-
Hand, foot and mouth disease (HFMD) is a common infectious disease that usually affects children less than 5 years of age. HFMD is caused by human enteroviruses (HEVs). HEVs, members of the Enterovirus genus of the Picornaviridae (small RNA virus) family, were traditionally classified into Poliovirus (PV), Echovirus (Echo), Coxsackievirus A and B (CVA and B) and new HEVs. However, since 1999, HEVs have been divided into four groups depending on their molecular, biological, and genetic characteristics. The four groups are enterovirus A, B, C, and D[1]. To date, more than 100 HEV serotypes have been reported. The major pathogens that cause HFMD include Enterovirus 71 (EV71) and Coxsackievirus A16 (CVA16). In addition, other enteroviruses account for a large proportion of the pathogenic spectrum of HFMD. In recent years, the number of HFMD cases and epidemic events caused by Coxsackievirus A6 and A10 (CVA6 and CVA10)[2] have increased worldwide. However, there have been relatively few reports of HFMD-related coxsackievirus B (CVB1-5).
In May 2008, HFMD was listed as a class C notifiable disease in China. Since then, HFMD epidemics have been reported online directly. The web-based HFMD outbreak reporting system in Fujian Province showed that EV71 and CVA16 were the major pathogens causing HFMD in the Fujian Province between 2008 and 2010. However, the percentage of HFMD cases caused by other enteroviruses in the pathogenic spectrum increased from less than 5% before 2011, to 27% in 2011, and subsequently increased by 30% every year.
This study was conducted in the Fujian province of south-eastern China, from 2011 to 2016. Clinical specimens were obtained from HFMD patients who were infected with non-EV71 and non-CVA16 enteroviruses and the specimens were examined in the study. The corresponding epidemiological information was collected using laboratory-based HFMD surveillance system in accordance with the 'Guidelines to Prevention and Control of Hand, Foot and Mouth Disease'. Enterovirus isolation and identification was performed in accordance with the standards and procedures provided by the 'Laboratory Manual for Hand, Foot and Mouth Disease'. Molecular typing was performed and the acquired nucleotide sequence of the VP1 gene of CVB1-5 was subjected to RT-PCR using an Access One Step RT-PCR Kit (Promega Corporation, USA) with primers[3, 4]. Phylogenetic trees were constructed using the maximum likelihood method and the Mega 6.0 software. The full-length genome sequences of the virus were sequenced using Ion torrent S5 second-generation sequencing platform and spliced using the program that maps to reference in the CLC software. Recombination analysis was performed using Simplot 3.5.1 software.
Twenty-two cases of HFMD were identified to be caused by coxsackievirus B1-5 (CVB1-5). These cases accounted for 3.0% of the 733 HFMD cases that were caused by other enteroviruses circulating in the Fujian Province from 2011 to 2016. The 22 cases of CVB1-5 related HFMD included 13 males and 9 females (the male-to-female ratio was 1.4:1). Males appeared to be more susceptible to infection with CVB1-5, which is consistent with the results from the analysis of the population most affected by HFMD. The majority of the patients with CVB1-5 related HFMD (21/22) were under the age of 5, which is also consistent with results from the analysis of the population most affected by HFMD. These cases were first detected between April and July (20/22), suggesting that CVB1-5 is relatively active during this period, thus causing infections in individuals. The 22 HFMD cases consisted of 15 sporadic children and 7 kindergarten children and 21 ordinary cases and 1 severe case caused by CVB5. Previous studies also reported that infections with CVB5[5, 6] can result in severe clinical manifestations, including inflammation of the central nervous system, aseptic meningitis, encephalitis, and bronchial pneumonia. Based on the regional and temporal distribution of the cases, some types of CVB1-5 viruses were found to be more prevalent in a certain area at a certain time; for example, the CVB3 in the Nanping prefecture in 2012, the CVB4 in the Ningde prefecture in 2012, and the CVB5 in the Longyan prefecture in 2012. Specific information about these cases is shown in Table 1.
Strain Name Sex Age
(years)Date of Accident ClinicaI Outcome Cluster Patterns Regions Serotype GenBank Accession No. 2011FJZZ059 Female 1.22 2011/5/25 Mild Sporadic Zhangzhou CVB1 MG922510 2011FJQZ127 Male 4.24 2011/6/13 Mild Kindergarten Quanzhou CVB1 MG922509 2011FJQZ165 Male 1.44 2011/7/6 Mild Sporadic Quanzhou CVB1 MG922508 2011FJQZ162 Female 3.99 2011/7/4 Mild Sporadic Quanzhou CVB2 MG922511 2012FJND182 Female 6.00 2012/6/6 Mild Kindergarten Ningde CVB3 MG922517 2012FJZZ015 Male 3.85 2012/4/17 Mild Sporadic Zhangzhou CVB3 MG922518 2012FJNP065 Male 2.32 2011/11/15 Mild Kindergarten Nanping CVB3 MG922516 2012FJNP124 Female 1.25 2012/5/14 Mild Sporadic Nanping CVB3 MG922515 2012FJNP264 Female 0.22 2012/6/9 Mild Sporadic Nanping CVB3 MG922514 2012FJSM017 Male 0.69 2012/4/21 Mild Sporadic Sanming CVB3 MG922512 2012FJQZ063 Female 4.50 2012/4/15 Mild Kindergarten Quanzhou CVB3 MG922513 2012FJND241 Male 5.01 2012/7/9 Mild Sporadic Ningde CVB4 MG922522 2012FJND259 Male 1.67 2012/7/4 Mild Sporadic Ningde CVB4 MG922521 2012FJND260 Male 1.43 2012/7/12 Mild Sporadic Ningde CVB4 MG922520 2012FJZZ185 Female 2.13 2012/6/1 Mild Sporadic Zhangzhou CVB4 MG922523 2012FJSM058 Male 2.48 2012/4/18 Mild Kindergarten Sanming CVB4 MG922519 2011FJLY159 Male 2.99 2011/6/6 Mild Sporadic Longyan CVB5 MG922526 2011FJLY126 Female 2.00 2011/5/26 Mild Kindergarten Longyan CVB5 MG922525 2011FJLY096 Male 2.97 2011/5/8 Mild Kindergarten Longyan CVB5 MG922524 2012FJZZ125 Male 1.98 2012/5/25 Mild Sporadic Zhangzhou CVB5 MG922527 2014FJPTN026 Female 3.16 2014/5/1 Mild Sporadic Pingtan CVB5 MG922529 2014FJFZ015 Male 0.59 2013/12/5 Severe Sporadic Fuzhou CVB5 MG922528 Table 1. Epidemiological Information about CVB1-5 Related HFMD that Occurred in the Fujian Province between 2011 and 2016
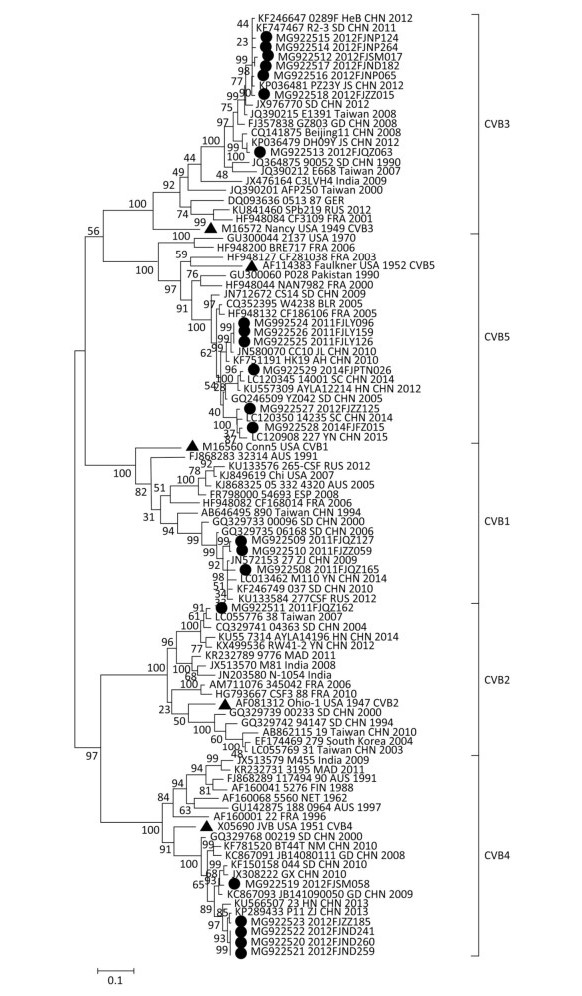
In this study, the 22 strains of CVB1-5 that were circulating between 2011 and 2016 were isolated, and the entire VP1 region of the isolates was amplified and sequenced. Compared to their corresponding prototype strains, the enteroviruses did not have any nucleotide deletions or insertions in the VP1 region. Sequence similarity analysis (Supplementary Table S1 available in www.besjournal.com) showed that there were low nucleotide and amino acid sequence similarities between the Fujian strains and their corresponding prototype strains from the same serotypes. The Fujian strains shared greater nucleotide and amino acid sequence similarities with national strains than with the international strains. The phylogenetic tree (Figure 1) based on the full sequence of the VP1 genomic region of Fujian CVB1-5 strains and corresponding prototype, national, and international strains (retrieved from GenBank) showed that virulent strains of the same serotype were clustered together. The Fujian CVB1-5 isolates had small genetic distances and close genetic relationships with each other and therefore exhibited a certain degree of clustering, while some of them also showed regional clustering. These Fujian isolates and their corresponding prototypes trains belonged to different clades. Large genetic distances and distant genetic relationships were detected between the Fujian isolates and corresponding prototype strains from the same serotype. Such results are consistent with the findings of recent domestic and foreign studies on the enteroviruses EV71, CVA16, and CVA6[7, 8], suggesting that enteroviruses have undergone more significant evolutionary changes during the recent years in comparison with the early prototype strains. Smaller genetic distances and closer genetic relationships existed between the Fujian isolates and national strains than those between Fujian isolates and the international strains. In addition, the fact that some Fujian isolates showed regional clustering on the phylogenetic tree was consistent with the epidemiological information of the specific Fujian isolate, further confirming that some types of CVB1-5 viruses were prevalent in a certain area during a certain time.

Table Supplementary Table 1. Analysis of the VP1 Genomic Region of Various Serotypes of CVB1-5 Circulating in Fujian Province during 2011-2016

Figure 1. A phylogenetic dendrogram of the complete VP1 gene sequences of CVB1-5 circulating in Fujian between 2011 and 2016, constructed using the maximum likelihood method. ▲ indicates prototype strains; ● indicates Fujian strains.
Full genomes for each of the CVB1-5 serotypes in Fujian (CVB1-2011FJQZ127, CVB2-2011FJQZ162, CVB3-2012FJZZ015, CVB4-2012FJSM058, and CVB5-2014FJFZ015) were obtained. The full genome length for the five strains (CVB1-2011FJQZ127, CVB2-2011FJQZ162, CVB3-2012FJZZ015, CVB4-2012FJSM058, and CVB5-2014FJFZ015) are 7, 378 nt, 7, 405 nt, 7, 395 nt, 7, 401 nt, and 7, 332 nt, respectively. The open reading frame (ORF) of the five strains are 6, 546 nt, 6, 561 nt, 6, 555 nt, 6, 549 nt, and 6, 555 nt, respectively and encode a polypeptide containing 2, 182, 2, 187, 2, 185, 2, 183, and 2, 185 amino acids, respectively. A comprehensive comparison of the similarities in nucleotide acid sequence between the Fujian CVB1-5 viruses and the corresponding prototype strains and other HEV-B prototype viruses is shown in Supplementary Table S2 (available in www.besjournal.com). In the VP1, VP2, and VP3 capsid protein, the Fujian CVB1-5 strains exhibits higher nucleotide similarity with the corresponding prototype strains than with the prototype strains from different serotypes; in the P2 and P3 regions, the Fujian CVB1-5 strains show greater similarity in identity with some prototype strains from different serotypes than with their corresponding prototype strains. These results suggest that the molecular typing of the enterovirus based on VP1 and P1 gene is reliable, and recombination possibly occurs in the non-capsid regions.

Table Supplementary Table 2. Pair-wise Nucleotide Acid Sequence Identities Between the Fujian CVB1-5 Strains and Prototype Strains of the HEV-B Species
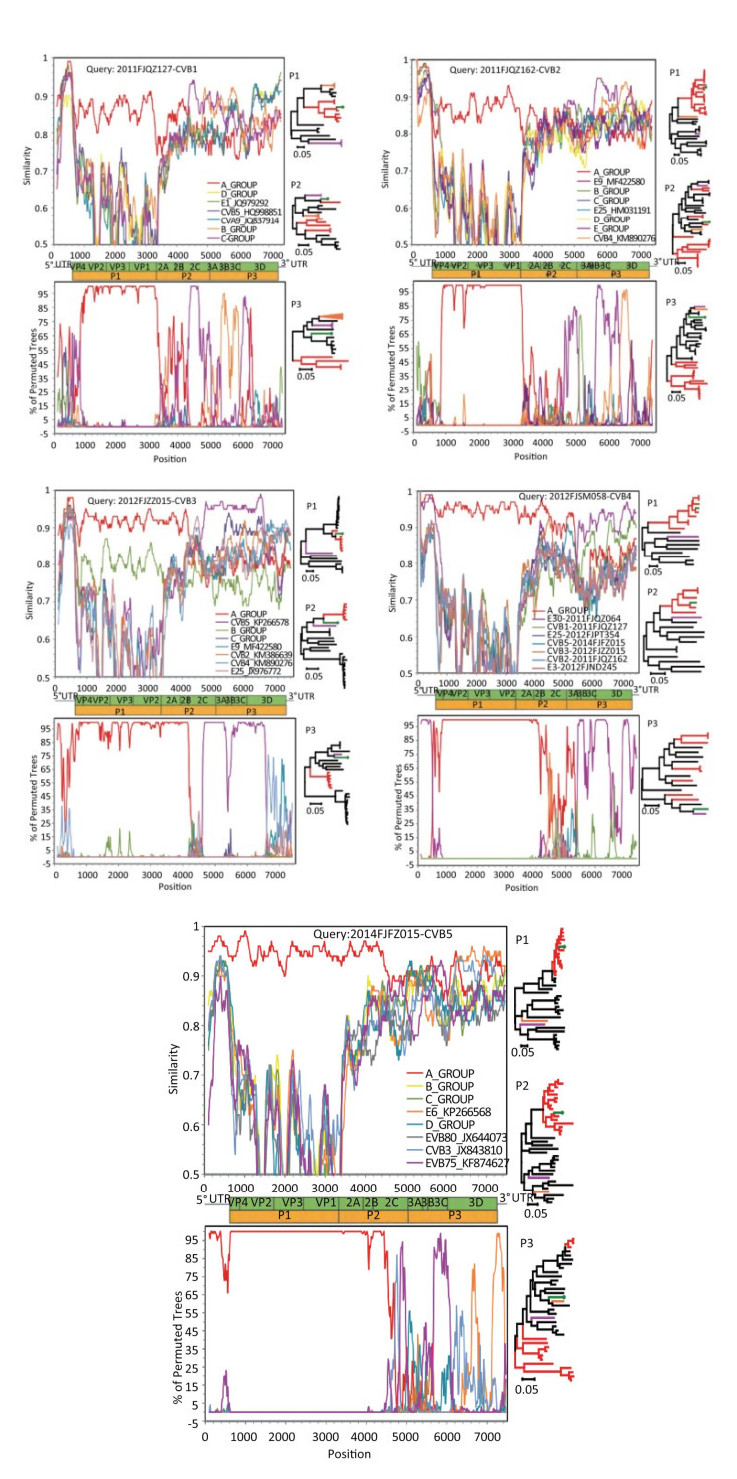
The analysis of similarity plots and boot-scanning demonstrated that the Fujian CVB1-5 strains showed the highest degree of similarity with strains from the same serotype in the P1 gene, but in the P2 and P3 region, the Fujian CVB1-5 strains were the most similar to some HEV-B strains from different serotypes. Simultaneously, maximum likelihood phylogenetic trees based on P1, P2, and P3 regions were constructed individually, using the five Fujian CVB1-5 isolates and potential recombinants (Supplementary Table S3 available in www.besjournal.com and Figure 2). The phylogenetic tree based on the P1 region also showed that the Fujian CVB1-5 strains clustered with strains from the same serotype, but in the P2 and P3 non-capsid region, the Fujian CVB1-5 strains did not cluster with strains from the same serotype and instead, clustered with some HEV-B strains from different serotypes. Using recombination analysis, we found clear evidence of intraspecies recombination in the Fujian CVB1-5 strains. Recombination is a frequent phenomenon in enteroviruses[9]. Frequent recombination events were the basis for the evolution of enteroviruses, thereby contributing to outbreaks of CVB1-5 infection.

Table Supplermentary Table 3. The Reference Strain and Possible Recombinant Strain of the Five Fujian CVB1-5 Strains

Figure 2. Recombination analysis of the five Fujian CVB1-5 strains. The five Fujian CVB1-5 strains in this study are in green, the strains from the same serotype are in red, and the possible donor strains are in pink and orange.
In conclusion, it has been reported that CVB1-5 infection can cause severe complications[5, 6, 10] such as myocarditis, aseptic meningitis, and encephalitis worldwide. Hence, the current HFMD surveillance and basic research efforts should not only focus on EV71 and CVA16, but should be more comprehensive and include other HEV serotypes. This study illustrates the epidemiological characteristics of HFMD cases associated with CVB1-5, the VP1 gene and all genomic characteristics of Fujian CVB1-5 strains, thus providing the basic data needed for the control and prevention of CVB1-5 based diseases.
The authors of this article did not perform any of the studies with human participants or animals.
We are grateful to the National HFMD Laboratory Monitoring Network, especially for the specimens and information regarding the cases provided by the center for disease control and prevention of each prefecture in the Fujian province.
The authors declare that they have no competing interests.
HTML
the Chinese National Science and Technology Major Project 2017ZX10104001
Fujian Provincial Natural Science Foundation 2016J01350
the Department of Science and Technology, Fujian Province 2016Y0011








 Quick Links
Quick Links
 DownLoad:
DownLoad: