Dox (n = 7) or Dox withdrawal (n = 7), P < 0.001. AML, acute myeloid leukemia; GFP, green fluorescent protein.
Figure 2. Features associated with peripheral WBC counts in human and mouse APL. (A) Relative miR-125a-5p and miR-125b-5p expression in bone marrow cells in wild-type mice, PR mice before APL onset, and PR mice with APL was determined by qRT-PCR, n = 3. (B) Mac1+ myeloid cells sorted from the bone marrow of wild-type and PR mice before APL onset. Relative miR-125a-5p and miR-125b-5p expression was determined by qRT-PCR. (C) WBC counts from a cohort of human AMLs at disease presentation plotted according to their FAB subtypes. For the M3 subtype, only PML-RARA+ (PR+) cases are shown. WBC data from individuals without hematological disorders were used as normal (n = 50). The horizontal lines represent the medians of the corresponding groups. FAB-M1 (n = 30); FAB-M2 (n = 57); FAB-M3 (PR+) (n = 109); FAB-M4 (n = 61); FAB-M5 (n = 59). (D) Bone marrow blast percentage data from the same patient cohort as in (C). (E) Ratios of WBC (103/μL) to bone marrow blast percentage for data in (C, D). (F) WBC from secondary PR+Ri125b AML mice treated with Dox (continued, n = 5) or without Dox (withdrawal, n = 9). WBC was evaluated only in moribund mice with APL. (G) Bone marrow blast percentage data from the same mice described in (F) are shown. *P < 0.05. (H) Ratios of WBC (103/uL) to bone marrow blast percentage. *P < 0.05. Error bars represent standard deviations. WBC, white blood cell. M1: Acute myeloid leukemia of FAB subtype M1; M2: Acute myeloid leukemia of FAB subtype M2; M3: Acute promyelocytic leukemia; M4: Acute myelomonocytic leukemia; M5: Acute monocytic leukemia
Table 1. Clinical characteristics of 366 clinical patients with AML
FAB subtype Patients (n) Gender (M/F) Median age at
diagnosis (Years)WBC
(1 x 103/μL)BM Blast% Ratio of
WBC/Blast%M1 30 23/7 41 75.89 ± 4.06 88.9 ± 4.56 0.83 ± 1.12 M2 57 38/19 41 28.43 ± 44.85 51.48 ± 20.93 0.56 ± 0.84 M3 (APL) 109 66/43 37 9.24 ± 15.60 79.03 ± 14.56 0.11 ± 0.18 M4 61 37/24 50 44.57 ± 60.24 52.79 ± 20.42 0.80 ± 1.06 M5 59 33/26 41 50.47 ± 79.37 68.80 ± 19.57 0.66 ± 0.88 Normal 50 35/15 55 6.15 ± 1.46 N/A N/A Note. FAB: French-American-Britiish1cooperative1group; WBC: White blood cells; BM: Bone marrow; AML, acute myeloid leukemia; APL, Acute promyelocytic leukemia. M1: Acute myeloid leukemia of FAB subtype M1; M2: Acute myeloid leukemia of FAB subtype M2; M3: Acute promyelocytic leukemia; M4: Acute myelomonocytic leukemia; M5: Acute monocytic leukemia. -
[1] Arber DA. The 2016 WHO classification of acute myeloid leukemia: what the practicing clinician needs to know. Semin Hematol, 2019; 56, 90−5. doi: 10.1053/j.seminhematol.2018.08.002 [2] Hwang SM. Classification of acute myeloid leukemia. Blood Res, 2020; 55, S1−4. doi: 10.5045/br.2020.S001 [3] Lo-Coco F, Avvisati G, Vignetti M, et al. Retinoic acid and arsenic trioxide for acute promyelocytic leukemia. N Engl J Med, 2013; 369, 111−21. doi: 10.1056/NEJMoa1300874 [4] Tallman MS, Altman JK. How I treat acute promyelocytic leukemia. Blood, 2009; 114, 5126−35. [5] Gaillard C, Surianarayanan S, Bentley T, et al. Identification of IRF8 as a potent tumor suppressor in murine acute promyelocytic leukemia. Blood Adv, 2018; 2, 2462−6. doi: 10.1182/bloodadvances.2018018929 [6] Westervelt P, Lane AA, Pollock JL, et al. High-penetrance mouse model of acute promyelocytic leukemia with very low levels of PML-RARα expression. Blood, 2003; 102, 1857−65. doi: 10.1182/blood-2002-12-3779 [7] Brown D, Kogan S, Lagasse E, et al. A PMLRARα transgene initiates murine acute promyelocytic leukemia. Proc Natl Acad Sci USA, 1997; 94, 2551−6. doi: 10.1073/pnas.94.6.2551 [8] He LZ, Tribioli C, Rivi R, et al. Acute leukemia with promyelocytic features in PML/RARα transgenic mice. Proc Natl Acad Sci USA, 1997; 94, 5302−7. doi: 10.1073/pnas.94.10.5302 [9] Welch JS, Yuan WL, Ley TJ. PML-RARA can increase hematopoietic self-renewal without causing a myeloproliferative disease in mice. J Clin Invest, 2011; 121, 1636−45. doi: 10.1172/JCI42953 [10] He CC, Luo B, Jiang N, et al. OncomiR or antioncomiR: role of miRNAs in acute myeloid leukemia. Leuk Lymphoma, 2019; 60, 284−94. doi: 10.1080/10428194.2018.1480769 [11] Liu Y, Cheng ZH, Pang YF, et al. Role of microRNAs, circRNAs and long noncoding RNAs in acute myeloid leukemia. J Hematol Oncol, 2019; 12, 51. doi: 10.1186/s13045-019-0734-5 [12] Guo SQ, Bai HT, Megyola CM, et al. Complex oncogene dependence in microRNA-125a-induced myeloproliferative neoplasms. Proc Natl Acad Sci USA, 2012; 109, 16636−41. doi: 10.1073/pnas.1213196109 [13] Sharifi H, Jafari Najaf Abadi MH, Razi E, et al. MicroRNAs and response to therapy in leukemia. J Cell Biochem, 2019; 120, 14233−46. doi: 10.1002/jcb.28892 [14] Liu J, Guo B, Chen Z, et al. miR-125b promotes MLL-AF9-driven murine acute myeloid leukemia involving a VEGFA-mediated non-cell-intrinsic mechanism. Blood, 2017; 129, 1491−502. doi: 10.1182/blood-2016-06-721027 [15] Iacovino M, Bosnakovski D, Fey H, et al. Inducible cassette exchange: a rapid and efficient system enabling conditional gene expression in embryonic stem and primary cells. Stem Cells, 2011; 29, 1580−8. doi: 10.1002/stem.715 [16] He YX, Guo YX, Zhang Y, et al. Ionizing radiation-induced RPL23a reduction regulates apoptosis via RPL11-MDM2-p53 pathway in mouse spermatogonia. Biomed Environ Sci, 2021; 34, 789−802. [17] Yu XB, Mansouri A, Liu ZD, et al. NRF2 activation induced by PML-RARα promotes microRNA 125b-1 expression and confers resistance to chemotherapy in acute promyelocytic leukemia. Clin Transl Med, 2021; 11, e418. [18] He JH, Huang ZY, He ML, et al. Circular RNA MAPK4 (circ-MAPK4) inhibits cell apoptosis via MAPK signaling pathway by sponging miR-125a-3p in gliomas. Mol Cancer, 2020; 19, 17. doi: 10.1186/s12943-019-1120-1 [19] Xu XB, Lai YY, Zhou WZ, et al. Lentiviral delivery of a shRNA sequence analogous to miR-4319/miR-125-5p induces apoptosis in NSCLC cells by arresting G2/M phase. Cell Biochem, 2019; 120, 14017−27. doi: 10.1002/jcb.28676 [20] Yao DH, Zhou ZY, Wang PF, et al. MiR-125-5p/IL-6R axis regulates macrophage inflammatory response and intestinal epithelial cell apoptosis in ulcerative colitis through JAK1/STAT3 and NF-κB pathway. Cell Cycle, 2021; 20, 2547−64. doi: 10.1080/15384101.2021.1995128 [21] Gu YZ, Zhao SG, Wan JP, et al. Hsa-miRNA-125b may induce apoptosis of HTR8/SVneo cells by targeting MCL1. Reprod Biol, 2019; 19, 368−73. doi: 10.1016/j.repbio.2019.09.004 [22] Lo YL, Wang CS, Chen YC, et al. Mitochondrion-directed nanoparticles loaded with a natural compound and a microRNA for promoting cancer cell death via the modulation of tumor metabolism and mitochondrial dynamics. Pharmaceutics, 2020; 12, 756. doi: 10.3390/pharmaceutics12080756 [23] Yu CW, Tang WB, Lu R, et al. Human adipose-derived mesenchymal stem cells promote lymphocyte apoptosis and alleviate atherosclerosis via miR-125b-1-3p/BCL11B signal axis. Ann Palliat Med, 2021; 10, 2123−33. doi: 10.21037/apm-21-49 [24] Enomoto Y, Kitaura J, Hatakeyama K, et al. Eμ/miR-125b transgenic mice develop lethal B-cell malignancies. Leukemia, 2011; 25, 1849−56. doi: 10.1038/leu.2011.166 [25] Shi XB, Xue LR, Ma AH, et al. MiR-125b promotes growth of prostate cancer xenograft tumor through targeting pro-apoptotic genes. Prostate, 2011; 71, 538−49. doi: 10.1002/pros.21270 [26] Surdziel E, Cabanski M, Dallmann I, et al. Enforced expression of miR-125b affects myelopoiesis by targeting multiple signaling pathways. Blood, 2011; 117, 4338−48. doi: 10.1182/blood-2010-06-289058 [27] Wang HJ, Guo YQ, Tan G, et al. MiR-125b regulates side population in breast cancer and confers a chemoresistant phenotype. J Cell Biochem, 2013; 114, 2248−57. doi: 10.1002/jcb.24574 [28] Biamonte F, Battaglia AM, Zolea F, et al. Ferritin heavy subunit enhances apoptosis of non-small cell lung cancer cells through modulation of miR-125b/p53 axis. Cell Death Dis, 2018; 9, 1174. doi: 10.1038/s41419-018-1216-3 [29] Hou PF, Li HL, Yong HM, et al. PinX1 represses renal cancer angiogenesis via the mir-125a-3p/VEGF signaling pathway. Angiogenesis, 2019; 22, 507−19. doi: 10.1007/s10456-019-09675-z [30] Incoronato M, Grimaldi AM, Mirabelli P, et al. Circulating miRNAs in untreated breast cancer: an exploratory multimodality morpho-functional study. Cancers (Basel), 2019; 11, 876. doi: 10.3390/cancers11060876 [31] Davari N, Ahmadpour F, Kiani AA, et al. Evaluation of microRNA-223 and microRNA-125a expression association with STAT3 and Bcl2 genes in blood leukocytes of CLL patients: a case-control study. BMC Res Notes, 2021; 14, 21. doi: 10.1186/s13104-020-05428-0 [32] Zhang Y, Liu YF, Xu XJ. Knockdown of LncRNA-UCA1 suppresses chemoresistance of pediatric AML by inhibiting glycolysis through the microRNA-125a/hexokinase 2 pathway. J Cell Biochem, 2018; 119, 6296−308. doi: 10.1002/jcb.26899 [33] Nie J, Jiang HC, Zhou YC, et al. MiR-125b regulates the proliferation and metastasis of triple negative breast cancer cells via the Wnt/β-catenin pathway and EMT. Biosci Biotechnol Biochem, 2019; 83, 1062−71. doi: 10.1080/09168451.2019.1584521 [34] El-Khazragy N, Elshimy AA, Hassan SS, et al. Dysregulation of miR-125b predicts poor response to therapy in pediatric acute lymphoblastic leukemia. J Cell Biochem, 2019; 120, 7428−38. doi: 10.1002/jcb.28017 [35] Li GD, So AYL, Sookram R, et al. Casellas R, Baltimore D. Epigenetic silencing of miR-125b is required for normal B-cell development. Blood, 2018; 131, 1920−30. [36] Fu K, Zhang L, Liu R, et al. MiR-125 inhibited cervical cancer progression by regulating VEGF and PI3K/AKT signaling pathway. World J Surg Oncol, 2020; 18, 115. doi: 10.1186/s12957-020-01881-0 [37] Yang MY, Tang XL, Wang Z, et al. miR-125 inhibits colorectal cancer proliferation and invasion by targeting TAZ. Biosci Rep, 2019; 39, BSR20190193. doi: 10.1042/BSR20190193 [38] Xu LH, Wang JW, Wang Y, et al. Hyperleukocytosis predicts inferior clinical outcome in pediatric acute myeloid leukemia. Hematology, 2020; 25, 507−14. doi: 10.1080/16078454.2020.1859169 [39] Bewersdorf JP, Zeidan AM. Hyperleukocytosis and leukostasis in acute myeloid leukemia: can a better understanding of the underlying molecular pathophysiology lead to novel treatments? Cells, 2020; 9, 2310. [40] Wang N, Desai A, Ge B, et al. Prognostic value of hypoalbuminemia at diagnosis in de novo non-M3 acute myeloid leukemia. Leuk Lymphoma, 2020; 61, 641−9. doi: 10.1080/10428194.2019.1686499 [41] Jahic A, Iljazovic E, Hasic S, et al. Prognostic parameters of acute myeloid leukaemia at presentation. Med Arch, 2017; 71, 20−4. doi: 10.5455/medarh.2017.71.20-24 [42] Chen X, Wang L, Qu J, et al. Predicting miRNA-disease association based on inductive matrix completion. Bioinformatics, 2018; 34, 4256−65. [43] Chen X, Xie D, Zhao Q, et al. MicroRNAs and complex diseases: from experimental results to computational models. Brief Bioinform, 2019; 20, 515−39. doi: 10.1093/bib/bbx130 [44] Chen X, Yin J, Qu J, et al. MDHGI: matrix decomposition and heterogeneous graph inference for miRNA-disease association prediction. PLoS Comput Biol, 2018; 14, e1006418. doi: 10.1371/journal.pcbi.1006418 [45] Chen X, Zhu CC, Yin J. Ensemble of decision tree reveals potential miRNA-disease associations. PLoS Comput Biol, 2019; 15, e1007209. doi: 10.1371/journal.pcbi.1007209 [46] Liu DD, Zhong L, Yuan Z, et al. miR-382-5p modulates the ATRA-induced differentiation of acute promyelocytic leukemia by targeting tumor suppressor PTEN. Cell Signal, 2019; 54, 1−9. doi: 10.1016/j.cellsig.2018.11.012 [47] Fu YY, Li LM, Hou JX, et al. miR-139-5p regulates the proliferation of acute promyelocytic leukemia cells by targeting MNT. J Oncol, 2021; 2021, 5522051. [48] Bai YY, Chen C, Guo XL, et al. miR-638 in circulating leukaemia cells as a non-invasive biomarker in diagnosis, treatment response and MRD surveillance of acute promyelocytic leukaemia. Cancer Biomark, 2020; 29, 125−37. doi: 10.3233/CBM-190899 -




 下载:
下载:



 Quick Links
Quick Links