-
Amyotrophic lateral sclerosis (ALS) is a fatal neurodegenerative disorder characterized by progressive degeneration of upper and lower motor neurons. Although the etiology and pathogenesis of ALS remain unclear, genetic factors are believed to play a significant role. The apolipoprotein E (APOE) genotype, a major genetic risk factor in Alzheimer’s disease (AD)[1-3], has also been implicated in other neurodegenerative conditions, including Parkinson’s disease[4,5], Lewy body dementia[6,7], and frontotemporal dementia[8]. In humans, the APOE gene has three common alleles (ε2, ε3, and ε4), with the ε4 allele conferring an increased risk for AD and the ε2 allele exhibiting a protective effect[9].
Given this background, the potential influence of APOE on ALS has been explored in numerous association studies over the past two decades; however, the findings have been inconsistent. While most reports indicate that the APOE genotype does not affect the susceptibility to ALS[10-13], in contrast to its established role in AD, emerging evidence suggests that it acts as a disease modifier. For instance, some studies have linked the ε4 allele to an earlier age of onset[14], bulbar-onset disease[15,16], and shorter survival[17]. In contrast, the ε2 allele has been associated with a later age of onset[18,19]. Ethnic and regional variations appear to contribute to this discrepancy: a study in China reported a significantly higher frequency of ε4 in patients with ALS than in controls[20], and a Tunisian cohort observed not only an elevated APOE-ε4 frequency among ALS cases—particularly those with bulbar onset—but also a significant correlation between the ε4 allele and the presence of cognitive deficits, especially dysexecutive functions[21]. Furthermore, Maranzano et al. demonstrated that APOE haplotypes, together with CSF Aβ biomarkers, are associated with cognitive impairment, particularly memory deficits, in patients with ALS[22]. Although one study found no association between APOE genotypes and clinical parameters, it identified a strong correlation between elevated plasma APOE levels and an accelerated rate of disease progression[23].
Therefore, the role of APOE polymorphisms as potential disease modifiers in ALS remains unclear and requires further investigation. To clarify this relationship, we evaluated the impact of APOE genotypes and serum APOE levels on cognitive characteristics and clinical phenotypes in a cohort of Chinese patients with sporadic ALS.
-
This was a single-center, retrospective, observational cohort study. A total of 368 patients with sporadic ALS classified as definite, probable, probable laboratory-supported, or possible according to the Revised El Escorial diagnostic criteria[24] were consecutively enrolled from the Department of Neurology, Chinese PLA General Hospital, between December 2011 and November 2021. APOE genotyping was successfully performed in 289 patients, and serum APOE levels were quantified in a subset of 222 participants. The unavailability of APOE genotyping (n = 79) or serum data (n = 146) in some patients was due to the absence of blood sampling for these specific tests during routine clinical visits, because these analyses were not part of the mandatory clinical protocol. This occurrence was random in the clinical setting. For each patient, the following clinical parameters were collected: sex, site of symptom onset (limb or bulbar), age at onset, and time to generalization (TTG), defined as the interval from initial spinal or bulbar symptom onset to generalized involvement, which serves as an early clinical marker of disease progression[25]. Functional disability was assessed using the ALS Functional Rating Scale-Revised (ALSFRS-R)[26]. The disease progression rate (DPR) was calculated as follows: [(48 − ALSFRS-R score at evaluation) / (disease duration from symptom onset to evaluation in months)][27]. Cognitive and behavioral assessments were conducted using the Edinburgh Cognitive and Behavioural ALS Screen (ECAS)-Chinese version[28], which evaluates multiple domains, including language, verbal fluency, executive functions, memory, and visuospatial abilities. The total ECAS score comprises ALS-specific (language, fluency, and executive functions) and ALS-nonspecific domains (memory and visuospatial functions).
-
Genomic DNA was extracted using a MagNA Pure 96 magnetic bead system (Roche, Basel, Switzerland). APOE genotypes were determined by real-time quantitative polymerase chain reaction (qPCR) using the Hongshi real-time PCR system (Hongshi, Shanghai, China). Participants were categorized based on APOE alleles: individuals with genotypes APOE2/3 or APOE2/4 were classified as ε2 carriers; those with APOE3/4 or APOE4/4 as ε4 carriers; and those with APOE3/3 as ε3 carriers. No APOE2/2 genotypes were identified in this cohort. Serum APOE concentrations were quantified by immunonephelometry using a Cobas 8000 automated analyzer (Roche Diagnostics, Mannheim, Germany).
