-
Nonalcoholic fatty liver disease (NAFLD) is a rapidly growing health problem affecting approximately one-quarter of the world's population[1]. It encompasses a range of clinical phenotypes, from hepatic steatosis to liver cirrhosis and even hepatocellular carcinoma[2]. NAFLD has been linked to not only liver-related health issues and death but also a range of other health problems, including cardiovascular and chronic kidney diseases[3].
NAFLD is a complex multifactorial disease involving genetic, metabolic, and environmental factors[4-6]. Over the past three decades, more than 400 candidate gene association studies have been published on the associations between common variants and NAFLD risk, and some common variants, including PNPLA3 rs738409, TM6SF2 rs58542926, GCKR rs780094, have been studied frequently[7-10]. An increasing number of genome-wide association studies (GWASs) and whole-exome sequencing (WES) have been conducted in recent years. GWASs provide a broader and unbiased approach for the discovery of genes involved in complex genetic traits, whereas WES studies can directly capture relevant variations not interrogated by common genotyping platform designs, including rare variants[11]. Despite hundreds of genetic association studies published to date, results regarding the specific variant remain inconsistent, hindering our understanding of the genetic architecture of NAFLD. This may be attributed to small sample sizes in most studies, ethnic differences, and other factors. Meta-analysis is a powerful tool for increasing statistical strength and precision in detecting gene-disease associations[12]. However, previous meta-analyses on NAFLD susceptibility have mainly focused on one genetic variant or genetic variants within one gene, and few meta-analyses have encompassed all relevant types of studies, including candidate gene association studies, GWASs, and WES studies. In addition, previous meta-analyses have not evaluated the epidemiological credibility of these associations[13-15].
This study aimed to comprehensively summarize all genetic association studies of NAFLD, including candidate gene association, genome-wide association, and whole exome sequencing studies. This study will also conduct meta-analyses for genetic variants with sufficient available data, categorize related genes according to their genetic functions, and evaluate the epidemiological credibility of these associations using the Venice criteria proposed by the Human Genome Epidemiology Network (HuGENet)[16]. This study represents a comprehensive synthesis of genetic variants and functions associated with NAFLD, and will facilitate the interpretation of findings in the quest for genuine genetic susceptibility factors.
-
A comprehensive literature search was conducted in the Web of Science, PubMed, and Embase electronic databases to identify studies that analyzed genetic variants associated with NAFLD. The systematic search was first conducted on January 5, 2021, and updated on September 30, 2022. The detailed search terms are presented in Appendix A.
Studies included in this systematic review must satisfy the following criteria: (i) the study design must be a case-control, cohort, or cross-sectional association study in human subjects; (ii) the study must focus on the associations between genetic variants and NAFLD risk; (iii) NAFLD cases must have been diagnosed by clinicians, and meet the diagnostic criteria of NAFLD defined by the presence of steatosis in > 5% hepatocytes without secondary causes such as significant alcohol consumption, chronic viral hepatitis, long-term use of steatogenic medications, or monogenic hereditary disorders (including lecithin-cholesterol acyltransferase deficiency, cholesterol ester storage disease, Wolman's disease, etc); (iv) absence of hepatic steatosis in the control group; (v) if data or data subsets were published in more than one publication, the article with largest sample size was included; and (vi) article published in English. All records obtained through the database search were collected using EndNote software. After removing duplicates, the titles and abstracts were screened to exclude irrelevant studies, and the full texts of the remaining records were reviewed for eligibility. The reference lists of publications were also reviewed. Two reviewers (LI Ya Mei and XIAO Xiang) independently selected the studies.
-
The following information was extracted independently by two reviewers from eligible studies: first author, year of publication, country of origin, ethnicity of the study population, diagnostic criteria for NAFLD, sample size, source of controls, genotype methods, significant association (Yes/No), confounding factors adjusted in multivariate analysis models, and allele and genotype frequencies in cases and controls. Specifically, ethnicity was classified as Asian, Caucasian, Hispanic, African, or mixed, based on the ethnicity of at least 80% of the study population. The most recent gene names and variant accession numbers were used according to the National Center for Biotechnology Information (NCBI) database[17].
Two reviewers independently assessed the quality of the included studies using the Newcastle–Ottawa Scale (NOS)[18]. The NOS criteria evaluated the methodological quality of studies using a “star” rating system, with scores ranging from 0 star (worst) to 9 stars (best). Disagreements in data extraction and quality assessment were discussed among all the investigators and resolved by consensus.
-
The locations, coding proteins, and functional annotation information of the genes identified in previous studies were obtained by searching the human gene database (https://www.genecards.org/). Genes were divided into eight categories based on their main related functions, including lipid synthesis and metabolism, insulin resistance and glucose metabolism, adipokines/adipokine receptors, energy metabolism and obesity, oxidative stress and antioxidants, inflammatory and immune responses, and liver fibrosis.
A meta-analysis was performed for genetic variants with data available from at least five independent studies. The GWAS-identified variants were not included in this meta-analysis because they have been convincingly replicated in many studies. Genotype distributions in the control subjects were tested for Hardy–Weinberg equilibrium using the chi-square (χ2) test. Studies that did not conform to Hardy–Weinberg equilibrium were excluded from the meta-analysis. The odds ratios (ORs) and 95% confidence intervals (CIs) were calculated using five genetic models: allele contrast model (mutant [M] allele versus wild [W] allele), heterozygous model (WM versus WW), homozygous model (MM versus WW), recessive model (MM versus WW + WM), and dominant model (WM + MM versus WW)[19]. Random-effects models (Der Simonian–Laird method) were used in all meta-analyses[20]. Cochrane's Q test and I2 index were used to assess heterogeneity across eligible studies[21,22]. Meta-analyses, stratified by ethnicity, were conducted for variants with at least two data sources in each subgroup. Funnel plots and Egger's linear regression tests were used to assess potential publication bias in the included studies[23]. All tests were two-sided, and P value < 0.05 was considered statistically significant. Sensitivity analyses were conducted to evaluate the influence of each study on the overall estimates. Meta-analyses were performed using the STATA version 12.0 software (Stata Corporation, College Station, TX, USA).
-
The Venice criteria were applied to evaluate the epidemiological credibility of the significant associations identified in the meta-analysis[16]. Credibility was defined as strong (A), moderate (B), or weak (C) in three categories: the amount of evidence, replication of the association, and protection from bias. The amount of evidence was graded by the sum of the test alleles or genotypes among the cases and controls in the meta-analysis: A grade of A for > 1,000, B for 100–1,000, and C for < 100. Replication of the association was graded using the heterogeneity statistic: A for I2 < 25%, B for I2 between 25 and 50%, and C for I2 > 50%[22]. Protection from bias was graded as A, if there was no observable bias and bias was unlikely to explain the presence of the association, B, if bias was present, and C, if bias was evident or likely to explain the presence of the association. We also considered the magnitude of association in the assessment of protection from bias, and a score of C was assigned to an association when the summary OR deviated less than 1.15-fold from the null unless the association had been replicated prospectively by several studies with no evidence of publication bias. Next, the cumulative epidemiological evidence for significant associations was considered strong if all three grades were A, moderate if all three grades were A or B, and weak if any grade was C.
-
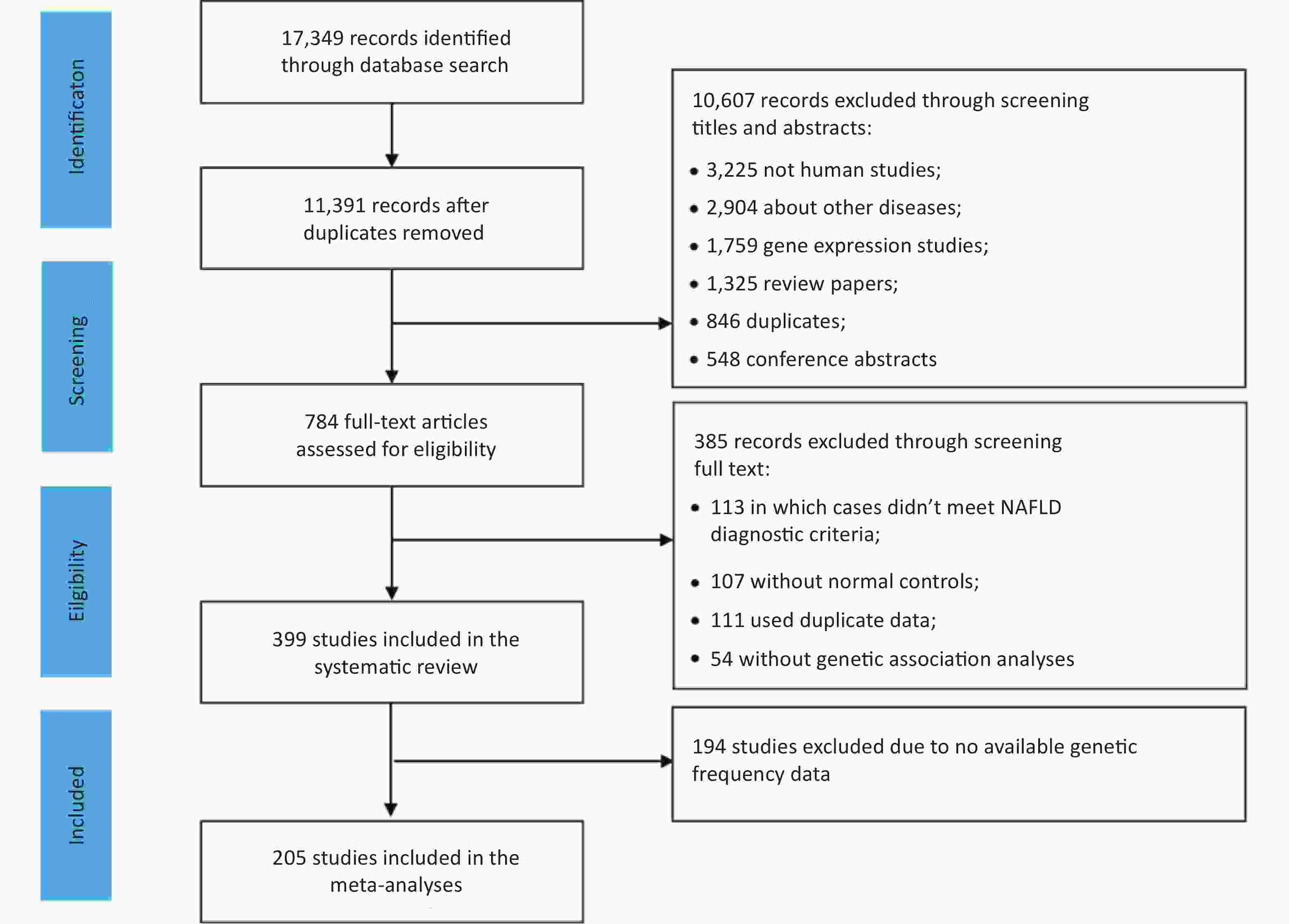
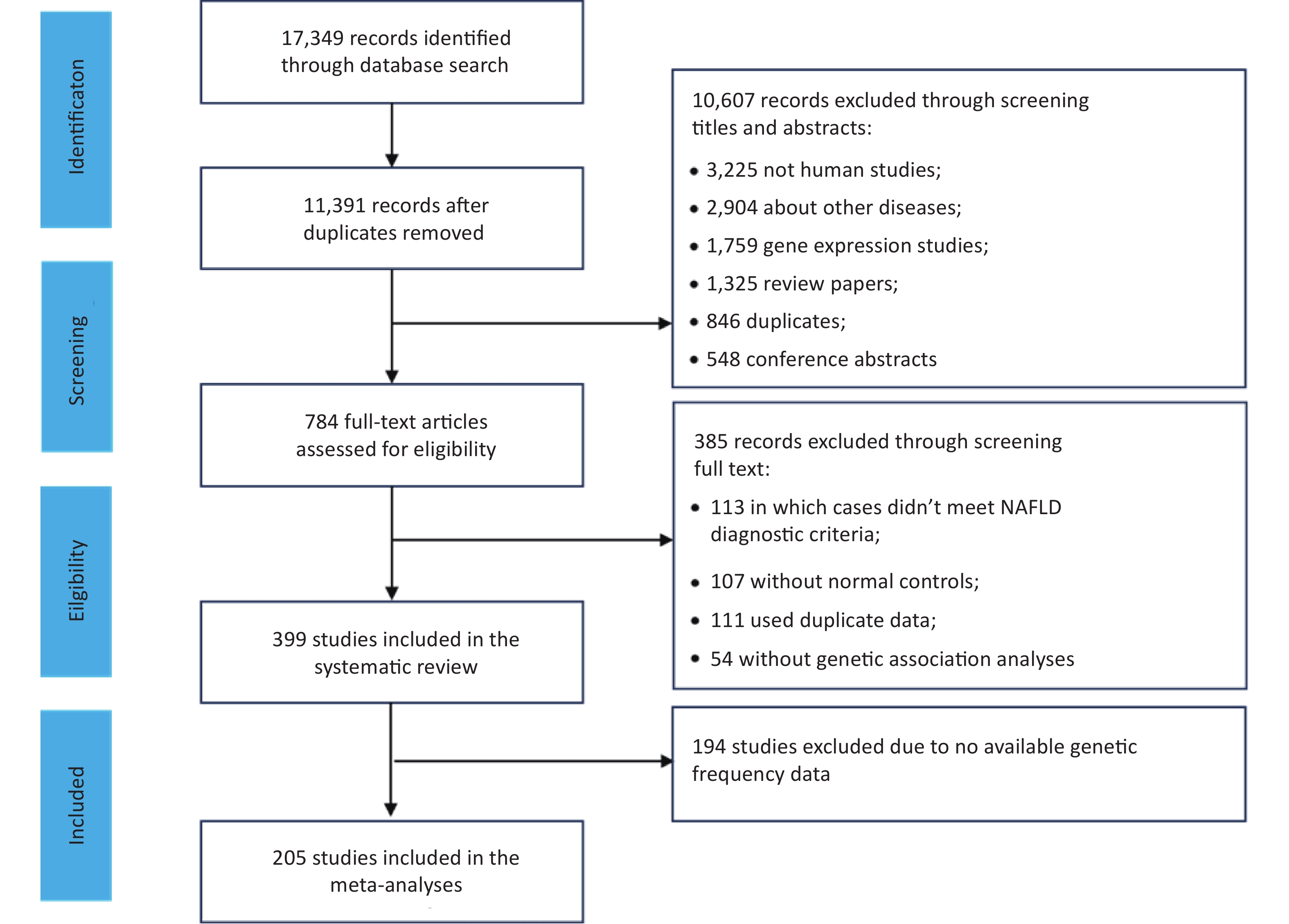
We identified 17,349 articles based on the literature searches, 11,391 non-duplicated articles were included in the title and abstract screening. Articles were excluded for non-human subjects (n = 3,225), irrelevant themes (n = 2,904), gene expression studies (n = 1,759), review articles (n = 1,325), duplicates (n = 846), or conference abstracts (n = 548), and the remaining 784 articles were included in the full-text review (Figure 1). In total, 399 eligible studies were identified, including 16 GWASs, 2 WES studies, and 381 candidate gene association studies. Of these, 205 studies with sufficient data were included in the meta-analysis of 25 variants of 17 genes.
Among the 16 GWASs, eight were conducted in the United States, four in Japan, two in South Korea, one in Australia, and one in Italy. Thirteen GWASs were conducted in adults, 2 studies were conducted in children, and 1 each in adults and children. The mean sample size was 5,710 ± 7,595 (ranged 234–27,374). At the same time, subsequent validation studies were performed followed by five GWAS studies. The basic characteristics of the GWASs are shown in Supplemental Table S1 (available in www.besjournal.com).
The characteristics of the 381 candidate gene association studies are presented in Supplemental Table S2 (available in www.besjournal.com). These studies analyzed 465 genetic variants in 173 candidate genes. The first candidate gene association study for NAFLD was published in 1998 and the number of such studies has gradually increased in recent decades. Most studies were conducted in China (n = 194), followed by Italy (n = 37), Japan (n = 21), Turkey (n = 18), Iran (n = 17), India (n = 16), the United States (n = 13), Egypt (n = 8), and other countries (n =57). The ethnic distribution was dominated by Asian (n = 251) and Caucasian (n = 102) populations. The vast majority of the studies included adults (n = 349), and the remaining 32 studies were conducted in children or a mixed population. The mean NOS score was 6.36 ± 1.11, suggesting that the overall research quality was acceptable.
-
In total, 72 variants were found to be significantly associated with NAFLD risk in the GWASs (Supplemental Table S1). Specifically, 52 variants were identified in adults, 15 variants were reported in children, and 6 variants were reported in both adults and children. Notably, PNPLA3 rs738409 has been reported not only in adults, but also in mixed adults and children. In addition, two WES studies have shown that three variants of two genes (PNPLA3, and PMPT) are associated with NAFLD susceptibility in adults.
-
Among the 465 genetic variants, common variants (416 variants in 157 genes) (MAF ≥ 5%) included synonymous single nucleotide polymorphisms (SNPs; n = 20), non-coding transcript SNPs (n = 8), intronic SNPs (n = 208), SNPs in 5´ or 3´ untranslated regions (UTRs; n = 28), missense variants (n = 66), upstream variants (n = 46), downstream variants (n = 4) and others (n = 36). Rare variants (49 variants in 34 genes) (MAF < 5%) included synonymous variants (n = 1), non-coding transcript variants (n = 2), intronic variants (n = 17), variants in 5´ or 3´ UTRs (n = 6), missense variants (n = 21), and two upstream variants (Supplemental Table S3, available in www.besjournal.com). Based on the main function of each gene, the 173 genes were divided into eight categories: lipid synthesis and metabolism (n = 49), insulin resistance and glucose metabolism (n = 10), adipokines/adipokine receptors (n = 6), energy metabolism and obesity (n = 6), oxidative stress and antioxidants (n = 22), inflammatory and immune response (n = 32), liver fibrosis (n = 11) and others (n = 37), as shown in Supplemental Table S4 (available in www.besjournal.com).
-
As shown in Table 1, 25 variants of the 17 genes were included in the meta-analysis. The information extracted for each variant used in the meta-analysis is presented in Supplemental Table S5 (available in www.besjournal.com). These meta-analyses included a mean of 14 studies (range: 6–75) and 3281 participants (range: 1,784–42,903). 11 variants in 10 genes showed statistically significant associations with NAFLD (P < 0.05), including four genes related to lipid synthesis and metabolism (MBOAT7, PEMT, PNPLA3, TM6SF2), 1 gene related to insulin resistance and glucose metabolism (GCKR), 2 genes related to adipokines/adipokine receptors (ADIPOQ, LEPR), 2 genes related to oxidative stress and antioxidants (HFE, MTHFR), and 1 gene related to inflammatory and immune response (TNF). Moreover, all 11 variants in 10 genes showed positive associations with NAFLD, and the highest risk factor for NAFLD was PNPLA3 rs738409 (OR: 1.841, 95% CI: 1.691-2.004). The cumulative epidemiological evidence of a significant association was graded as strong for two variants in two genes (HFE, TNF), moderate for four variants in three genes (TM6SF2, GCKR, and ADIPOQ), and weak for five variants in five genes (MBOAT7, PEMT, PNPLA3, LEPR, and MTHFR), based on the Venice criteria.
Table 1. Results of Random Effects Meta-Analyses Using Allelic Contrasts for Polymorphisms.
Gene and function Variant Alleles MAF Number assessed Allelic contrasts Heterogeneity Venice criteria grade c Cumulative evidence of association d Studies Cases Controls OR (95% CI) a P value P value b I2 Lipid synthesis and metabolism related genes APOC3 rs2854116 T vs C 0.55 (T) 11 3792 4601 0.99 (0.89-1.10) 0.826 0.004 63 ACC + rs2854117 C vs T 0.67 (C) 9 3538 3819 1.02 (0.95-1.10) 0.577 0.370 8 AAC + LYPLAL1* rs12137855 T vs C 0.20 (T) 13 4369 5292 0.98 (0.89-1.09) 0.753 0.326 12 AAC + MBOAT7 rs641738 C vs T 0.57 (C) 12 4351 10,830 1.07 (1.00-1.14) 0.048 0.528 0 AAC + MTTP rs1800591 T vs G 0.15 (T) 11 1483 1490 0.89 (0.59-1.36) 0.592 < 0.001 88 ACC + PEMT rs7946 T vs C 0.70 (T) 10 1090 1390 1.51 (1.11-2.06) 0.008 < 0.001 73 ACA + PNPLA3 rs738409 G vs C 0.26 (G) 75 18,193 24,710 1.84 (1.69-2.00) < 0.001 < 0.001 85 ACA + PPARG rs1801282 G vs C 0.11 (G) 9 2108 2740 0.88 (0.70-1.10) 0.260 0.084 43 ABC + PPARGC1A* rs8192678 T vs C 0.33 (T) 6 726 1058 1.05 (0.78-1.41) 0.735 0.011 66 ACC + TM6SF2 rs58542926 T vs C 0.07 (T) 24 5499 12,677 1.69 (1.47-1.93) < 0.001 0.068 33 ABA ++ Insulin resistance and glucose metabolism related genes GCKR rs780094 T vs C 0.40 (T) 23 6401 9983 1.18 (1.12-1.26) < 0.001 0.030 39 ABA ++ rs1260326 T vs C 0.40 (T) 9 1655 2527 1.48 (1.67-1.87) 0.001 < 0.001 46 ABA ++ PPP1R3B* rs4240624 G vs A 0.11(G) 7 2362 3292 0.88 (0.68-1.14) 0.339 0.367 8 AAC + Adipokines/adipokine receptors related genes ADIPOQ rs1501299 T vs G 0.27 (T) 13 2261 2190 0.97 (0.74-1.29) 0.849 < 0.001 88 ACC + rs266729 G vs C 0.23 (G) 8 1875 1466 1.61 (1.36-1.91) < 0.001 0.142 40 ABA ++ rs2241766 G vs T 0.10 (G) 13 2187 2072 1.09 (0.92-1.28) 0.335 0.012 54 ACC + LEPR rs1137100 G vs A 0.27 (G) 7 1382 1304 1.02 (0.83-1.26) 0.863 0.131 39 ABC + rs1137101 G vs A 0.46 (A) 6 1591 1535 1.82 (1.41-2.54) < 0.001 0.019 63 ACA + Oxidative stress and antioxidants related genes HFE rs1800562 A vs G 0.05 (A) 15 2261 5508 1.83 (0.99-3.40) 0.056 < 0.001 63 ACA + rs1799945 G vs C 0.14 (G) 14 1993 2475 1.24 (1.04-1.48) 0.019 0.192 24 AAA +++ MTHFR rs1801131 C vs A 0.30 (C) 7 2122 1157 1.24 (0.93-1.65) 0.141 < 0.001 76 ACA + rs1801133 T vs C 0.34 (T) 9 2303 1399 1.30 (1.06-1.59) 0.012 0.006 63 ACA + Inflammatory and immune response related genes TNF rs3615525 A vs G 0.05 (A) 14 2055 1594 1.82 (1.42-2.34) < 0.001 0.219 23 AAA +++ rs1800629 A vs G 0.15 (A) 13 2176 1789 1.29 (0.99-1.69) 0.064 0.115 34 ABA ++ Other functional genes NCAN* rs2228603 T vs C 0.07 (T) 11 4251 6105 0.97 (0.86-1.08) 0.544 0.712 0 AAC + Note. OR: odds ratio; CI: confidence interval; G: guanine; A, adenine; C: cytosine; T: thymine; MAF: minor-allele frequency. * Genes and loci that have not been meta-analyzed in published literature; a Summary ORs are based on random-effects allelic contrasts comparing minor and major alleles (based on frequencies in the control samples); b Based on the Q statistic across crude ORs calculated for each study; c Degree of ‘epidemiological credibility’ based on the interim Venice guidelines (A, strong; B, modest; C, weak); d Cumulative epidemiological evidence as graded by Venice criteria as strong (+++), moderate (++), or weak (+) for association with NAFLD risk. Of the 25 variants in meta-analyses, seven variants had little or no inter-study heterogeneity (I2 < 25%), 7 variants showed moderate heterogeneity (25% ≤ I2 ≤ 50%), and 11 variants showed high inter-study heterogeneity (I2 > 50%). The results of the dominant, recessive, heterozygous, and homozygous genetic models are shown in Supplemental Table S6 (available in www.besjournal.com). The allele contrast model identified more significant associations than the other models did.
All meta-analyses included a population of mixed ethnicities, except for LEPR rs1137100 and LEPR rs1137101, for which data were available only among Asian adults. Stratified meta-analyses according to ethnicity were conducted for the five variants of the four genes (Table 2). We found that PNPLA3 rs738409 was associated with NAFLD in all ethnicities, but showed the highest OR value (OR: 2.45, 95% CI: 2.04–2.95) in Caucasians and the lowest OR value (OR: 1.66, 95% CI: 1.51–1.83) in Asians. The three variants showed significant associations in only one ethnic population in the stratified analysis. Despite significant associations with NAFLD in the total population, significant associations were observed for GCKR rs780094 in Asians, GCKR rs1260326 in Caucasians, and MTHFR rs1801133 in Asians.
Table 2. Results of Random Effects Meta-Analyses of Alleles Using Allelic Contrasts for Polymorphisms Stratified by Ethnicity.
Gene and function Variant Alleles Subgroup Number assessed Allelic contrasts Heterogeneity Venice criteria grade c Cumulative evidence of association d Studies Cases Controls OR (95% CI) a P value P value b I2 Lipid synthesis and metabolism related genes MBOAT7 rs641738 C vs T Total 12 4351 10,830 1.07 (1.00-1.14) 0.048 0.528 0 AAC + Caucasians 7 2464 2166 1.07 (0.98-1.16) 0.144 0.448 0 AAC + Asians 5 1887 8664 1.07 (0.97-1.19) 0.188 0.374 6 AAC + PNPLA3 rs738409 G vs C Total 75 18,193 24,710 1.84 (1.69-2.00) < 0.001 < 0.001 85 ACA + Caucasians 21 4144 3327 2.45 (2.04-2.95) < 0.001 < 0.001 77 ACA + Asians 45 12,792 20,320 1.66 (1.51-1.83) < 0.001 < 0.001 86 ACA + Others 4 742 299 1.89 (1.52-2.37) < 0.001 0.961 0 AAA +++ Insulin resistance and glucose metabolism related genes GCKR rs780094 T vs C Total 23 6401 9983 1.18 (1.12-1.26) < 0.001 0.030 39 ABA ++ Asians 21 5603 9456 1.17 (1.10-1.26) < 0.001 0.040 38 ABA ++ Caucasians 2 798 527 1.34 (0.91-1.98) 0.142 0.065 71 ABC + rs1260326 T vs C Total 9 1655 2527 1.48 (1.67-1.87) 0.001 < 0.001 46 ABA +++ Asians 4 1023 992 1.61 (0.99-2.62) 0.057 < 0.001 93 ACA + Caucasians 3 393 552 1.42 (1.17-1.71) < 0.001 0.776 0 BAA ++ Others 2 239 983 1.33 (0.74-2.36) 0.338 0.038 77 ACA + Oxidative stress and antioxidants related genes MTHFR rs1801133 T vs C Total 9 2303 1399 1.30 (1.06-1.59) 0.012 0.006 63 ACA + Caucasians 4 1663 714 1.20 (0.85-1.68) 0.296 0.013 72 ACA + Asians 3 471 506 1.51 (1.07-2.12) 0.019 0.077 61 BCA + Hispanics 2 169 179 1.28 (0.63-2.58) 0.499 0.058 72 BCA + Note. OR: odds ratio; G: guanine; C: cytosine; T: thymine. a Summary ORs are based on random-effects allelic contrasts comparing minor and major alleles (based on frequencies in the control samples); b Based on the Q statistic across crude ORs calculated for each study; c Degree of ‘epidemiological credibility’ based on the interim Venice guidelines (A, strong; B, modest; C, weak); d Cumulative epidemiological evidence as graded by Venice criteria as strong (+++), moderate (++), or weak (+) for association with NAFLD risk. -
By 30 September, 2022, a total of 75 meta-analyses on NAFLD genetic associations were published involving 21 variants of 13 genes. The results of meta-analyses that were published most recently or included the largest number of studies were extracted for each variant (Table 3). Compared with previously published meta-analyses, meta-analyses for four variants in four genes were conducted for the first time in this study, and all four variants showed no association with NAFLD risk (LYPLAL1 rs12137855, PPARGC1A rs8192678, PPP1R3B rs4240624, NCAN rs2228603). Among the variants that have been meta-analyzed in the past, consistent results were observed for 15 variants in 11 genes, where 10 variants in 9 genes were associated with increased risk of NAFLD (TNF rs3615525, ADIPOQ rs266729, GCKR rs780094, GCKR rs1260326, PNPLA3 rs738409, MTHFR rs1801133, TM6SF2 rs58542926, PEMT rs7946, LEPR rs1137101, HFE rs1799945) and 5 variants in 4 genes (TNF rs1800629, PPARG rs1801282, APOC3 rs2854116, APOC3 rs2854117, LEPR rs1137100) showed no associations with NAFLD. Five variants in four genes (ADIPOQ rs1501299, ADIPOQ rs2241766, MTTP rs1800591, MTHFR rs1801131, HFE rs1800562) showed significant associations in previous meta-analyses but showed insignificant associations in this study. In contrast, MBOAT7 rs641738 was not associated with NAFLD in previous meta-analyses but was significantly associated with NAFLD in this study.
Table 3. Previously published meta-analyses results compared to meta-analyses in this study.
Gene Study Polymorphism Prior meta sample size: cases; controls (number of samples) Model Published meta OR (95% CI) Published Het. Meta Our new meta sample size: cases; controls (number of samples) Model New meta OR (95% CI) Het. Meta TNF Wang et al. 2011 rs3615525 771; 787 (7) GA/AA vs GG 2.06 (1.58-2.69) a 0.160 2055; 1594 (14) A vs G 1.82 (1.42-2.34) 0.219 rs1800629 837; 990 (8) GA/AA vs GG 1.08 (0.82-1.42) a 0.860 2176; 1789 (13 A vs G 1.29 (0.99-1.69) 0.115 PPARG Zhang et al. 2015 rs1801282 1697; 2427 (8) GC/GG vs CC 0.93 (0.63-1.38) <0.001 2108; 2740 (9) G vs C 0.88 (0.70-1.10) 0.084 ADIPOQ Wang et al. 2014 rs266729 876; 989 (7) GG/GC vs CC 1.52 (1.10-2.09) a 0.280 1875; 1466 (8) G vs C 1.61 (1.36-1.91) 0.142 Wang et al. 2016 rs1501299 1117; 1555 (10) G vs T 1.27 (1.10-1.48) a 0.533 2261; 2190 (13) T vs G 0.97 (0.74-1.29) <0.001 rs2241766 1117; 1555 (10) T vs G 1.33 (1.12-1.58) a 0.151 2187; 2072 (13) G vs T 1.09 (0.92-1.28) 0.012 APOC3 Li et al. 2017 rs2854116 2111; 1866 (9) C vs T 1.39 (0.96-2.02) a 0.001 3792; 4601 (11) T vs C 0.99 (0.89-1.10) 0.004 rs2854117 2111; 1866 (9) T vs C 1.05 (0.92-1.19) a 0.840 3538; 3819 (9) C vs T 1.02 (0.95-1.10) 0.370 GCKR Li et al. 2021 rs780094 5115; 11,812 (20) T vs C 1.20 (1.11-1.29) 0.020 6401; 9983 (23) T vs C 1.17 (1.10-1.24) <0.001 rs1260326 2238; 8995 (9) T vs C 1.32 (1.22-1.42) 0.560 1655; 2527 (9) T vs C 1.27 (1.14-1.42) <0.001 MTHFR Sun et al. 2016 rs1801131 364; 611 (5) C vs A 1.53 (1.13-2.07) 0.001 2122; 1157 (7) C vs A 1.24 (0.93-1.65) <0.001 rs1801133 737; 1160 (8) TT vs TC/CC 1.42 (1.07-1.88) 0.160 2303; 1399 (9) T vs C 1.30 (1.06-1.59) 0.006 MBOAT7 Xia et al. 2019 rs641738 2560; 8738 (5) C vs T 0.99 (0.93-1.05) a - 4351; 10,830 (12) C vs T 1.07 (1.00-1.14) 0.528 TM6SF2 Chen et al. 2019 rs58542926 3075; 3000 (13) Unknown 0.55 (0.48-0.63) - 5499; 12,677 (24) T vs C 1.69 (1.47-1.93) 0.068 PEMT Tan et al. 2016 rs7946 792; 2722 (6) TT/TC vs CC 1.62 (1.10-2.39) - 1090; 1390 (10) T vs C 1.51 (1.11-2.06) <0.001 LEPR Pan et al. 2018 rs1137100 1111; 1132 (6) A vs G 1.01 (0.87-1.18) 0.110 1382; 1304 (7) G vs A 1.02 (0.83-1.26) 0.131 rs1137101 1298; 1348 (5) A vs G 0.57 (0.50-0.65) 0.140 1591; 1535 (6) G vs A 1.82(1.41-2.54) 0.019 HFE Ye et al. 2016 rs1800562 1846; 7037 (11) A vs G 1.95 (1.16-3.28) <0.001 2261; 5508 (15) A vs G 1.83 (0.99-3.40) <0.001 rs1799945 3945; 12,332 (16) G vs C 1.21 (1.07-1.38) a 0.338 1993; 2475 (14) G vs C 1.24(1.04-1.48) 0.192 Note. a Fixed effects model was used in prior meta-analysis-: P-value is not reported. -
This study comprehensively reviewed the genetic association with NAFLD susceptibility by analyzing 381 candidate gene studies, 16 GWASs, and two WESs. 72 and 465 variants of over 100 genes related to multiple functional pathways were identified in the GWASs and candidate gene studies, respectively. We conducted meta-analyses of 25 variants and found significant associations with NAFLD risk for 11 variants, including six variants that showed moderate to strong cumulative epidemiological evidence for a true association. Additionally, meta-analyses provided convincing evidence of no association between NAFLD risk and 14 variants in 11 genes.
Our study identified 4 SNPs in 4 genes related to lipid synthesis and metabolism that were significantly associated with NAFLD risk. An association with TM6SF2 rs58542926 has been identified in several GWASs and a meta-analysis by Chen et al. (2019)[24]. The SNP TM6SF2 rs58542926 leads to a reduction in the expression and activity of TM6SF2, which normally promotes triglyceride secretion from hepatocytes[10,25]. The PNPLA3 rs738409 SNP causes a missense coding of methionine instead of isoleucine, leading to reduced lipidation of Apolipoprotein B100 and promotion of hepatic fat accumulation[26,27], and has been reported as a genetic risk factor for NAFLD in a large number of studies, including several GWASs[7-9] and previous meta-analyses[28,29]. MBOAT7 rs641738 was found to have borderline significance in this study, whereas a previous meta-analysis by Xia et al. (2019)[30] reported insignificant associations. The SNP rs641738 is believed to affect the acyl chain composition and remodeling of phospholipids in membranes[31]. The PEMT rs7946 variant leads to an amino acid substitution and reduced PEMT activity, causing the accumulation of triacylglycerols in the liver[32]. A significant association between PEMT rs7946 polymorphism and NAFLD has only been reported in Asians in a meta-analysis conducted by Tan et al. (2016)[33], and this association was confirmed in a mixed population in this study, although only weak cumulative epidemiological evidence was found.
Among insulin resistance and glucose metabolism-related genes, this study found moderate evidence of an association between two GCKR variants (rs780094 and rs1260326) and NAFLD risk, supported by previous GWASs and meta-analyses[34,35]. GCKR encodes a protein that regulates glucokinase activity[36], and the polymorphism rs1260326 leads to increased glucokinase activity and promotes hepatic glucose metabolism and lipid synthesis[37]. The intronic SNP rs780094 has been linked to fasting serum triacylglycerol, insulin, and type 2 diabetes risk[38] and is in strong linkage disequilibrium with rs1260326, resulting in reduced regulation of glucokinase activity by GCKR[39]. In addition, an epigenetic study suggested that rs780094 functions as a transcriptional enhancer of GCKR expression[40].
Two genes related to adipokines or adipokine receptors were identified in this study (ADIPOQ, LEPR), and moderate evidence was observed for ADIPOQ rs266729. Adiponectin, the protein product of the ADIPOQ gene, is an anti-inflammatory adipokine that functions as an insulin sensitizer and regulates energy homeostasis and glucose and lipid metabolism[41]. A GWAS found that the ADIPOQ gene was the major contributor to plasma adiponectin, accounting for 6.7% of its variation[42]. The rs266729 SNP in the ADIPOQ promoter region regulates adiponectin levels and promoter activity[43].
We also identified two genes related to oxidative stress and antioxidants (HFE, MTHFR) that were associated with NAFLD. HFE regulates cellular iron uptake[44] and the rs11799945 polymorphism is significantly linked to NAFLD with strong cumulative epidemiological evidence, which is consistent with a previous meta-analysis[14]. Additionally, HFE rs1800562 was only observed among Caucasians and was not detected in studies on Asian populations. Consistent with this, the online dbSNP database showed that the percentages of mutated A alleles reported in Caucasians and Asians were 0.059 and 0.0003, respectively. MTHFR rs1801133 showed a significant association with NAFLD; however, a meta-analysis of nine studies showed moderate heterogeneity. Consistent results for MTHFR rs1801133 polymorphism were reported in a meta-analysis by Sun et al. (2016)[45], although they also identified a significant association with MTHFR rs1801131, which was not significant in this study. Rs1801133 is a common missense variant (C677T) that results in a less active MTHFR enzyme[46,47] and is associated with a higher concentration of homocysteine, which may induce oxidative stress and endoplasmic reticulum stress[48].
We found that TNF rs361525 had strong cumulative evidence of an association with NAFLD, whereas TNF rs1800629 showed no significant association, consistent with a previous meta-analysis by Wang et al. (2012)[49]. TNF gene encodes tumor necrosis factor alpha (TNFα), a pro-inflammatory cytokine involved in the development of NAFLD[50]. However, the role of TNF rs361525 remains elusive, and mixed findings were reported regarding rs361525 and the level of TNFα in blood[51,52]. Further studies are required to elucidate the potential mechanisms for TNF polymorphisms in the development of NAFLD.
This study compared the results to those of previous meta-analyses and found consistency in 14 variants in 10 genes, and inconsistency in six variants in five genes. These differences may be attributed to the updated number of included studies and differences in the inclusion and exclusion criteria. For example, some previous studies contained data that did not conform to the Hardy-Weinberg equilibrium or included samples with other chronic liver diseases. In addition, the inconsistent results may also be a result of different selections of genetic models. In this meta-analysis, we used the results based on allele contrast models for comparison with previous studies.
To the best of our knowledge, this is the first study to comprehensively review studies analyzing genetic susceptibility in NAFLD and to extract data from GWASs, WES studies, and candidate gene studies. Potential susceptibility genes were categorized into eight categories based on their genetic functions, which may have facilitated the interpretation of the findings. Our study has certain limitations. First, although a thorough literature search was conducted using several strategies, it is likely that some publications were overlooked. Second, studies not published in English and publications without resolvable genotype counts were not included, which may have resulted in a publication bias. However, these studies probably represent a small percentage of all publications that investigated NAFLD-associated variants, and we did not detect significant publication bias in most meta-analyses. Third, the meta-analyses were conducted only for variants with a minimum of five data sources. Fourth, the use of genotype counts and crude estimates of effects precludes more sophisticated analyses, including adjusted estimates of association and gene-gene and gene-environment interactions, for which raw genotype data are required. Fifth, although we conducted a stratified analysis by ethnicity, we were unable to examine the potential sources of heterogeneity, which were limited by the number of studies available. Sixth, there was heterogeneity between the studies on the diagnostic methods used to define NAFLD. Finally, most of the positive variants in this study did not reach very high levels of statistical significance, and those with modest P values should be considered cautiously. Issues such as multiple testing, linkage disequilibrium among associated variants, and undetected publications or other reporting biases may mask the observed association.
In conclusion, this systematic review and meta-analysis provides a comprehensive summary of the genetic susceptibility to NAFLD. Recent GWASs and WES studies have identified 72 variants associated with the risk of NAFLD. Meta-analyses identified 11 genetic variants with strong or moderate evidence of association, and further study of these variants is required. We also identified 14 variants in 11 genes that showed no evidence of association with NAFLD risk. These findings provide valuable information for future studies to assess genetic factors associated with the risk of NAFLD.
-
Web of science:
TS=("non-alcoholic fatty liver disease" OR "nonalcoholic fatty liver disease" OR NAFLD OR "non-alcoholic fatty liver" OR "nonalcoholic fatty liver" OR NAFL OR "non-alcoholic steatohepatitis" OR "nonalcoholic steatohepatitis" OR NASH OR "hepatic steatosis") AND TS=(SNP* OR polymorphism* OR variant* OR variation* OR mutation*) AND language: (English OR Chinese) The index=SCI-EXPANDED, CPCI-S, ESCI timescale=1980-2022
Pubmed:
(("non-alcoholic fatty liver disease" OR "nonalcoholic fatty liver disease" OR NAFLD OR "non-alcoholic fatty liver" OR "nonalcoholic fatty liver" OR NAFL OR "non-alcoholic steatohepatitis" OR "nonalcoholic steatohepatitis" OR NASH OR "hepatic steatosis") AND (SNP* OR polymorphism* OR variant* OR variation* OR mutation*)) AND ((humans) AND (chinese OR english) AND (1980:2022))
Embase:('non-alcoholic fatty liver disease':ab,ti OR 'nonalcoholic fatty liver disease':ab,ti OR nafld:ab,ti OR 'non-alcoholic fatty liver':ab,ti OR 'nonalcoholic fatty liver':ab,ti OR nafl:ab,ti OR 'non-alcoholic steatohepatitis':ab,ti OR 'nonalcoholic steatohepatitis':ab,ti OR nash:ab,ti OR 'hepatic steatosis':ab,ti) AND (snp*:ab,ti OR polymorphism*:ab,ti OR variant*:ab,ti OR variation*:ab,ti OR mutation*:ab,ti) AND (/lim OR /lim) AND /lim AND /lim AND /py
doi: 10.3967/bes2024.079
Genetic Variations and Nonalcoholic Fatty Liver Disease: Field Synopsis, Systematic Meta-Analysis, and Epidemiological Evidence
-
Abstract:
Objective To systematically summarize the published literature on the genetic variants associated with nonalcoholic fatty liver disease (NAFLD). Methods Literature from Web of Science, PubMed, and Embase between January 1980 and September 2022 was systematically searched. Meta-analyses of the genetic variants were conducted using at least five data sources. The epidemiologic credibility of the significant associations was graded using the Venice criteria. Results Based on literature screening, 399 eligible studies were included, comprising 381 candidate gene association, 16 genome-wide association, and 2 whole-exome sequencing studies. We identified 465 genetic variants in 173 genes in candidate gene association studies, and 25 genetic variants in 17 genes were included in the meta-analysis. The meta-analysis identified 11 variants in 10 genes that were significantly associated with NAFLD, with cumulative epidemiological evidence of an association graded as strong for two variants in two genes (HFE, TNF), moderate for four variants in three genes (TM6SF2, GCKR, and ADIPOQ), and weak for five variants in five genes (MBOAT7, PEMT, PNPLA3, LEPR, and MTHFR). Conclusions This study identified six variants in five genes that had moderate to strong evidence of an association with NAFLD, which may help understand the genetic architecture of NAFLD risk. -
Key words:
- Nonalcoholic fatty liver disease /
- Genetic association study /
- Genetic variant /
- Systematic review /
- Meta-analysis
&These authors contributed equally to this work.
注释:1) AUTHOR CONTRIBUTIONS: -
Table 1. Results of Random Effects Meta-Analyses Using Allelic Contrasts for Polymorphisms.
Gene and function Variant Alleles MAF Number assessed Allelic contrasts Heterogeneity Venice criteria grade c Cumulative evidence of association d Studies Cases Controls OR (95% CI) a P value P value b I2 Lipid synthesis and metabolism related genes APOC3 rs2854116 T vs C 0.55 (T) 11 3792 4601 0.99 (0.89-1.10) 0.826 0.004 63 ACC + rs2854117 C vs T 0.67 (C) 9 3538 3819 1.02 (0.95-1.10) 0.577 0.370 8 AAC + LYPLAL1* rs12137855 T vs C 0.20 (T) 13 4369 5292 0.98 (0.89-1.09) 0.753 0.326 12 AAC + MBOAT7 rs641738 C vs T 0.57 (C) 12 4351 10,830 1.07 (1.00-1.14) 0.048 0.528 0 AAC + MTTP rs1800591 T vs G 0.15 (T) 11 1483 1490 0.89 (0.59-1.36) 0.592 < 0.001 88 ACC + PEMT rs7946 T vs C 0.70 (T) 10 1090 1390 1.51 (1.11-2.06) 0.008 < 0.001 73 ACA + PNPLA3 rs738409 G vs C 0.26 (G) 75 18,193 24,710 1.84 (1.69-2.00) < 0.001 < 0.001 85 ACA + PPARG rs1801282 G vs C 0.11 (G) 9 2108 2740 0.88 (0.70-1.10) 0.260 0.084 43 ABC + PPARGC1A* rs8192678 T vs C 0.33 (T) 6 726 1058 1.05 (0.78-1.41) 0.735 0.011 66 ACC + TM6SF2 rs58542926 T vs C 0.07 (T) 24 5499 12,677 1.69 (1.47-1.93) < 0.001 0.068 33 ABA ++ Insulin resistance and glucose metabolism related genes GCKR rs780094 T vs C 0.40 (T) 23 6401 9983 1.18 (1.12-1.26) < 0.001 0.030 39 ABA ++ rs1260326 T vs C 0.40 (T) 9 1655 2527 1.48 (1.67-1.87) 0.001 < 0.001 46 ABA ++ PPP1R3B* rs4240624 G vs A 0.11(G) 7 2362 3292 0.88 (0.68-1.14) 0.339 0.367 8 AAC + Adipokines/adipokine receptors related genes ADIPOQ rs1501299 T vs G 0.27 (T) 13 2261 2190 0.97 (0.74-1.29) 0.849 < 0.001 88 ACC + rs266729 G vs C 0.23 (G) 8 1875 1466 1.61 (1.36-1.91) < 0.001 0.142 40 ABA ++ rs2241766 G vs T 0.10 (G) 13 2187 2072 1.09 (0.92-1.28) 0.335 0.012 54 ACC + LEPR rs1137100 G vs A 0.27 (G) 7 1382 1304 1.02 (0.83-1.26) 0.863 0.131 39 ABC + rs1137101 G vs A 0.46 (A) 6 1591 1535 1.82 (1.41-2.54) < 0.001 0.019 63 ACA + Oxidative stress and antioxidants related genes HFE rs1800562 A vs G 0.05 (A) 15 2261 5508 1.83 (0.99-3.40) 0.056 < 0.001 63 ACA + rs1799945 G vs C 0.14 (G) 14 1993 2475 1.24 (1.04-1.48) 0.019 0.192 24 AAA +++ MTHFR rs1801131 C vs A 0.30 (C) 7 2122 1157 1.24 (0.93-1.65) 0.141 < 0.001 76 ACA + rs1801133 T vs C 0.34 (T) 9 2303 1399 1.30 (1.06-1.59) 0.012 0.006 63 ACA + Inflammatory and immune response related genes TNF rs3615525 A vs G 0.05 (A) 14 2055 1594 1.82 (1.42-2.34) < 0.001 0.219 23 AAA +++ rs1800629 A vs G 0.15 (A) 13 2176 1789 1.29 (0.99-1.69) 0.064 0.115 34 ABA ++ Other functional genes NCAN* rs2228603 T vs C 0.07 (T) 11 4251 6105 0.97 (0.86-1.08) 0.544 0.712 0 AAC + Note. OR: odds ratio; CI: confidence interval; G: guanine; A, adenine; C: cytosine; T: thymine; MAF: minor-allele frequency. * Genes and loci that have not been meta-analyzed in published literature; a Summary ORs are based on random-effects allelic contrasts comparing minor and major alleles (based on frequencies in the control samples); b Based on the Q statistic across crude ORs calculated for each study; c Degree of ‘epidemiological credibility’ based on the interim Venice guidelines (A, strong; B, modest; C, weak); d Cumulative epidemiological evidence as graded by Venice criteria as strong (+++), moderate (++), or weak (+) for association with NAFLD risk. Table 2. Results of Random Effects Meta-Analyses of Alleles Using Allelic Contrasts for Polymorphisms Stratified by Ethnicity.
Gene and function Variant Alleles Subgroup Number assessed Allelic contrasts Heterogeneity Venice criteria grade c Cumulative evidence of association d Studies Cases Controls OR (95% CI) a P value P value b I2 Lipid synthesis and metabolism related genes MBOAT7 rs641738 C vs T Total 12 4351 10,830 1.07 (1.00-1.14) 0.048 0.528 0 AAC + Caucasians 7 2464 2166 1.07 (0.98-1.16) 0.144 0.448 0 AAC + Asians 5 1887 8664 1.07 (0.97-1.19) 0.188 0.374 6 AAC + PNPLA3 rs738409 G vs C Total 75 18,193 24,710 1.84 (1.69-2.00) < 0.001 < 0.001 85 ACA + Caucasians 21 4144 3327 2.45 (2.04-2.95) < 0.001 < 0.001 77 ACA + Asians 45 12,792 20,320 1.66 (1.51-1.83) < 0.001 < 0.001 86 ACA + Others 4 742 299 1.89 (1.52-2.37) < 0.001 0.961 0 AAA +++ Insulin resistance and glucose metabolism related genes GCKR rs780094 T vs C Total 23 6401 9983 1.18 (1.12-1.26) < 0.001 0.030 39 ABA ++ Asians 21 5603 9456 1.17 (1.10-1.26) < 0.001 0.040 38 ABA ++ Caucasians 2 798 527 1.34 (0.91-1.98) 0.142 0.065 71 ABC + rs1260326 T vs C Total 9 1655 2527 1.48 (1.67-1.87) 0.001 < 0.001 46 ABA +++ Asians 4 1023 992 1.61 (0.99-2.62) 0.057 < 0.001 93 ACA + Caucasians 3 393 552 1.42 (1.17-1.71) < 0.001 0.776 0 BAA ++ Others 2 239 983 1.33 (0.74-2.36) 0.338 0.038 77 ACA + Oxidative stress and antioxidants related genes MTHFR rs1801133 T vs C Total 9 2303 1399 1.30 (1.06-1.59) 0.012 0.006 63 ACA + Caucasians 4 1663 714 1.20 (0.85-1.68) 0.296 0.013 72 ACA + Asians 3 471 506 1.51 (1.07-2.12) 0.019 0.077 61 BCA + Hispanics 2 169 179 1.28 (0.63-2.58) 0.499 0.058 72 BCA + Note. OR: odds ratio; G: guanine; C: cytosine; T: thymine. a Summary ORs are based on random-effects allelic contrasts comparing minor and major alleles (based on frequencies in the control samples); b Based on the Q statistic across crude ORs calculated for each study; c Degree of ‘epidemiological credibility’ based on the interim Venice guidelines (A, strong; B, modest; C, weak); d Cumulative epidemiological evidence as graded by Venice criteria as strong (+++), moderate (++), or weak (+) for association with NAFLD risk. Table 3. Previously published meta-analyses results compared to meta-analyses in this study.
Gene Study Polymorphism Prior meta sample size: cases; controls (number of samples) Model Published meta OR (95% CI) Published Het. Meta Our new meta sample size: cases; controls (number of samples) Model New meta OR (95% CI) Het. Meta TNF Wang et al. 2011 rs3615525 771; 787 (7) GA/AA vs GG 2.06 (1.58-2.69) a 0.160 2055; 1594 (14) A vs G 1.82 (1.42-2.34) 0.219 rs1800629 837; 990 (8) GA/AA vs GG 1.08 (0.82-1.42) a 0.860 2176; 1789 (13 A vs G 1.29 (0.99-1.69) 0.115 PPARG Zhang et al. 2015 rs1801282 1697; 2427 (8) GC/GG vs CC 0.93 (0.63-1.38) <0.001 2108; 2740 (9) G vs C 0.88 (0.70-1.10) 0.084 ADIPOQ Wang et al. 2014 rs266729 876; 989 (7) GG/GC vs CC 1.52 (1.10-2.09) a 0.280 1875; 1466 (8) G vs C 1.61 (1.36-1.91) 0.142 Wang et al. 2016 rs1501299 1117; 1555 (10) G vs T 1.27 (1.10-1.48) a 0.533 2261; 2190 (13) T vs G 0.97 (0.74-1.29) <0.001 rs2241766 1117; 1555 (10) T vs G 1.33 (1.12-1.58) a 0.151 2187; 2072 (13) G vs T 1.09 (0.92-1.28) 0.012 APOC3 Li et al. 2017 rs2854116 2111; 1866 (9) C vs T 1.39 (0.96-2.02) a 0.001 3792; 4601 (11) T vs C 0.99 (0.89-1.10) 0.004 rs2854117 2111; 1866 (9) T vs C 1.05 (0.92-1.19) a 0.840 3538; 3819 (9) C vs T 1.02 (0.95-1.10) 0.370 GCKR Li et al. 2021 rs780094 5115; 11,812 (20) T vs C 1.20 (1.11-1.29) 0.020 6401; 9983 (23) T vs C 1.17 (1.10-1.24) <0.001 rs1260326 2238; 8995 (9) T vs C 1.32 (1.22-1.42) 0.560 1655; 2527 (9) T vs C 1.27 (1.14-1.42) <0.001 MTHFR Sun et al. 2016 rs1801131 364; 611 (5) C vs A 1.53 (1.13-2.07) 0.001 2122; 1157 (7) C vs A 1.24 (0.93-1.65) <0.001 rs1801133 737; 1160 (8) TT vs TC/CC 1.42 (1.07-1.88) 0.160 2303; 1399 (9) T vs C 1.30 (1.06-1.59) 0.006 MBOAT7 Xia et al. 2019 rs641738 2560; 8738 (5) C vs T 0.99 (0.93-1.05) a - 4351; 10,830 (12) C vs T 1.07 (1.00-1.14) 0.528 TM6SF2 Chen et al. 2019 rs58542926 3075; 3000 (13) Unknown 0.55 (0.48-0.63) - 5499; 12,677 (24) T vs C 1.69 (1.47-1.93) 0.068 PEMT Tan et al. 2016 rs7946 792; 2722 (6) TT/TC vs CC 1.62 (1.10-2.39) - 1090; 1390 (10) T vs C 1.51 (1.11-2.06) <0.001 LEPR Pan et al. 2018 rs1137100 1111; 1132 (6) A vs G 1.01 (0.87-1.18) 0.110 1382; 1304 (7) G vs A 1.02 (0.83-1.26) 0.131 rs1137101 1298; 1348 (5) A vs G 0.57 (0.50-0.65) 0.140 1591; 1535 (6) G vs A 1.82(1.41-2.54) 0.019 HFE Ye et al. 2016 rs1800562 1846; 7037 (11) A vs G 1.95 (1.16-3.28) <0.001 2261; 5508 (15) A vs G 1.83 (0.99-3.40) <0.001 rs1799945 3945; 12,332 (16) G vs C 1.21 (1.07-1.38) a 0.338 1993; 2475 (14) G vs C 1.24(1.04-1.48) 0.192 Note. a Fixed effects model was used in prior meta-analysis-: P-value is not reported. -
[1] Manikat R, Ahmed A, Kim D. Up-to-date global epidemiology of nonalcoholic fatty liver disease. Hepatobiliary Surg Nutr, 2023; 12, 956−9. doi: 10.21037/hbsn-23-548 [2] Feng G, Valenti L, Wong VWS, et al. Recompensation in cirrhosis: unravelling the evolving natural history of nonalcoholic fatty liver disease. Nat Rev Gastroenterol Hepatol, 2024; 21, 46−56. doi: 10.1038/s41575-023-00846-4 [3] Younossi ZM, Wong G, Anstee QM, et al. The global burden of liver disease. Clin Gastroenterol Hepatol, 2023; 21, 1978−91. doi: 10.1016/j.cgh.2023.04.015 [4] Liu L, Shao YH, Feng EQ, et al. Risk of developing non-alcoholic fatty liver disease over time in a cohort of the elderly in Qingdao, China. Biomed Environ Sci, 2023; 36, 760−7. [5] Zhao H, Qiu X, Li HZ, et al. Association between serum uric acid to HDL-cholesterol ratio and nonalcoholic fatty liver disease risk among Chinese adults. Biomed Environ Sci, 2023; 36, 1−9. [6] Pourteymour S, Drevon CA, Dalen KT, et al. Mechanisms behind NAFLD: a system genetics perspective. Curr Atheroscler Rep, 2023; 25, 869−78. doi: 10.1007/s11883-023-01158-3 [7] Romeo S, Kozlitina J, Xing C, et al. Genetic variation in PNPLA3 confers susceptibility to nonalcoholic fatty liver disease. Nat Genet, 2008; 40, 1461−5. doi: 10.1038/ng.257 [8] Speliotes EK, Yerges-Armstrong LM, Wu J, et al. Genome-wide association analysis identifies variants associated with nonalcoholic fatty liver disease that have distinct effects on metabolic traits. PLoS Genet, 2011; 7, e1001324. doi: 10.1371/journal.pgen.1001324 [9] Kitamoto T, Kitamoto A, Yoneda M, et al. Genome-wide scan revealed that polymorphisms in the PNPLA3, SAMM50, and PARVB genes are associated with development and progression of nonalcoholic fatty liver disease in Japan. Hum Genet, 2013; 132, 783−92. doi: 10.1007/s00439-013-1294-3 [10] Kozlitina J, Smagris E, Stender S, et al. Exome-wide association study identifies a TM6SF2 variant that confers susceptibility to nonalcoholic fatty liver disease. Nat Genet, 2014; 46, 352−6. doi: 10.1038/ng.2901 [11] Chen W, Coombes BJ, Larson NB. Recent advances and challenges of rare variant association analysis in the biobank sequencing era. Front Genet, 2022; 13, 1014947. doi: 10.3389/fgene.2022.1014947 [12] Lee YH. Meta-analysis of genetic association studies. Ann Lab Med, 2015; 35, 283−7. doi: 10.3343/alm.2015.35.3.283 [13] Sookoian S, Pirola CJ. Meta-analysis of the influence of I148M variant of patatin-like phospholipase domain containing 3 gene (PNPLA3) on the susceptibility and histological severity of nonalcoholic fatty liver disease. Hepatology, 2011; 53, 1883−94. doi: 10.1002/hep.24283 [14] Ye Q, Qian BX, Yin WL, et al. Association between the HFE C282Y, H63D polymorphisms and the risks of non-alcoholic fatty liver disease, liver cirrhosis and hepatocellular carcinoma: an updated systematic review and meta-analysis of 5, 758 cases and 14, 741 controls. PLoS One, 2016; 11, e0163423. doi: 10.1371/journal.pone.0163423 [15] Teo K, Abeysekera KWM, Adams L, et al. rs641738C>T near MBOAT7 is associated with liver fat, ALT and fibrosis in NAFLD: a meta-analysis. J Hepatol, 2021; 74, 20−30. doi: 10.1016/j.jhep.2020.08.027 [16] Ioannidis JPA, Boffetta P, Little J, et al. Assessment of cumulative evidence on genetic associations: interim guidelines. Int J Epidemiol, 2008; 37, 120−32. doi: 10.1093/ije/dym159 [17] The International HapMap Consortium. A second generation human haplotype map of over 3.1 million SNPs. Nature, 2007; 449, 851−61. doi: 10.1038/nature06258 [18] Stang A. Critical evaluation of the Newcastle-Ottawa scale for the assessment of the quality of nonrandomized studies in meta-analyses. Eur J Epidemiol, 2010; 25, 603−5. doi: 10.1007/s10654-010-9491-z [19] Zintzaras E, Lau J. Synthesis of genetic association studies for pertinent gene-disease associations requires appropriate methodological and statistical approaches. J Clin Epidemiol, 2008; 61, 634−45. doi: 10.1016/j.jclinepi.2007.12.011 [20] DerSimonian R, Laird N. Meta-analysis in clinical trials revisited. Contemp Clin Trials, 2015; 45, 139−45. doi: 10.1016/j.cct.2015.09.002 [21] Whitehead A, Whitehead J. A general parametric approach to the meta-analysis of randomized clinical trials. Stat Med, 1991; 10, 1665−77. doi: 10.1002/sim.4780101105 [22] Higgins JPT, Thompson SG. Quantifying heterogeneity in a meta-analysis. Stat Med, 2002; 21, 1539−58. doi: 10.1002/sim.1186 [23] Egger M, Davey Smith G, Schneider M, et al. Bias in meta-analysis detected by a simple, graphical test. BMJ, 1997; 315, 629−34. doi: 10.1136/bmj.315.7109.629 [24] Chen XP, Zhou PC, De L, et al. The roles of transmembrane 6 superfamily member 2 rs58542926 polymorphism in chronic liver disease: a meta-analysis of 24, 147 subjects. Mol Genet Genomic Med, 2019; 7, e824. doi: 10.1002/mgg3.824 [25] Mahdessian H, Taxiarchis A, Popov S, et al. TM6SF2 is a regulator of liver fat metabolism influencing triglyceride secretion and hepatic lipid droplet content. Proc Natl Acad Sci USA, 2014; 111, 8913−8. doi: 10.1073/pnas.1323785111 [26] He SQ, McPhaul C, Li JZ, et al. A sequence variation (I148M) in PNPLA3 associated with nonalcoholic fatty liver disease disrupts triglyceride hydrolysis. J Biol Chem, 2010; 285, 6706−15. doi: 10.1074/jbc.M109.064501 [27] Kumari M, Schoiswohl G, Chitraju C, et al. Adiponutrin functions as a nutritionally regulated lysophosphatidic acid acyltransferase. Cell Metab, 2012; 15, 691−702. doi: 10.1016/j.cmet.2012.04.008 [28] Lu FB, Hu ED, Xu LM, et al. The relationship between obesity and the severity of non-alcoholic fatty liver disease: systematic review and meta-analysis. Expert Rev Gastroenterol Hepatol, 2018; 12, 491−502. doi: 10.1080/17474124.2018.1460202 [29] Wu PB, Shu YX, Guo F, et al. Association between patatin-like phospholipase domain-containing protein 3 gene rs738409 polymorphism and non-alcoholic fatty liver disease susceptibility: a Meta-analysis. Chin J Epidemiol, 2015; 36, 78−82. (In Chinese [30] Xia Y, Huang CX, Li GY, et al. Meta-analysis of the association between MBOAT7 rs641738, TM6SF2 rs58542926 and nonalcoholic fatty liver disease susceptibility. Clin Res Hepatol Gastroenterol, 2019; 43, 533−41. doi: 10.1016/j.clinre.2019.01.008 [31] Caddeo A, Jamialahmadi O, Solinas G, et al. MBOAT7 is anchored to endomembranes by six transmembrane domains. J Struct Biol, 2019; 206, 349−60. doi: 10.1016/j.jsb.2019.04.006 [32] Song JN, Da Costa KA, Fischer LM, et al. Polymorphism of the PEMT gene and susceptibility to nonalcoholic fatty liver disease (NAFLD). FASEB J, 2005; 19, 1266−71. doi: 10.1096/fj.04-3580com [33] Tan HL, Mohamed R, Mohamed Z, et al. Phosphatidylethanolamine N-methyltransferase gene rs7946 polymorphism plays a role in risk of nonalcoholic fatty liver disease: evidence from meta-analysis. Pharmacogenet Genomics, 2016; 26, 88−95. doi: 10.1097/FPC.0000000000000193 [34] Zain SM, Mohamed Z, Mohamed R. A common variant in the glucokinase regulatory gene rs780094 and risk of nonalcoholic fatty liver disease: a meta-analysis. J Gastroenterol Hepatol, 2015; 30, 21−7. doi: 10.1111/jgh.12714 [35] Cai W, Weng DH, Yan P, et al. Genetic polymorphisms associated with nonalcoholic fatty liver disease in Uyghur population: a case-control study and meta-analysis. Lipids Health Dis, 2019; 18, 14. doi: 10.1186/s12944-018-0877-3 [36] Hayward BE, Dunlop N, Intody S, et al. Organization of the human glucokinase regulator geneGCKR. Genomics, 1998; 49, 137−42. doi: 10.1006/geno.1997.5195 [37] Beer NL, Tribble ND, McCulloch LJ, et al. The P446L variant in GCKR associated with fasting plasma glucose and triglyceride levels exerts its effect through increased glucokinase activity in liver. Hum Mol Genet, 2009; 18, 4081−8. doi: 10.1093/hmg/ddp357 [38] Sparsø T, Andersen G, Nielsen T, et al. The GCKR rs780094 polymorphism is associated with elevated fasting serum triacylglycerol, reduced fasting and OGTT-related insulinaemia, and reduced risk of type 2 diabetes. Diabetologia, 2008; 51, 70−5. [39] Tan HL, Zain SM, Mohamed R, et al. Association of glucokinase regulatory gene polymorphisms with risk and severity of non-alcoholic fatty liver disease: an interaction study with adiponutrin gene. J Gastroenterol, 2014; 49, 1056−64. doi: 10.1007/s00535-013-0850-x [40] López Rodríguez M, Kaminska D, Lappalainen K, et al. Identification and characterization of a FOXA2-regulated transcriptional enhancer at a type 2 diabetes intronic locus that controls GCKR expression in liver cells. Genome Med, 2017; 9, 63. doi: 10.1186/s13073-017-0453-x [41] Trujillo ME, Scherer PE. Adiponectin--journey from an adipocyte secretory protein to biomarker of the metabolic syndrome. J Intern Med, 2005; 257, 167-75. [42] Heid IM, Henneman P, Hicks A, et al. Clear detection of ADIPOQ locus as the major gene for plasma adiponectin: results of genome-wide association analyses including 4659 European individuals. Atherosclerosis, 2010; 208, 412−20. doi: 10.1016/j.atherosclerosis.2009.11.035 [43] Gu HF. Biomarkers of adiponectin: plasma protein variation and genomic DNA polymorphisms. Biomark Insights, 2009; 4, 123−33. [44] Bennett MJ, Lebrón JA, Bjorkman PJ. Crystal structure of the hereditary haemochromatosis protein HFE complexed with transferrin receptor. Nature, 2000; 403, 46−53. doi: 10.1038/47417 [45] Sun MY, Zhang L, Shi SL, et al. Associations between methylenetetrahydrofolate reductase (MTHFR) polymorphisms and Non-Alcoholic Fatty Liver Disease (NAFLD) risk: a meta-analysis. PLoS One, 2016; 11, e0154337. doi: 10.1371/journal.pone.0154337 [46] Guenther BD, Sheppard CA, Tran P, et al. The structure and properties of methylenetetrahydrofolate reductase from Escherichia coli suggest how folate ameliorates human hyperhomocysteinemia. Nat Struct Biol, 1999; 6, 359−65. doi: 10.1038/7594 [47] Van Der Put NMJ, Gabreëls F, Stevens EMB, et al. A second common mutation in the methylenetetrahydrofolate reductase gene: an additional risk factor for neural-tube defects? Am J Hum Genet, 1998; 62, 1044-51. [48] Santilli F, Davì G, Patrono C. Homocysteine, methylenetetrahydrofolate reductase, folate status and atherothrombosis: a mechanistic and clinical perspective. Vascul Pharmacol, 2016; 78, 1−9. doi: 10.1016/j.vph.2015.06.009 [49] Wang JK, Feng ZW, Li YC, et al. Association of tumor necrosis factor-α gene promoter polymorphism at sites -308 and -238 with non-alcoholic fatty liver disease: a meta-analysis. J Gastroenterol Hepatol, 2012; 27, 670−6. doi: 10.1111/j.1440-1746.2011.06978.x [50] Hajeer AH, Hutchinson IV. Influence of TNFα gene polymorphisms on TNFα production and disease. Hum Immunol, 2001; 62, 1191−9. doi: 10.1016/S0198-8859(01)00322-6 [51] Wilson AG, Symons JA, McDowell TL, et al. Effects of a polymorphism in the human tumor necrosis factor α promoter on transcriptional activation. Proc Natl Acad Sci USA, 1997; 94, 3195−9. doi: 10.1073/pnas.94.7.3195 [52] Wong VWS, Wong GLH, Tsang SWC, et al. Genetic polymorphisms of adiponectin and tumor necrosis factor-alpha and nonalcoholic fatty liver disease in Chinese people. J Gastroenterol Hepatol, 2008; 23, 914−21. doi: 10.1111/j.1440-1746.2008.05344.x -




 下载:
下载:



 Quick Links
Quick Links