-
Normality and homogeneity of variance for continuous variables were assessed using the Kolmogorov–Smirnov and Levene’s tests, respectively. Data are presented as mean ± standard deviation for normally distributed variables and median (interquartile range) for non-normal variables. Categorical variables (e.g., sex and site of onset) were compared across APOE genotype groups using the chi-square test. The Kruskal–Wallis test or Wilcoxon rank-sum test was applied to evaluate the effect of the APOE genotype on non-normally distributed continuous variables (DPR, TTG, and ECAS scores). For normally distributed variables (age of onset), one-way analysis of variance was used. To further examine allele-specific effects, patients were grouped by the presence or absence of specific APOE alleles (ε2, ε3, ε4). Intergroup comparisons of non-normal variables (DPR, TTG) were conducted using the Mann–Whitney U test, and the age of onset was compared using Student’s t-test. Associations between serum APOE levels and clinical parameters in patients with sporadic ALS were evaluated using Spearman’s correlation analysis. Differences in ECAS scores across APOE allele carrier status (ε2 vs. non-ε2, ε3 vs. non-ε3, ε4 vs. non-ε4) were examined through partial correlation analysis, with adjustments for age and years of education. The analysis testing the primary hypothesis—that the APOE ε4 allele is associated with cognitive impairment—was performed using partial correlation, controlling for age and years of education. This was a prespecified, focused analysis, and its P-value was reported without further multiplicity correction. All statistical analyses were conducted using SPSS (version 22.0; IBM Corp., Armonk, NY, USA), with a significance threshold of P < 0.05.
-
A total of 368 participants were included in this study. Among them, APOE genotyping was successfully performed in 289 individuals. The distribution of APOE genotypes within the cohort is summarized in Table 1.
Characteristic Genotypes Alleles ε2-ε3 ε2-ε4 ε3-ε3 ε3-ε4 ε4-ε4 Total ε2 ε3 ε4 Total n 22 5 215 46 1 289 27 498 53 578 Frequency (%) 7.6 1.7 74.4 15.9 0.3 100 4.6 86.2 9.2 100 Note. APOE: apolipoprotein E. Table 1. Genotype and allele frequencies of APOE gene in sporadic amyotrophic lateral sclerosis
-
No significant differences were observed in the distribution of APOE genotypes with respect to sex or the site of symptom onset. Furthermore, age at onset, DPR, and TTG showed no association with the APOE genotype (classified as 2/3 and 2/4 vs. 3/3 vs. 3/4 and 4/4) (Table 2). Similarly, comparisons between all carriers of a specific allele and non-carriers (e.g., E2 carriers vs. non-E2 carriers) also revealed no statistically significant associations, which is consistent with the results presented in Table 3.
Characteristic APOE genotypes ε2 carriers (2/3, 2/4) (n = 27) ε3 carriers (3/3) (n =215) ε4 carriers (3/4, 4/4) (n =47) P-value Gender (Male/female) 16/11 134/81 32/15 0.696 Site of onset (Bulbar/limb) 3/24 41/174 8/39 0.587 Age of onset (years) (n) 50.61 ± 11.42 (27) 53.20 ± 10.09 (215) 51.34 ± 10.46 (47) 0.296 Rate of disease progression (n) 0.73 (0.46–1.22) (9) 0.85 (0.34–1.39) (76) 0.57 (0.40–1.25) (19) 0.797 TTG (months) (n) 10.00 (5.00–15.00) (13) 9.00 (5.00–16.00) (101) 11.00 (6.00–18.00) (25) 0.832 Note. APOE: apolipoprotein E; TTG: time to generalization (interval from symptom onset to generalized involvement). Data for age at onset are presented as mean ± standard deviation. Data for disease progression rate and TTG are presented as median (interquartile range). Table 2. Clinical characteristics of ALS patients by APOE genotype group
APOE allele carrier status Age of onset (years) (n) Rate of disease progression (n) TTG (months) (n) ε2 carriers (n = 27) 50.61 ± 11.42 (27) 0.73 (0.46–1.22) (9) 10.00 (5.00–15.00) (13) Non-ε2 carriers (n = 262) 52.87 ± 10.17 (262) 0.75 (0.37–1.33) (95) 10.00 (5.00–16.00) (126) P-value 0.279 0.954 0.871 ε3 carriers (n = 283 ) 52.55 ± 10.20 (283) 0.75 (0.37–1.33) (99) 10.00 (5.00–16.00) (135) Non-ε3 carriers (n = 6) 57.89 ± 13.74 (6) 0.73 (0.41–1.29) (5) 15.00 (6.00–20.25) (4) P-value 0.209 0.945 0.476 ε4 carriers (n = 52) 51.72 ± 10.71 (52) 0.58 (0.40–1.25) (23) 12.00 (6.00–18.00) (29) Non-ε4 carriers (n = 237) 52.86 ± 10.20 (237) 0.78 (0.35–1.36) (81) 9.00 (5.00–14.50) (110) P-value 0.469 0.652 0.389 Note. APOE: apolipoprotein E; TTG: time to generalization (interval from symptom onset to generalized involvement). Data for age at onset are presented as mean ± standard deviation. Data for disease progression rate and TTG are presented as median (interquartile range). Table 3. Clinical characteristics of ALS patients by APOE allele carrier status
-
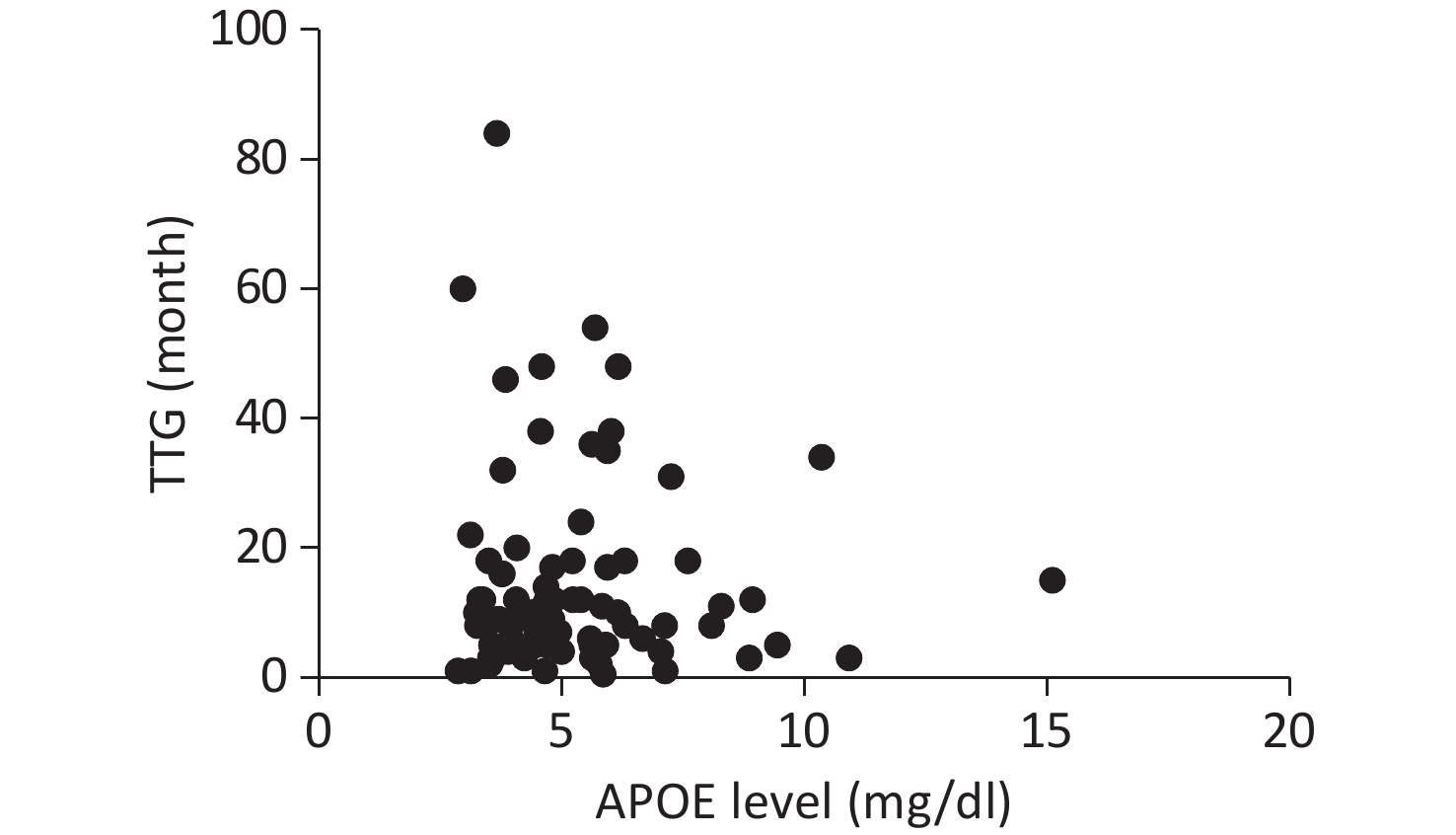
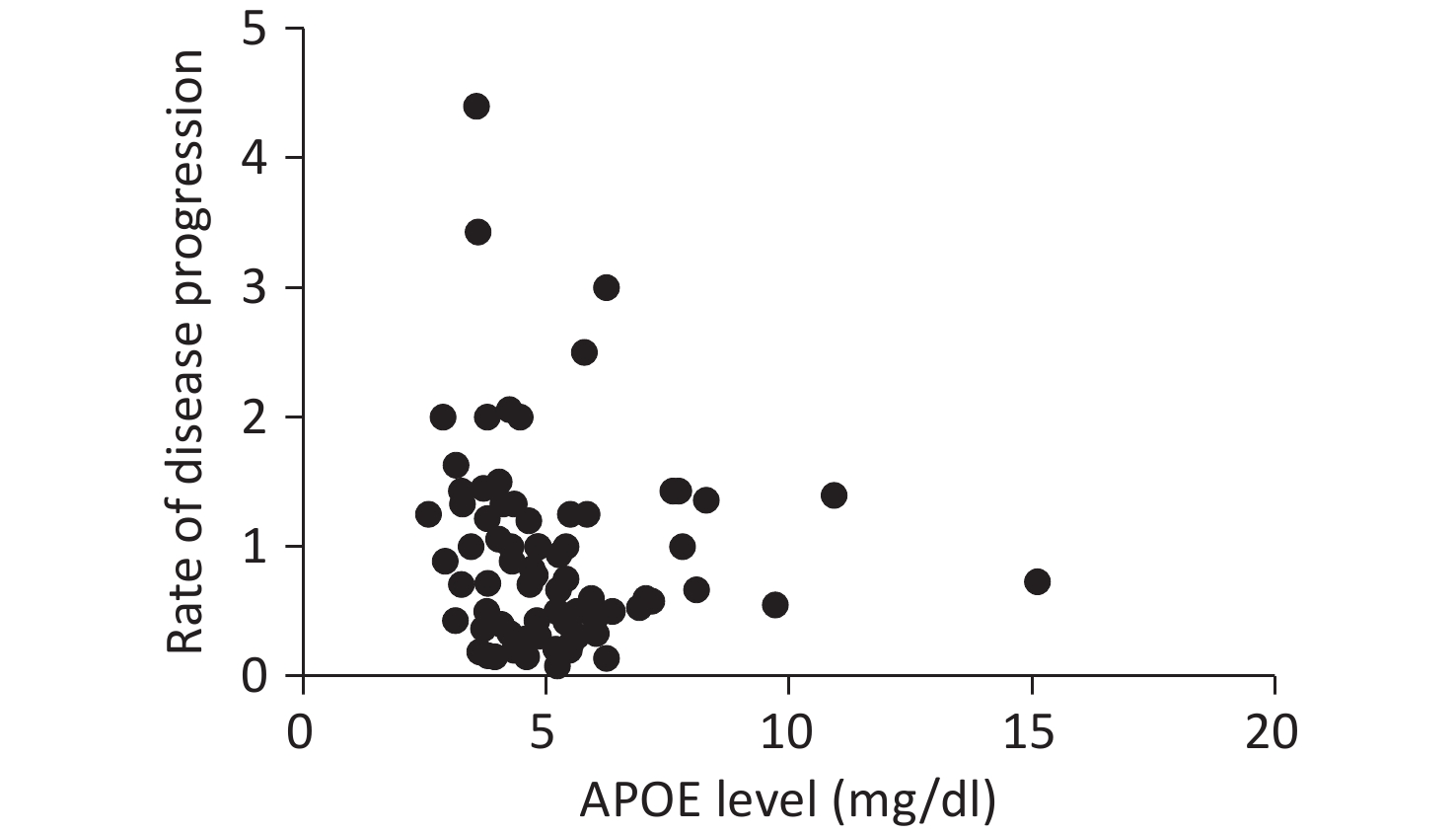
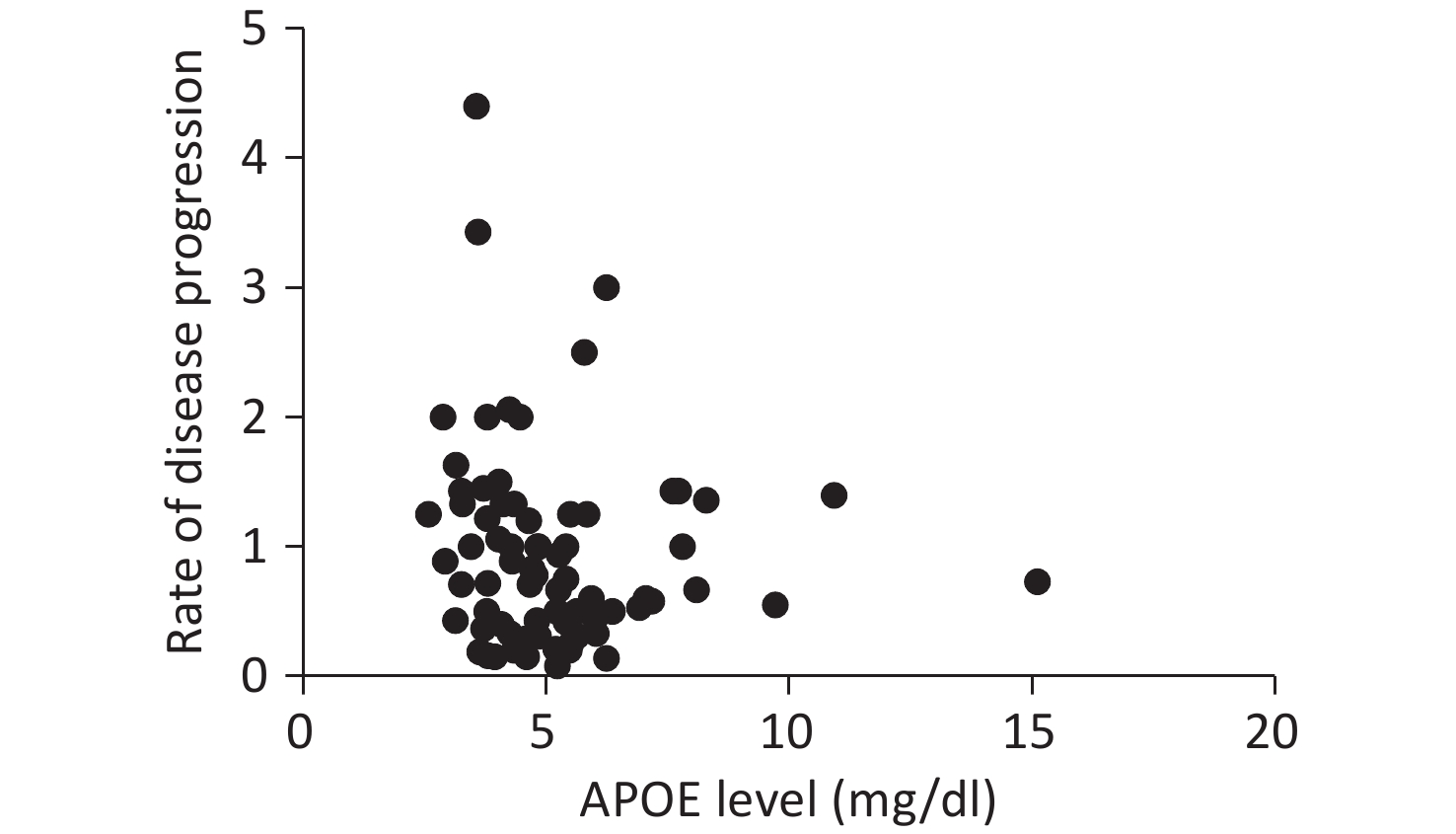
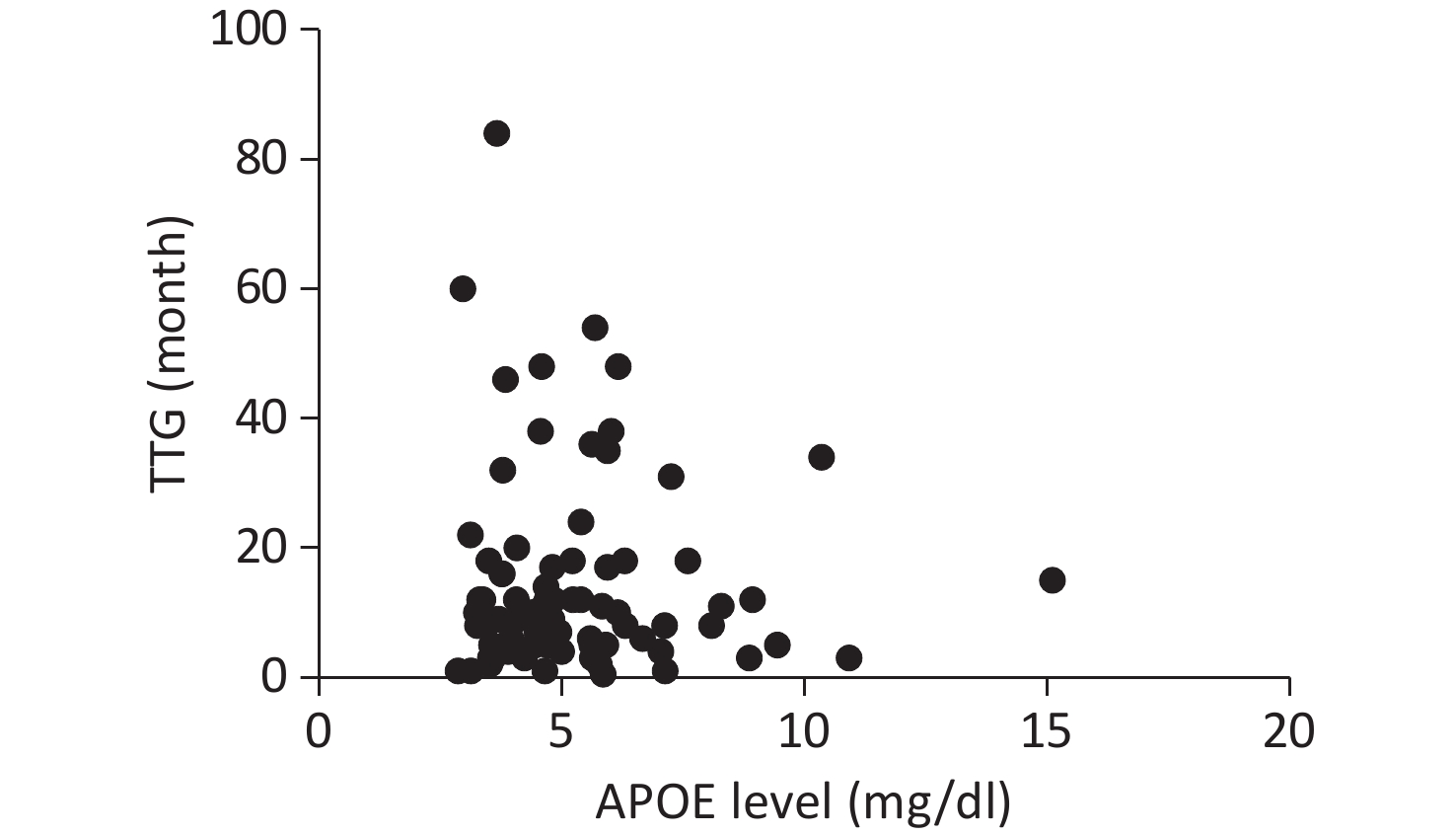
Serum APOE levels were measured in 222 patients. Complete clinical data, including ALSFRS-R scores, DPR, and TTG, were available for 107 participants. No significant associations were observed between APOE serum levels and either DPR or TTG, as illustrated in Figures 1 and 2.
-
Cognitive function was assessed using the ECAS in a subset of 78 patients in the overall cohort. No carriers of E2/E4 or E4/E4 genotypes were identified in this group. The allele frequency distributions were as follows: E2/E3 (14/78, 17.9%), E3/E3 (51/78, 65.4%), and E3/E4 (13/78, 16.7%). After adjustment for age and education level, known covariates of the cognitive performance, partial correlation analysis revealed no significant association between the APOE genotypes (E2/E3, E3/E3, or E3/E4) and either the ECAS total score or its subdomain scores (language, verbal fluency, executive function, ALS-specific, memory, visuospatial, and ALS-nonspecific functions) (Table 4).
Domain APOE genotypes ε2 carriers (ε2/ε3) (n = 14) ε3/ε3 (n = 51) ε4 carriers (ε3/ε4) (n = 13) P-value Language 22 (19–25) 23 (19–25) 23 (19–26.5) 0.875 Fluency 20 (13–20) 20 (16–22) 18 (4–21) 0.324 Executive 42.5 (38.5–44.25) 42 (35–44) 40 (29.5–47.5) 0.098 ALS specific score 80.5 (77–87) 82 (75–90) 78 (57.5–92) 0.179 Memory 16.5 (12–18) 14 (10–17) 14 (6–20) 0.249 Visuospatial 12 (12–12) 12 (12–12) 12 (11.5–12) 0.205 ALS non-specific score 28 (23.5–30) 26 (22–29) 26 (17.5–32) 0.199 ECAS total score 109 (100–115) 106 (97–118) 111 (76.5–119) 0.145 Note. APOE: apolipoprotein E; ALS: amyotrophic lateral sclerosis; ECAS: Edinburgh Cognitive and Behavioural ALS Screen. Data are presented as median (interquartile range). Table 4. Cognitive performance assessed by ECAS in ALS patients by APOE genotype group
However, when comparing carriers with non-carriers of specific alleles, individuals carrying at least one E4 allele exhibited significantly lower median scores for verbal fluency (P = 0.015) and ALS-specific function (P = 0.037) than non-carriers. Furthermore, the E4 allele showed a borderline association with poorer performance in visuospatial function (P = 0.060) and total ECAS scores (P = 0.051), suggesting that E4 carriers exhibit more pronounced cognitive deficits. In contrast, no significant association was observed between the E2 allele and any domain of ECAS (Tables 5 and 6).
APOE allele carrier status Language Fluency Executive ALS specific score ε2 carriers > (n =14 ) 22 (19-25) 20 (13-20) 42.5 (38.5-44.25) 80.5 (77-87) Non-ε2 carriers > (n = 64) 23 (19-25) 20 (14.5-22) 42 (32.5-44.75) 81.5 (69-91) P-value 0.667 0.426 0.370 0.960 ε3 carriers > (n = 51) 23 (19-25) 20 (16-22) 42 (35-44) 82 (75-90) Non-ε3 carriers > (n =27 ) 22 (19-25) 20 (10-20) 42 (35-45) 80 (67-90) P-value 0.617 0.011 0.563 0.114 ε4 carriers > (n =13) 23 (19-26.5) 18 (4-21) 40 (29.5-47.5) 78 (57.5-92) Non-ε4 carriers > (n =65 ) 22 (19-25) 20 (16-22) 42 (36-44) 81 (75-89) P-value 0.845 0.015 0.093 0.037 Note. APOE: apolipoprotein E; ALS, amyotrophic lateral sclerosis; ECAS, Edinburgh Cognitive and Behavioural ALS Screen. Data are presented as median (interquartile range). Bold P-values indicate statistical significance (P < 0.05). Table 5. Comparison of ECAS domain scores (language, fluency, executive function, and ALS-specific) by APOE allele carrier status
APOE allele carrier status Memory Visuospatial ALS non-specific score ECAS total score ε2 carriers (n =14 ) 16.5 (12-18) 12 (12-12) 28 (23.5-30) 109 (100-115) Non-ε2 carriers (n = 64) 14 (9.25-17.75) 12 (12-12) 26 (21.25-29.75) 106.5 (91.5-118) P-value 0.315 0.896 0.347 0.727 ε3 carriers (n =51 ) 14 (10-17) 12 (12-12) 26 (22-29) 106 (97-118) Non-ε3 carriers (n =27 ) 16 (12-18) 12 (12-12) 28 (22-30) 109 (88-117) P-value 0.841 0.177 0.951 0.220 ε4 carriers (n =13) 14 (6-20) 12 (11.5-12) 26 (17.5-32) 111 (76.5-119) Non-ε4 carriers (n =65 ) 15 (11.5-17.5) 12 (12-12) 27 (22.5-29.5) 109 (100-118) P-value 0.436 0.060 0.293 0.051 Note. APOE: apolipoprotein E; ALS: amyotrophic lateral sclerosis; ECAS: Edinburgh Cognitive and Behavioural ALS Screen. Data are presented as median (interquartile range). Table 6. Comparison of ECAS domain scores (memory, visuospatial function, ALS-nonspecific, and total) by APOE allele carrier status
-
Our analysis revealed no significant differences in APOE genotype distribution based on sex or site of disease onset in ALS. Furthermore, no associations were detected between APOE genetic profiles (grouped as 2/3 and 2/4 vs. 3/3 vs. 3/4 and 4/4) and clinical features such as age at onset, DPR, or TTG. Consistent results were obtained when comparing individual APOE alleles against non-carriers (e.g., ε2 carriers vs. non-ε2 carriers). Similarly, serum APOE levels showed no correlation with disease characteristics, including age at onset, TTG, or DPR. In contrast, carriers of the APOE ε4 allele exhibited significantly greater cognitive impairment compared to non-carriers, with particularly notable deficits in verbal fluency. This observation is consistent with previous studies that reported the adverse cognitive effects associated with the ε4 allele[21,22]. The allele frequencies in our sporadic ALS cohort (ε2, 4.6%; ε3, 86.2%; and ε4, 9.2%) are broadly comparable to those reported in healthy Chinese controls from large cerebrovascular studies (ε2, 16.7%; ε3, 71.1%; and ε4, 12.3%)[29].
Several large-cohort studies have evaluated the association between the APOE genotype and ALS. However, studies measuring blood APOE levels in patients with ALS remain scarce. Lacomblez et al. reported a significant correlation between APOE plasma levels and both disease progression and outcomes in a cohort of 403 patients with ALS; however, they found no association between these clinical parameters and the APOE phenotype[23]. No correlation was observed between serum APOE concentrations and the rate of deterioration in our study. In contrast to serum APOE, other serum components, such as trace elements, have shown significant associations with disease progression in Chinese patients with ALS[30]. In addition to the differences in the blood samples tested, the relatively small number of patients included in the current study may have caused these conflicting results. Nonetheless, blood APOE concentration does not appear to be a highly sensitive biomarker for ALS disease risk, in contrast to its established role in AD, where peripheral APOE is recognized as a risk factor[31], potentially mediated through oxidative stress mechanisms[32]. Further investigation is warranted to elucidate the potential role of peripheral APOE in the pathophysiology of ALS.
As previously documented, multiple studies have investigated the potential association between APOE genotype and ALS, with some reporting a significantly elevated risk of ALS linked to specific APOE variants, whereas others have reported contradictory findings. However, much of the literature supporting such an association originates from earlier preliminary studies. A comprehensive meta-analysis of all available studies on the role of APOE in ALS—including 4,249 patients and 10,397 controls—concluded that none of the APOE alleles significantly influenced the risk of ALS. Specifically, the ε4 allele was not associated with a statistically significant increase in susceptibility to ALS[10]. Additionally, a large-scale study involving 504 patients demonstrated that the presence of the APOE ε2 allele significantly increased the risk of frontotemporal dementia[8]. This finding was further corroborated by another investigation utilizing cerebral 18F-FDG-PET, which identified ε2 as a risk factor for cognitive impairment in ALS[33]. Therefore, we hypothesized that APOE alleles exert a stronger influence on cognitive function than on the clinical progression of motor symptoms in ALS. The present study further substantiated the association between APOE polymorphisms and cognitive performance. Specifically, the E4 haplotype remained significantly correlated with ECAS scores, even after adjusting for covariates affecting the cognitive profiles. This result is consistent with reports by Maranzano et al.[21,22], which indicated that carrying at least one APOE-E4 allele was associated with reduced levels of Aβ42/40 and Aβ42, as well as more severe cognitive impairment—particularly in memory domains—in nonspecific ALS presentations. These findings support the role of APOE as a major genetic modulator of cognitive decline via Aβ-dependent pathways and strengthen the evidence linking APOE polymorphisms to dementia in ALS. Some biological mechanisms may explain our findings. The APOE ε4 allele could interact with core ALS pathologies, such as TDP-43 aggregation, worsening cortical neurodegeneration. It can also impair glial cell function, increase neuroinflammation, and reduce neuronal support. These pathways are likely to reduce neuronal resilience, making patients more vulnerable to cognitive decline. Further studies using biomarker data are required to confirm these mechanisms.
This study had several limitations. First, the sample size for cognitive assessment was modest (n = 78), and the cognitive subgroup lacked certain genotypes (e.g., E4/E4), which may limit the interpretation of a full APOE ε4 dosage effect. Second, its cross-sectional design precludes causal inferences. Third, owing to the retrospective nature of the study, we could not systematically exclude patients with potential confounders of serum APOE levels, such as hyperlipidemia or diabetes. The influence of these metabolic conditions on serum APOE concentrations, which may affect the interpretability of serum APOE-related findings, cannot be ruled out. Furthermore, discrepancies between our findings and those of other populations may be attributable to differences in genetic background, environmental factors, and clinical heterogeneity. Future studies should (1) recruit larger, multicenter cohorts to enhance statistical power; (2) incorporate longitudinal cognitive assessments; and (3) investigate the underlying molecular mechanisms (e.g., CSF biomarkers) linking APOE ε4 to cognitive decline in ALS. Despite these limitations, our study provides preliminary evidence linking the APOE ε4 allele to cognitive impairment in Chinese patients with sporadic ALS.
In conclusion, our study provides initial evidence linking the APOE ε4 allele to cognitive impairment in Chinese patients with ALS. These findings further reinforce the hypothesis that APOE genotype plays a role in the etiology of ALS.
HTML
Patients and Clinical Parameters
APOE Genotyping and Serum Level Measurement
Statistical Analysis
Demographic Characteristics of the Participants
Associations between APOE Genotype and ALS Clinical Parameters
Associations between APOE Serum Levels and ALS Clinical Parameters
Association of APOE Genotype with Cognitive Performance
Competing Interests The authors declare no conflicts of interest.
Ethics This study was approved by the Ethics Committee of the First Medical Center of Chinese PLA General Hospital (No. S2022-119) and was conducted in accordance with the Declaration of Helsinki. Written informed consent was obtained from all the recruited patients.
Authors’ Contributions Conception, design and execution of the study, and writing of the manuscript: Qionghua Sun; Analysis of data and execution of the study: Xuan Xuan; Review and editing: Yuguo Du and Yucui Zhai; Summarized the materials and curated the data: Tie Ma, Feng Duan, Cuiqiao Xia; Provided guidance for the entire study: Xusheng Huang; In charge of the project and responsible for the project, design, and checking of the article: Yonghua Huang and Hongfen Wang.







 Quick Links
Quick Links
 DownLoad:
DownLoad: