-
Oral implant restoration has markedly improved the lives of patients with missing teeth, becoming a preferred treatment option[1]. Despite titanium (Ti) implants have been proven to have a good success rate, there are still many unfavourable factors that can easily lead to loosening or shedding of the implant[2,3]. Contemporary researches primarily examines the effects of oral implant complications, including peri-implant inflammation and diabetes, on the early/long-term osseointegration of implants[4,5], Additionally, there is a significant focus on the innovation of sophisticated implant materials designed to enhance bone regeneration and increase the success rates of implant procedures[6]. However, little attention has been given to the potential harm and mechanism of suboptimal dietary patterns, especially a high-sodium (HS) diet. According to the health guidelines of the World Health Organization (WHO), a daily salt intake for adults should be less than 5 g, yet the average intake in China is roughly double the advised amount, posing a significant dietary risk factor[7]. A high intake of sodium (Na) has been proven to significantly increase the risk of cardiovascular and cerebrovascular diseases, including heart disease and stroke, and is known to contribute to bone degradation and osteoporosis[8,9]. This issue is particularly relevant for the aging population, who are the primary recipients of dental implants and may have a predilection for saltier foods due to diminished taste and saliva production[10]. Therefore, an in-depth exploration of the detrimental impacts and underlying mechanism of HS diets on oral implant restoration is essential. This knowledge is pivotal for advising a healthy postoperative diet for patients with missing teeth, ultimately enhancing the success rate of implants.
In the past few decades, the challenge of regenerating hard tissue has been a major issue in global healthcare[11]. Although most bone injuries can heal naturally with minimal intervention[12], the presence of HS load in the body can profoundly affect the healing process. Bone histomorphometry has indicated that excessive sodium can increase the osteoclast numbers while decreasing osteoblasts and osteoid volume[13], yet the impact of a HS diet on the early osseointegration of Ti-based implants remains contentious. According to Baldisserotto et al., in an aged rat model, a diet with 1 wt% sodium chloride (NaCl) had no significant effect on the early osseointegration of Ti implants[8]. However, Schröder's study found that a HS diet (4 wt% NaCl) seemed to activate osteoclasts and enhance periodontal bone loss[14]. These varying outcomes suggest that the interference of a HS diet on implant repairing could be contingent on NaCl concentration. In addition, little is known about the potential mechanism underlying the effect of a HS microenvironment on the skeletal system, with current researches primarily addressing the acceleration of urinary calcium excretion and osteolysis. Consequently, the potential harm and mechanism by which a HS diet affects oral implant restoration need to be further explored.
Bones have an incredible ability to repair themselves and fully support the structural system of the body[15]. The repair process of bone injuries involves angiogenesis and bone remodelling, which is a complex process. In recent years, our understanding of bone regeneration has evolved. Instead of the traditional "dual" regulatory mechanism of osteogenesis/bone resorption, it is now recognized that bone injury repair involves multiple regulations, including the interaction between vascular regeneration, osteogenesis and bone resorption[16,17,18]. As an important network system in bone, blood vessels not only provide essential nutrients and oxygen for new bone but also provide necessary signaling molecules for bone regeneration[19,20]. Ramasamy and others have demonstrated that vascular endothelial cells (VECs) not only support the function of osteoblasts through cell-matrix signaling but also promote the osteogenic ability of bone progenitor cells by secreting a variety of bone growth regulatory factors[21]. Vuornos et al. have confirmed that VECs can significantly enhance osteogenic differentiation and promote microvessel/new bone formation[22]. Our previous studies have also shown that normal communication between VECs and mesenchymal stem cells (MSCs) is crucial for bone healing and remodeling[23]. Furthermore, studies have revealed that insufficient blood supply occurs when the distance between osteocytes and capillaries exceeds 500 μm, which can easily lead to osteonecrosis and cell death[24]. Therefore, rapid reconstruction of the microvascular system after surgery is vital for achieving early osseointegration of implants[25,26,27].
The intake of high sodium has been conclusively linked to the occurrence and progression of cardiovascular disease[28]. Elevated extracellular Na+ levels is often accompanied by changes in vascular tissue properties[29,30]. Among these changes, VECs are particularly susceptible to stimulation by cardiovascular risk factors, such as high sodium and high glucose, leading to inflammation, cell damage and thereby accelerating the occurrence of vascular diseases[8,31]. In cardiovascular disease studies, it has been observed that a high sodium microenvironment could significantly and negatively regulate VEC migration and angiogenesis. For example, Torres’ research has shown that a HS microenvironment damaged the migration ability of VECs through reactive oxygen species (ROS) and the angiotensin II type 1 receptor AT1R[32]. Fu’s reports also demonstrated that a HS diet can inhibit angiogenesis by activating the expression of NLR family pyrin domain containing 3 (NALP3) in aortic endothelial cells, and the thoracic aortic ring length of NALP3-/- mice treated with HS is significantly longer than that of normal mice[33]. To date, the potential mechanisms of bone loss caused by HS has been partially explained, including: 1) promotion of osteoclast formation[10,34]; 2) induction of urinary calcium loss[13,35]; and 3) impairment of Treg cell function and interference with M2 macrophage formation[36]. However, based on relevant clues, we speculate that a HS diet interferes with early postoperative microangiogenesis, presenting another potential mechanism that hinders the early osseointegration of implants. In this study, we studied the effect of NaCl on the early osseointegration of implants and its mechanism, both in vitro and in vivo.
-
Alpha-minimum essential medium (α-MEM) and Dulbecco's modified Eagle’s medium (DMEM) were purchased from Gibco Co. (United States). Foetal bovine serum (FBS) was obtained from Gibco Co. (Australia). Sodium chloride (NaCl) and cetylpyridinium chloride were provided by Sigma Co., Ltd. (United States). The PrimeScriptTM RT kit and TBGreenTM PremixExTaqTM Ⅱ kit were purchased from TaKara (Japan). The RNA extraction kit was obtained from Tiangen Biochemical Technology Company (China). Thiazolyl blue (MTT), Triton X-100, 4% tissue cell fixation solution, 4’,6-diamidino-2-phenylindole dihydrochloride (DAPI), and fluorescein isothiocyanate (FITC)-labelled ghost pen cyclic peptide were provided by Beijing Solarbio Science & Technology Co., Ltd. (China). The Bicinchoninic acid (BCA) protein concentration determination kit was purchased from Biyuntian Biotechnology Co., Ltd. (China). The Matrigel matrix adhesive was provided by Corning Co. (United States). Other chemicals were obtained from Muke Experimental Equipment Co., Ltd. (China).
-
The animal study protocol was approved by the Animal Ethics Committee of Wenzhou Medical University and was in adherence to the national and local laws, as well as guidelines issued by the Animal Experimental Center of Wenzhou Medical University. Initially, clean Ti rods (diameter: 0.8 mm, length: 10 mm) were underwent a 60 s sandblasting process under 0.45 MPa. Then, these Ti rods were corroded in a mixed acid solution of 37 wt% hydrochloric acid, 98 wt% sulfuric acid, and deionized water (v/v/v = 2:1:1) at 60 °C. The final sandblasted and acid-etched (SLA) titanium was obtained after ultrasonic cleaning and drying. In this study, 18 male Sprague-Dawley (SD) rats, aged 3 weeks, were randomly assigned to 3 groups (n = 6) and fed diets containing 0.8 wt% (control group), 2 wt%, and 6 wt% NaCl. The implantation surgery was performed 2 weeks later. Briefly, rats were anaesthetized before the operation, and then the epiphyseal ends of bilateral femurs were surgically exposed. A round hole (1 mm diameter) was drilled into the centre of the femur using a surgical drill, following which sterilized SLA implants were inserted. Finally, the surgical sites were carefully sutured in layers, and all operations were performed under aseptic conditions. Penicillin was injected intramuscularly for three days post-surgery to prevent potential infection.
-
To evaluate new bone formation around the implants after implantation for 8 weeks, femurs from each rat were collected and fixed with 4% paraformaldehyde. Microcomputerized tomography was used to analyze and reconstruct the three-dimensional structure around the implant, focusing on a region measuring 1 mm in thickness and 2 mm in width. Then, the percentage of bone volume to total volume ratio (BV/TV%), connectivity density, trabecular number (Tb.N), and trabecular spacing (Tb.Sp) were quantified by CTVox, CTAn and CTVol software. These metrics served as key indicators of the newly formed bone mass and its spatial distribution.
-
The fixed femur specimens were decalcified with formic acid for one week, dehydrated through a series of graded alcohol solutions, embedded in paraffin, sliced, and stained with haematoxylin and eosin (HE) and a CD34 monoclonal antibody. Histological observation was performed using an optical microscope.
-
Human umbilical vein endothelial cells (HUVECs) were obtained from the American type culture collection (ATCC) Biological Standard Resource Center. NaCl was dissolved in double distilled water, sterilized, and then added to DMEM under sterile conditions to create HS medium with final Na+ concentrations of 140 mmol/L, 145 mmol/L, and 150 mmol/L. The cells were subcultured twice once cell confluence reached 90%. The passages were named generations 1, 2 and 3 accordingly (Figure 3A).
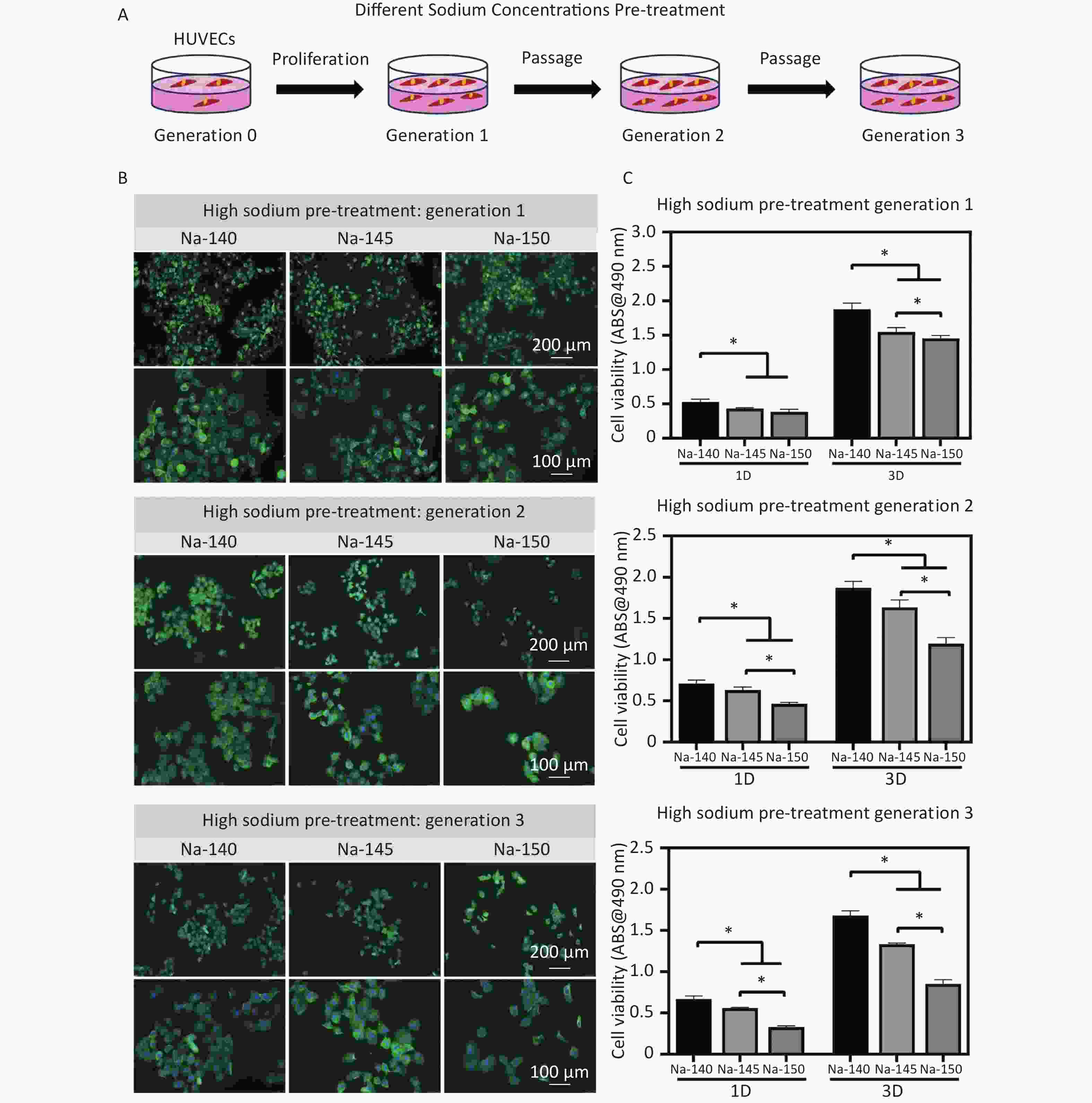
Figure 3. (A) Schematic diagram of pre-treatment of HUVECs (generation 1, 2 & 3) using culture media supplemented with different concentrations of Na+; (B) Fluorescence staining images of HUVECs in each group after pretreating with different concentrations of Na+; (C) Cell viability of HUVECs (generation 1, 2 & 3) in different groups, *P < 0.05.
-
To observe the cell morphology, each group of HUVECs was seeded in a 24-well plate at a density of 8,000 cells per well. After 2 days of culture, the cytoskeleton and nucleus were stained using ghost pen cyclic peptide and DAPI, respectively. Five visual fields were randomly selected for fluorescence imaging. Green fluorescence represents the cytoskeleton, and blue fluorescence represents the nucleus.
Additionally, MTT was used to assess cell viability. Cells were inoculated in 48-well plates at a density of 10,000 cells per well and cultured in HS DMEM for 1 and 3 days. The culture medium was refreshed every two days. For the MTT assay, the culture medium was first removed, and then each well was supplemented with 200 μL of serum-free DMEM containing 10% MTT. After incubation for 4 hours, the 10% MTT medium was removed, and 500 μL of dimethyl sulfoxide (DMSO) was added to each well. The plate was then placed on a shaker for 10 minutes. The absorbance at 490 nm was measured by a microplate reader.
-
The horizontal migration capability of HUVECs was measured. For this purpose, cells from different groups were seeded in 24-well plates at a density of 40,000 cells per well. When the degree of convergence reached 80%–90%, a white gun head was used to cross the line in the centre of the hole plate. Cell migration was recorded at 0 (initial state), 12, and 24 hours using an optical microscope. The cell migration area was measured and calculated by Image J software.
-
Different groups of HUVECs were seeded in 96-well plates, each well covered with matrix glue, at a density of 50,000 cells per well. After being cultured in varied media for 3 and 6 hours, the cells were washed 3 times with phosphate buffered solution (PBS). The resultant vascular rings were observed and analyzed using an optical microscope. Angiogenesis Analyser in Image J software was used to measure the number of computing nodes (points adjacent to three pixels) and connections (boundaries superimposed by four nodes).
-
Real-time quantitative polymerase chain reaction (RT‒qPCR) was employed to detect the expression of key angiogenic genes, including VEGF, FGFR, KDR, CXCR4, and TIE-1. Different groups of HUVECs were seeded in 24-well plates at a density of 20,000 cells per well. After being cultured in different media for 3 days, the total RNA from the cells was extracted using TRIzol, and then the mRNA was reverse-transcribed into complementary DNA (cDNA) using the PrimeScriptTM RT kit. Finally, the expression of different genes was analyzed through RT‒qPCR. Glyceraldehyde 3-phosphate dehydrogenase (GAPDH) was used as the internal reference. The primer sequences are detailed in Table 1.
Angiogenesis-related genes Gene sequence GAPDH F:5’- TCAAGAAGGTGGTGAAGCAGG-3’
R:5’- AGCGTCAAAGGTGGAGGAGTG-3’VEGF F:5’- AGGGCAGAATCATCACGAAGT-3’
R:5’- AGGGTCTCGATTGGATGGCA-3’FGFR F:5’- AATGAGTACGGCAGCATCAAC-3′
R:5’- ACCTCGA TGTGCTTTAGCCAC-3′CXCR4 F:5’- CACTGTTGCCTGAACCCCAT-3′
R:5’- TGTCCACCCCGTTTCCCTT-3′TIE-1 F:5’- AAGCAGACAGACGTGATCTGG-3′
R:5’- GCACGATGAGCCGAAAGAAG-3′Table 1. The primer sequence of angiogenic genes (VEGF, FGFR, CXCR4, TIE-1)
-
Mouse embryonic osteoblast progenitor cells (MC3T3-E1 cells) were obtained from the American ATCC Biological Standard Resource Center. α-MEM medium with Na+ concentrations of 140 mM, 145 mM, and 150 mM was prepared and then used to culture MC3T3-E1 cells for 1, 5, and 10 generations (Figure 5A). The cells were cultured at 37 °C and sub-cultured 9 times once cell confluence reached 90%.
-
To observe the cell morphology, MC3T3-E1 cells from each group were seeded in 24-well plates at a density of 8,000 cells per well. After a 3-day culture, the medium was changed every two days. The staining procedure was the same as that of HUVECs, and the cell morphology was observed using a fluorescence microscope. In addition, MTT assay was utilized to assess cell viability. Different groups of cells were seeded in 48-well plates at a density of 10,000 cells per well and cultured for 4 and 7 days. The detection procedure was also the same as that of HUVECs.
-
The ALP activity was used to evaluate the early osteogenic ability of MC3T3-E1 cells. Cells were seeded in 24-well plates at a density of 20,000 cells per well. After 4 and 7 days of culture, the culture medium was refreshed every two days. Triton X-100 (1%, wt/v) was used to lyse the cells. A BCA kit was used to quantify the total protein in the cell lysate (absorption at 562 nm). An ALP kit was used to measure the ALP activity in the cell lysate (absorption at 520 nm).
-
To measure the mineralization level, MC3T3-E1 cells from each group were seeded in 24-well plates at a density of 20,000 cells per well. After 14 days of culture, the culture medium was changed every two days. Then, the cells were rinsed thrice with sterile PBS, fixed with 4% paraformaldehyde for 30 min, and stained with alizarin red until red calcium nodules appeared. These calcified nodules were dissolved in 10% cetylpyridinium chloride, and the absorbance was measured at 540 nm by a spectrophotometer.
-
All in vitro experiments were repeated at least 3 times. All data were analyzed by one-way analysis of variance (ANOVA) and Student’s t test using SPSS Statistics 22, and presented as the mean ± standard deviation (mean ± SD). The confidence coefficient was set to 95% (*P < 0.05).
-
The 3D contour imaging data (Figure 1A) revealed a notable reduction in new bone formation around the implants in the Na-2% and Na-6% groups compared to the Na-0.8% control group. A dose-dependent decline was observed in key parameters such as BV/TV%, connectivity density, and Tb.N (Figure 1B-D). Particularly in the Na-6% group, the decreases in BV/TV% (8.0% vs. 14.0% of control), connectivity density (7.5% vs. 13% of control), and Tb.N (0.7 vs. 0.9 of control) were considered substantial. In contrast, the Tb.Sp in the Na-6% group was 10% higher than that in the control group (Figure 1E).
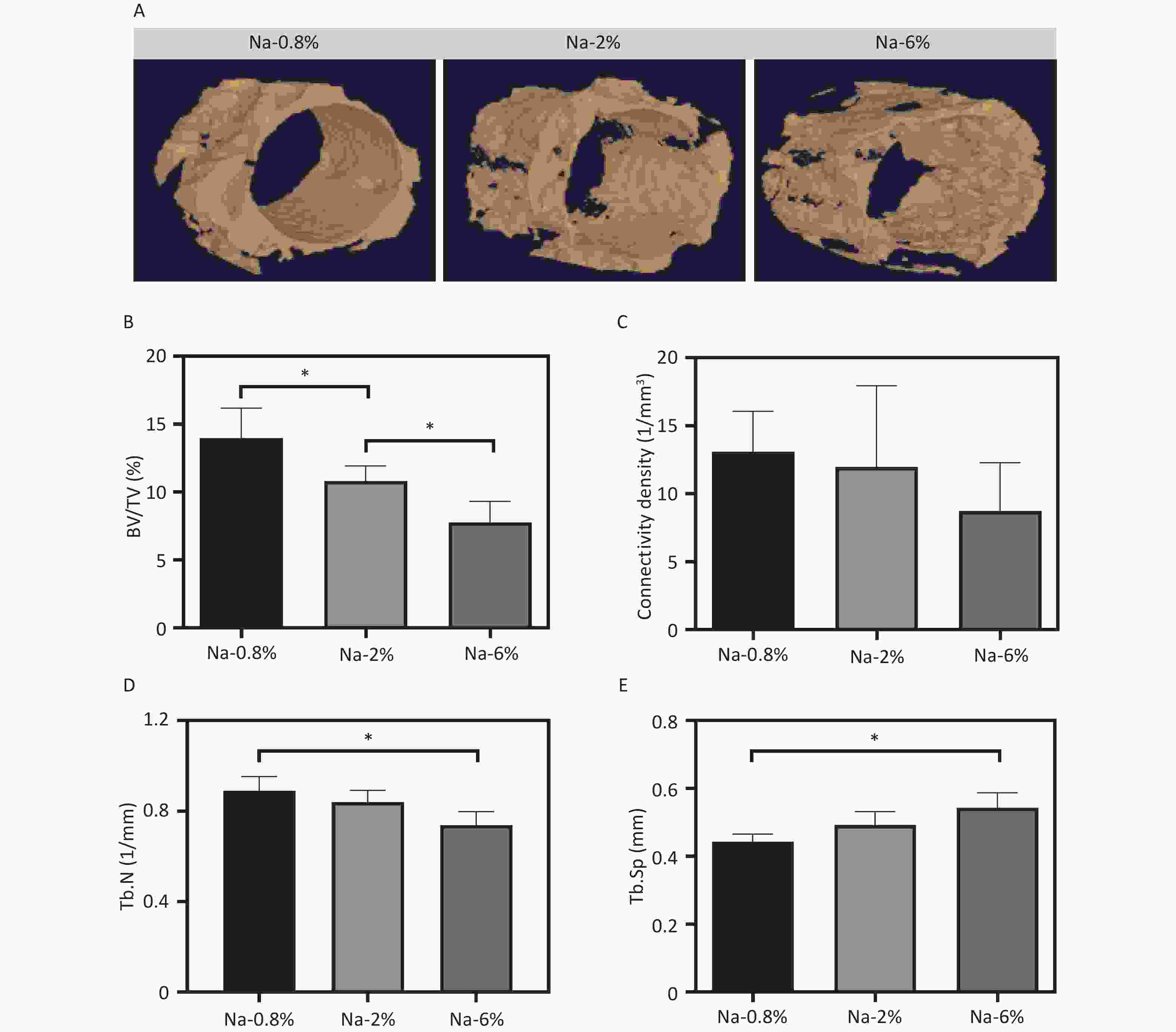
Figure 1. 3D reconstruction images of newly formed bone around the SLA implants; Quantitative results of bone volume/tissue volume (BV/TV%, B), connectivity density (C), trabecular number (Tb.N, D), and trabecular space (Tb. Sp, E) in different groups, *P < 0.05.
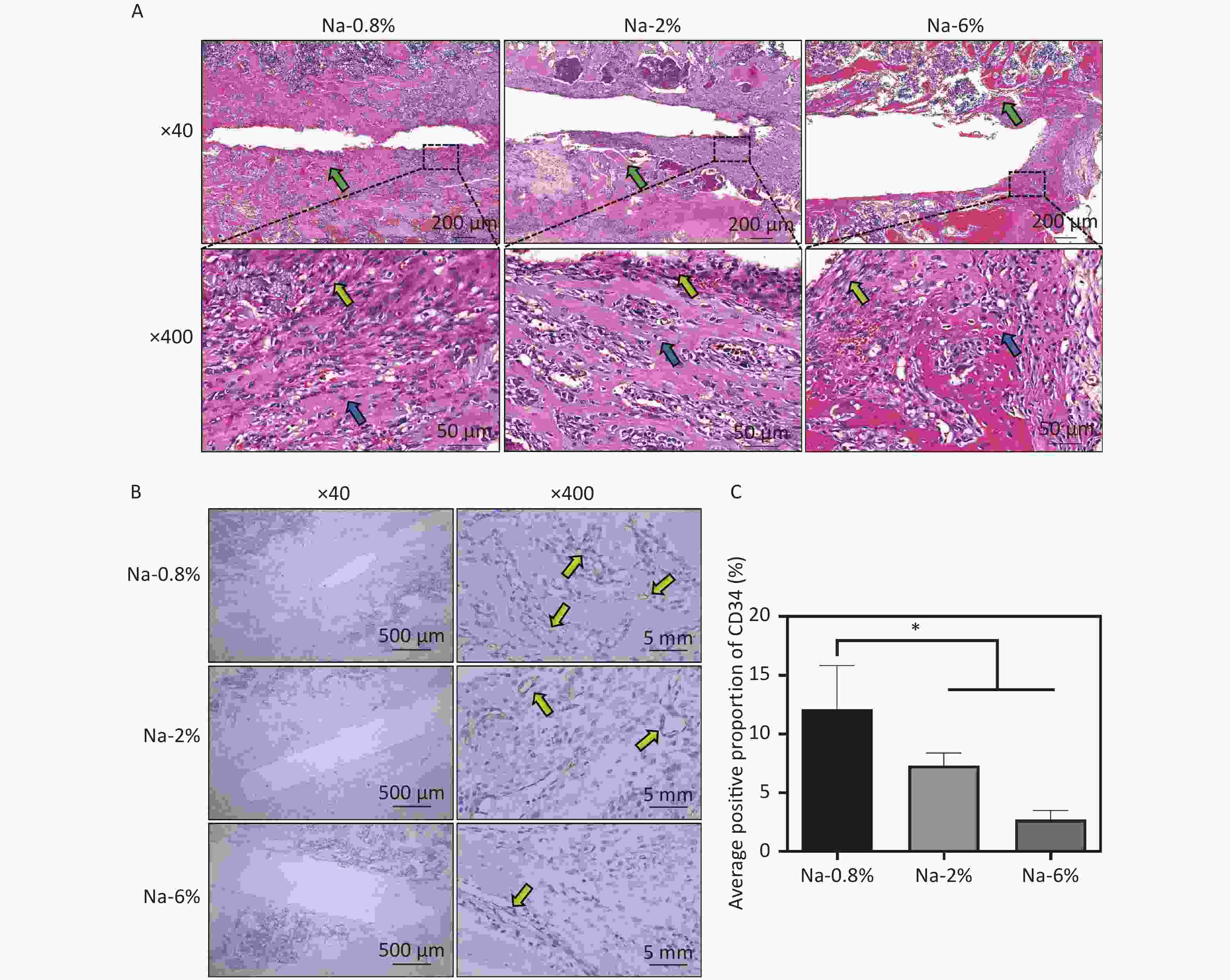
Furthermore, HE staining images (Figure 2A) indicated that the Na-0.8% group had prominent bone trabeculae and fibrous tissues, suggesting that normal repair was underway. Compared with those in the Na-0.8% group, wider bone trabecular space (green arrows), increased inflammatory cells (lymphocytes and neutrophils, blue arrows) and reduced fibroblasts (yellow arrows) were noted in the Na-4% and Na-6% groups (especially the latter). Immunohistochemical analysis further revealed that CD34+ staining (yellow arrows) in the Na-6% group was significantly lower than that in the Na-0.8% group and Na-2% groups (Figure 2B). The trend of the quantitative results (Na-6% < Na-2% < Na-0.8%) was consistent with the microscopic observations (Figure 2C). In summary, both peri-implant neovascularization and new bone formation in the Na-6% group were markedly inferior to those in the Na-0.8% group.
-
After 2 days of culture, the morphology of HUVECs stained with green fluorescence was depicted in Figure 3B. The cells in the control group (Na-140) exhibited robust spreading, indicative of healthy growth. In contrast, the HS groups (Na-145 and Na-150) exhibited signs of cellular atrophy and reduced spreading areas (Figure 3B).
Next, cell viability was measured to determine the cytotoxicity under HS conditions. When HUVECs were cultured for 1 or 3 days, a significant decline in cell proliferation was observed in the HS groups. This decline in cell viability became more pronounced with increasing Na+ concentration. A notable difference was observed between the control group and the HS groups. Additionally, with increasing cell culture time, the cell proliferation of the HS group decreased more obviously (Figure 3C).
-
In the Na-150 group, there were fewer migrating cells than in the other groups at both 12 and 24 hours post cross-scratching (Figure 4A). A dose-dependent decrease in cell migration percentage was evident upon Na+ incubation, as shown in the quantitative analysis in Figure 4B. Specifically, the Na-150 group experienced a decline of approximately 28.3%, 29.6%, and 34.1% in migration after culturing for 1, 2, and 3 generations, respectively, compared to the control group. This trend indicated that with increased cell culture time, the migration efficiency of cells in each group was lower than that of the control group.
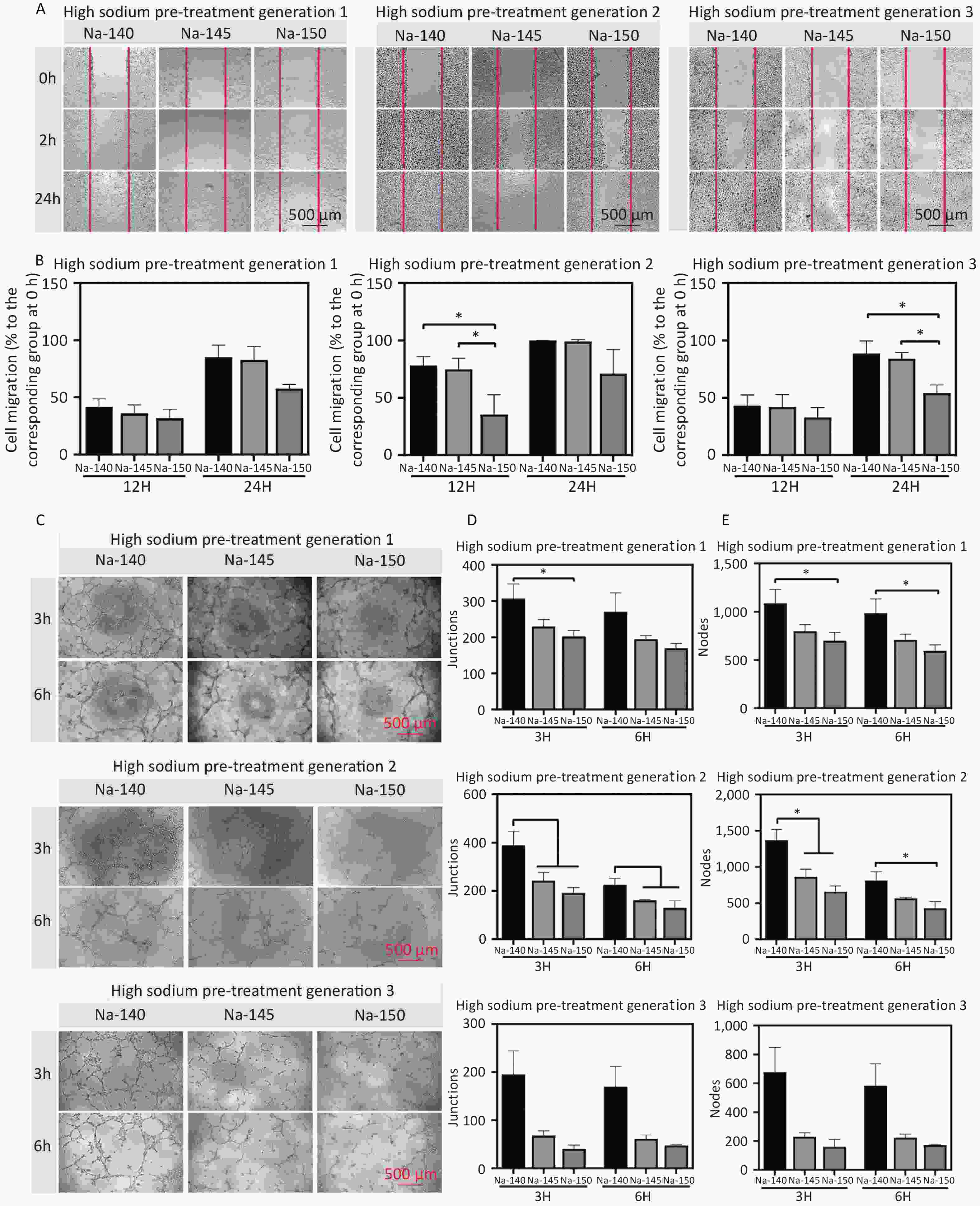
Figure 4. (A) Migration images of HUVECs in different groups at 0, 12, and 24 h; (B) Quantitative analysis of cell migration area after 12 and 24 h of cultivation using culture media supplemented with different concentrations of Na+; (C) Images of tubule formation at 3 and 6 h in each group; Quantitative analysis of the number of nodes (D) and branches (E) formed by tubules, *P < 0.05.
Furthermore, to detect the effect of Na+ on the angiogenesis of HUVECs, vascular ring formation was assessed. The results revealed that the angiogenic ability of the Na-150 group was significantly lower than that of the Na-140 and Na-145 groups, following an order of Na-150 < Na-145 < Na-140. Compared with that of the control group, the ring formation capability of cells cultured in a HS concentration environment was significantly weaker (Figure 4C). The quantitative data on junctions and nodes further supported this trend (Figure 4D & 4E). Furthermore, the vascular ring formation ability of third generation HUVECs cultured in a HS environment was significantly weaker than that of the control group. These results collectively suggest that higher Na+ concentrations and prolonged exposure to a high-sodium environment adversely affect the angiogenic ability of HUVECs.
-
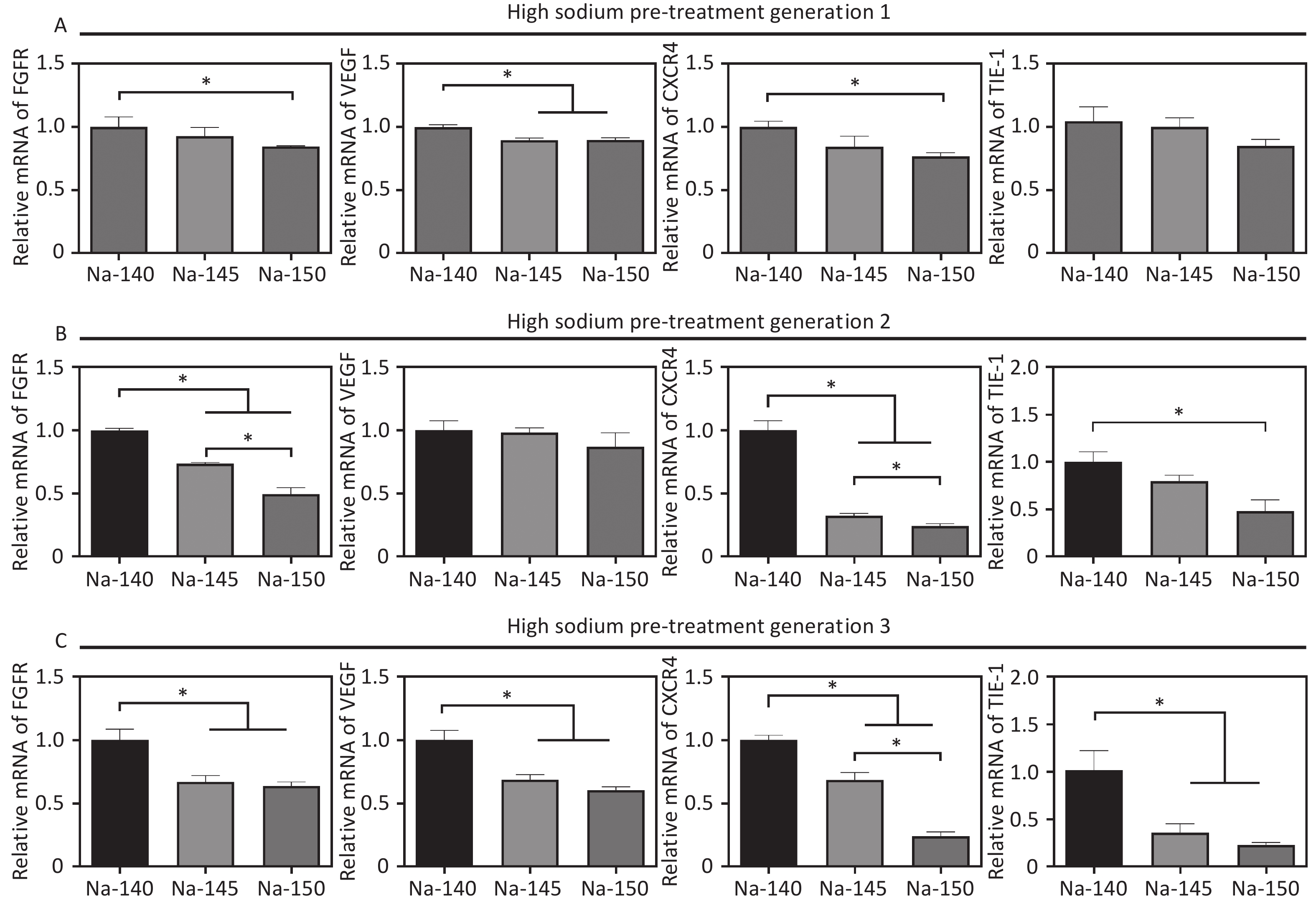
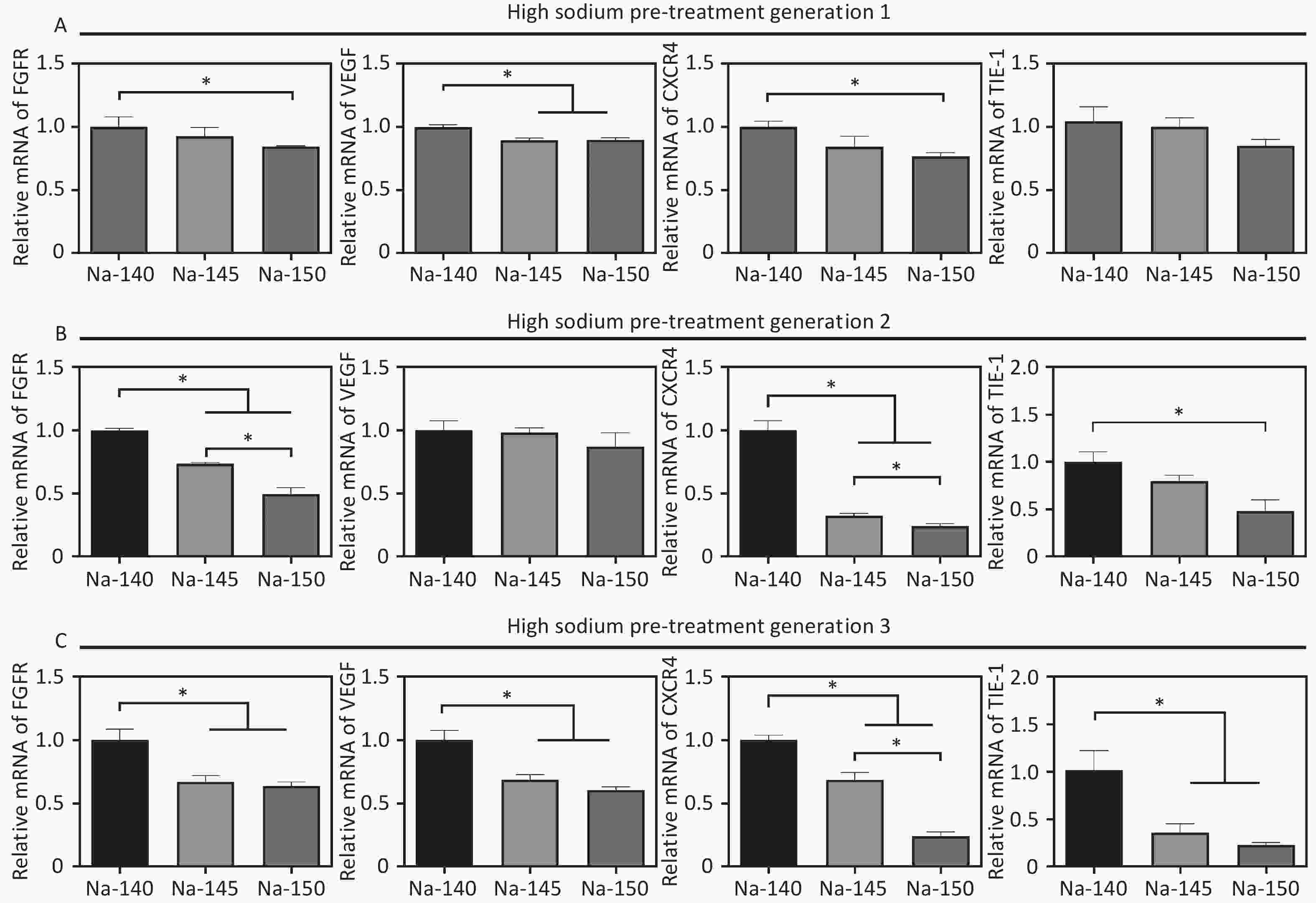
To study the effects of varying Na+ concentrations on angiogenesis, the expression of angiogenesis-related genes (VEGF, FGFR, eNOS, CXCR4) was measured after culturing HUVECs for 3 days. The results were in line with those from the scratch migration and vascular ring formation assays. Specifically, gene expression in the Na-150 group was significantly lower than that in the other groups, with the cells cultured for three generations showing more pronounced sensitivity to sodium concentration (Figure 5). In detail, in the Na-150 group, the FGFR gene expression decreased by approximately 16.2%, 50.7%, and 37.7% for 1, 2, and 3 generations, respectively, in comparison with the control. Likewise, VEGF gene expression in the Na-150 group declined by about 11.6%, 14.3%, and 40.8% across 1, 2, and 3 generations, respectively. The expression of the CXCR4 gene in the Na-150 group showed a decrease of roughly 24.1%, 76.5%, and 77.2% for the respective generations. For the TIE-1 gene, the expression levels in the Na-150 group reduced by about 15.7%, 53.7%, and 77.8% across 1, 2, and 3 generations, respectively. Overall, the presence of a HS microenvironment and prolonged exposure time exerted a concentration-dependent inhibitory effect on the expression of the FGFR, VEGF, CXCR4, and TIE-1 genes. Notably, the impact on CXCR4 and TIE-1 gene expressions was more significant.
-
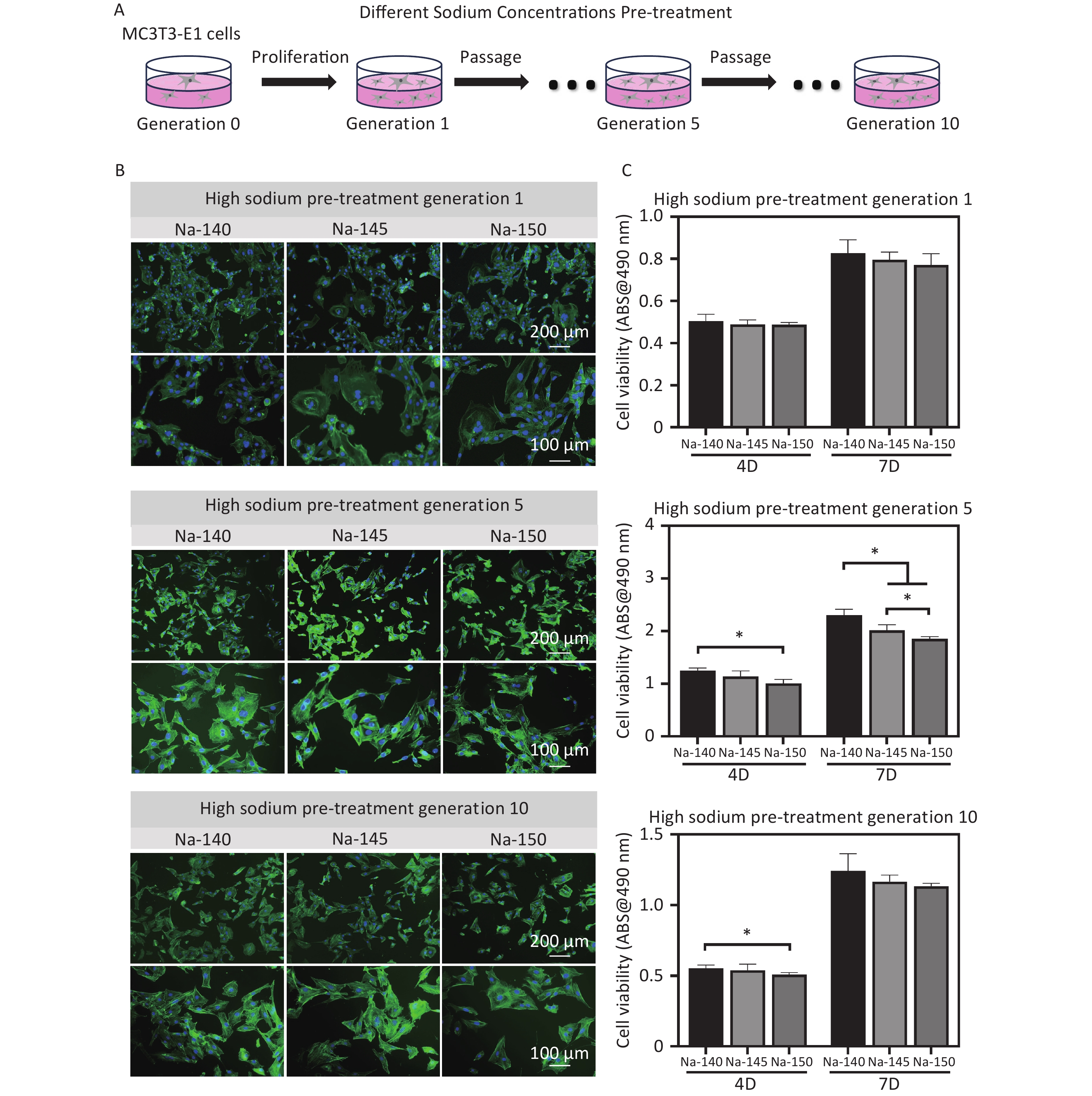
After 3 days of culture, the morphology of MC3T3-E1 cells stained by fluorescence was observed, as shown in Figure 6B. It was noted that there was no substantial difference in cell morphology between the HS groups and the control group. However, in cells of the 5th and 10th generations, the spreading area of cells in the HS groups, excluding the control, was observed to be smaller and had a fusiform/triangular shape. Notably, the spreading area in the Na-150 group was significantly smaller than that in the Na-140 and Na-145 groups (Na-150 < Na-145 < Na-140). In terms of cell viability, no significant difference was initially observed between the high sodium group and the control group. Yet, as the culture time increased, the viability of MC3T3-E1 cells cultured for 4 or 7 days in the Na-150 group was significantly lower than that in the Na-140 group and Na-145 groups (Figure 6C). This trend suggests that prolonged exposure to HS conditions, particularly in the Na-150 group, adversely affects the viability of MC3T3-E1 cells.
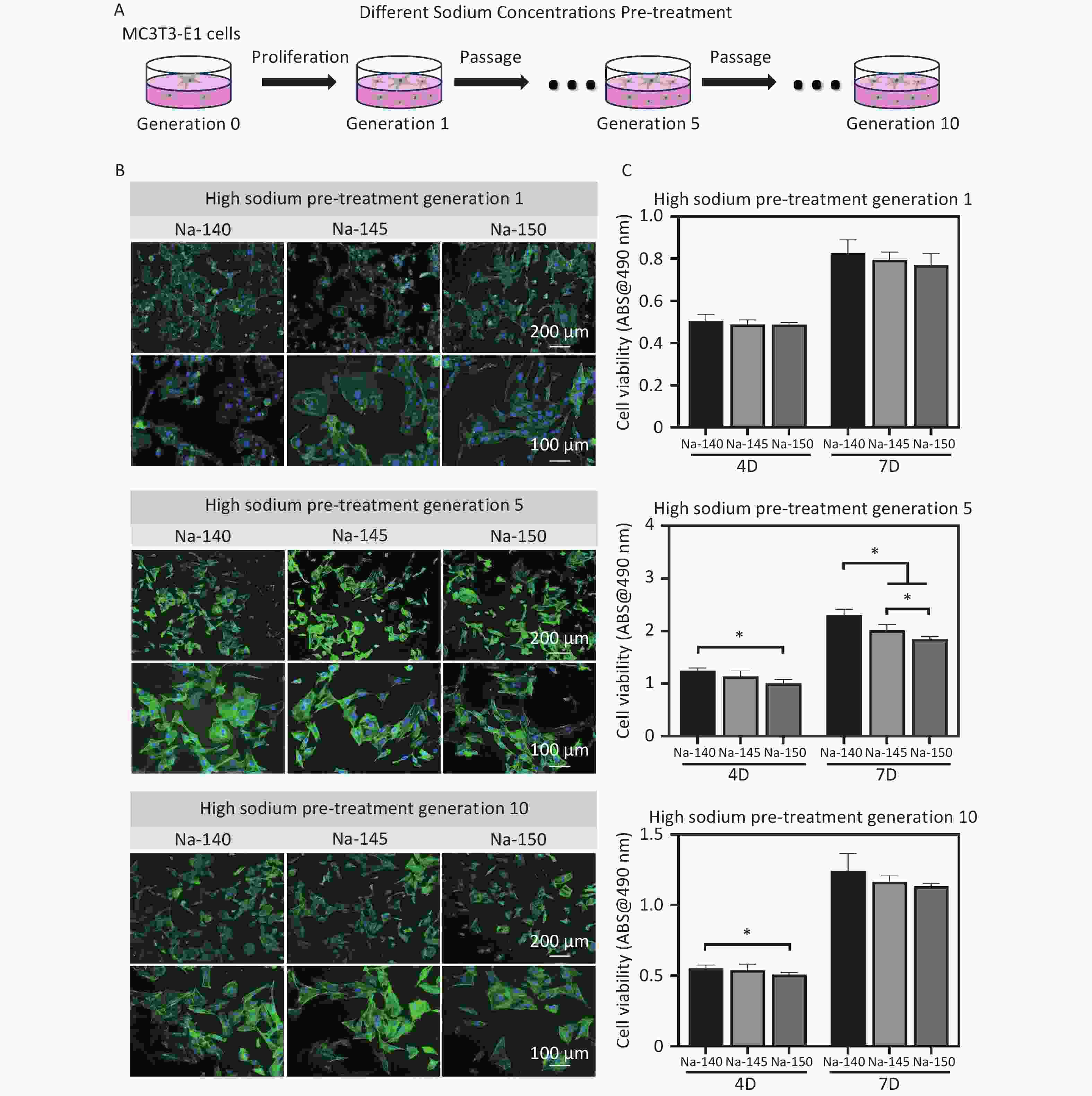
Figure 6. (A) Schematic diagram of pre-treatment of MC3T3-E1 cells (generation 1, 5 & 10) using culture media supplemented with different concentrations of Na+; (B) Fluorescence staining images of MC3T3-E1 cells in each group after pretreating with different concentrations of Na+; (C) Cell viability of MC3T3-E1 cells (generation 1, 5 & 10) in different groups, *P < 0.05.
-
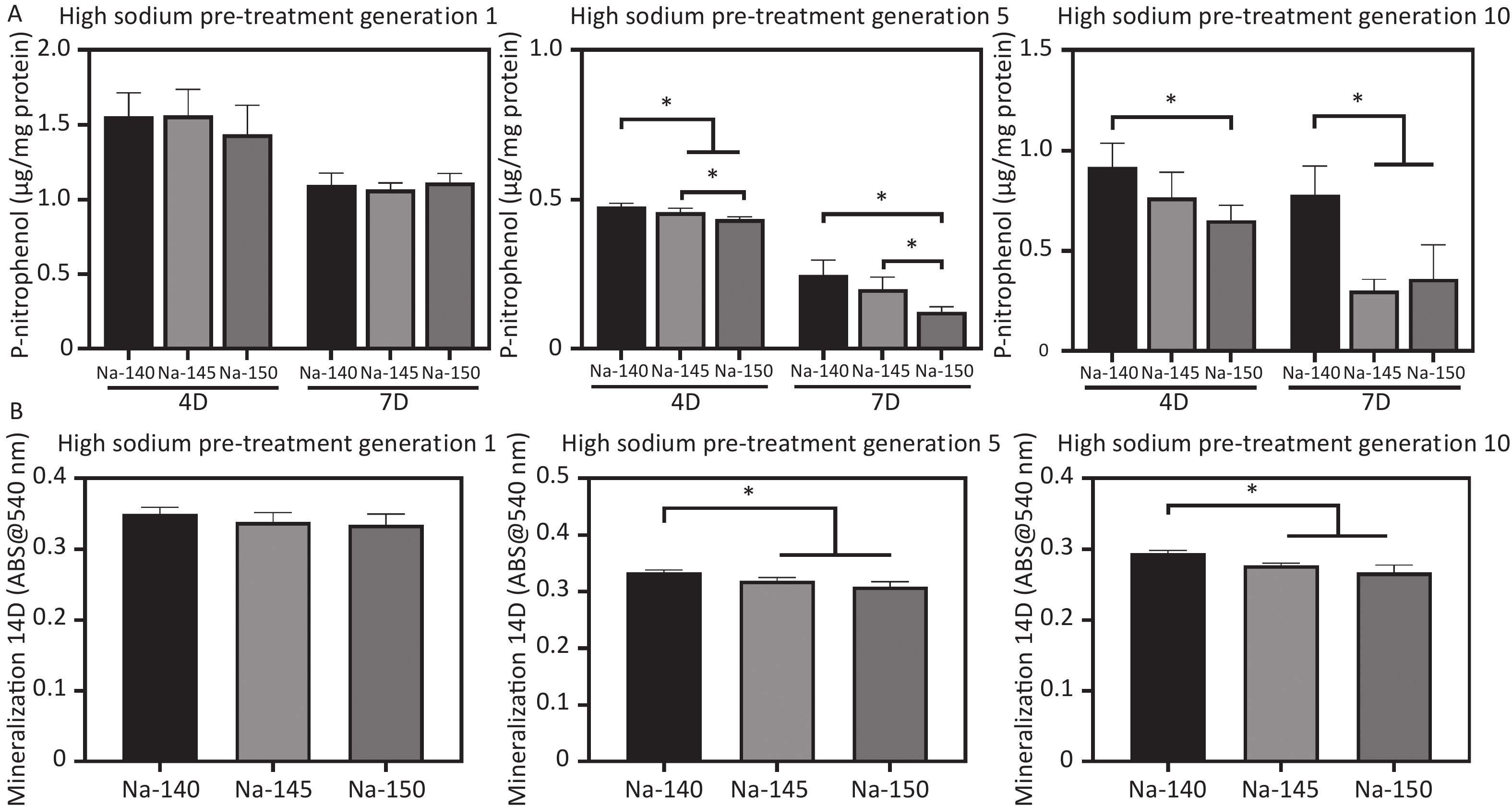
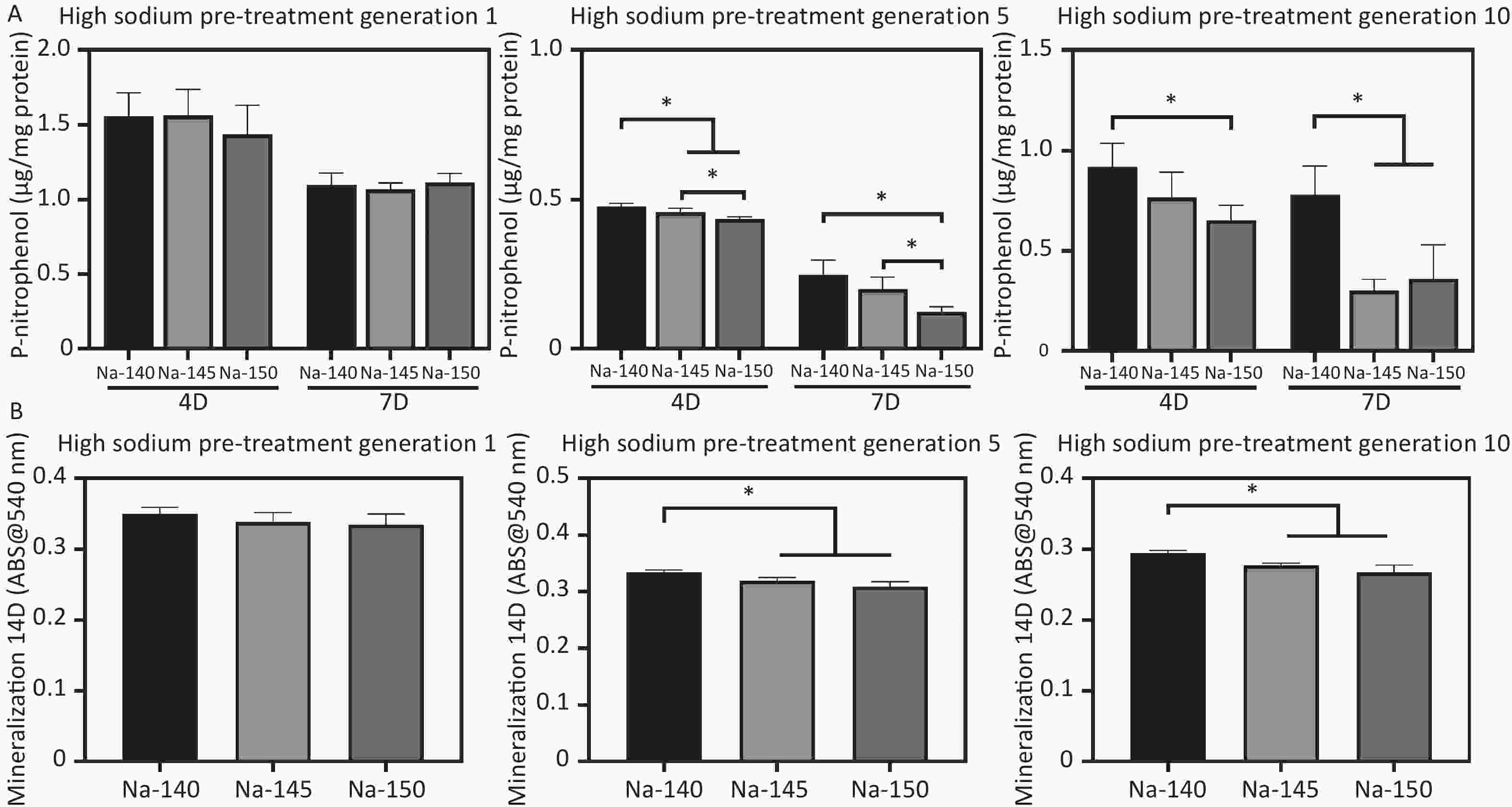
ALP activity was used to evaluate the early osteogenic differentiation of MC3T3-E1 cells, as depicted in Figure 7A. The ALP activity in the high sodium group cultured for one generation did not show a significant decrease compared to the control group. However, as the duration of culture in high sodium conditions increased, a notable reduction in ALP activity was observed in the Na-150 group, compared to the Na-140 and Na-145 groups. In line with the ALP results, the mineralization levels in the Na-150 group, after being cultured in a HS environment for five or ten generation, were found to be the lowest. This difference was statistically significant, as shown in Figure 7B. These results indicate that the effect of a HS environment on the osteogenic ability of MC3T3-E1 cells was Na+ concentration-dependent and time-dependent.
-
Recent research has highlighted that increased salt intake can lead to prolonged retention of Na+ in various human tissues, consequently elevating blood Na+ levels in the human body[37]. In addition, numerous studies have linked HS concentrations to bone destruction, as well as to several inflammatory and cardiovascular diseases, and immunological disorders, suggesting a correlation with excessive dietary sodium intake[38-41]. Some have reported that elevated sodium levels directly augment osteoclast differentiation and fuel inflammation[42]. Additionally, another research has identified components that stimulate the immune system, potentially exacerbating certain autoimmune diseases or bolstering defenses against special infections, such as cutaneous leishmaniasis[43]. Given this backdrop, the specific impact of high sodium concentrations on the early osseointegration of postoperative implants remains largely unexplored.
In this study, we investigated the impact of a high-sodium diet on vascularized bone regeneration around titanium implants, with a particular focus on the Na+ concentration-dependent effects. Our findings revealed that, 8 weeks post-implantation, there was a noticeable decrease in peri-implant blood vessels and bone formation in rats fed diets containing 2% and 6% NaCl (wt/wt), compared to the control group. Titanium-based implants are known to form a fibrous tissue layer between the implant and surrounding tissue[44]. Studies have shown that 1-3 months after implantation, collagen fibers begin to form around the implant and then develop into a reticular fibre structure[45]. Disruption in the vascular composition of the periosteum around the implant can trigger inflammation, resulting in haematoma and cell death[46,47]. The damaged blood vessels are then filled with fibrin, creating a matrix. After platelets are activated in the matrix, monocytes/macrophages and neutrophils invade the blood clot and begin to be gradually replaced by granulation tissue[48]. Monocytes in this environment release inflammatory cytokines, such as interleukin-6 and interleukin-10, as well as tumour necrosis factor-a, thereby increasing the synthesis of prostaglandin[49,50]. These prostaglandins can cause benign inflammatory reactions, such as vasodilation, angiogenesis, and macrophage aggregation at the site of bone injury, thereby enhancing local blood and oxygen supply and promoting fracture healing[51-53]. In the stage of bone union repair and remodeling, most inflammatory cells are replaced by fibroblasts and osteoblasts, progressively completing the bone union process[54]. In our HE staining test, we found a decrease in peri-implant fibroblasts and an increase in inflammatory cells, like neutrophils and lymphocytes, in rats fed a 6% NaCl (wt/wt) diet. This suggests that a HS diet may impede collagen fiber formation around the implant and exacerbate the inflammatory response, thereby slowing osseointegration. Moreover, our immunohistochemical findings showed that the positivity rate of CD34 in the Na-6% group was significantly lower than in the Na-0.8% control group. CD34 is a marker of endothelial cells and is crucial for assessing new capillary (microvessel) formation[55]. It has been noted that CD34 expression is altered in inflammation, particularly in chronic inflammatory diseases. Additionally, activated vascular endothelial cells migrate under the influence of adhesion molecules and chemokines, contributing to endothelial repair and vascular remodeling[56]. This study thus underscores the complex interplay between high sodium intake, inflammation, and bone healing processes.
Excessive salt intake has been implicated in various health issues, including hypertension, cardiovascular disease, and kidney disease, which are known to promote bone resorption[57]. Research by Cui et al. found that a HS diet could deteriorate the microstructure of bone by increasing bone resorption and affecting certain ion channels in bone tissue and renal tubules in ovariectomized rats[58]. Increasingly, studies are suggesting that a HS diet not only affects haemodynamic changes but also destroys immune homeostasis. Liu et al. studied the effect of dietary sodium on the immune response related to food allergies. Their study revealed that excessive salt intake could enhance the Th2 response in a food-allergic mouse model[59]. High sodium intake may also affect blood vessels in the body by affecting the immune system, potentially resulting in organ damage[60,61]. Lee et al. discovered that hypertension caused by a HS diet could result in tissue and vascular damage, often occurring silently for years or decades before any increase in blood pressure is detected[62]. This is consistent with our findings in this study that angiogenesis around the implant was notably reduced in the high NaCl group. This suggests that NaCl influences vascularized bone regeneration around the implant in a concentration-dependent manner.
So far, we have recognized that the normal concentration of serum sodium in the human body ranges from 135 mM to 145 mM. The sodium concentration in the medium we purchased was approximately 140 mM, which is within the normal range, while 145 mM represents the upper limit of normal serum sodium. A serum sodium concentration of 150 mM is considered mild hypernatremia. Therefore, we chose Na+ concentrations of 140 mM, 145 mM and 150 mM for the culture media in our in vitro experiments, aiming to simulate the in vivo microenvironment following a HS diet. This setup allowed us to analyze the concentration-dependent and time-dependent effects of Na+ on MC3T3-E1 cells and HUVECs. In this study, we speculated that long-term HS stimulation might exert a more pronounced inhibitory effect on vascularized bone regeneration. Indeed, we observed that when the 1st, 2nd, and 3rd generation HUVECs were cultured in HS concentrations, the cell viability, proliferation, migration, and tubule formation were significantly decreased compared to the control group in a NaCl concentration-dependent manner. Furthermore, the longer the HUVECs were cultured in a HS environment, the more pronounced the impact on their biological behaviors. Zhang et al. reported that HS intake alone did not cause atherosclerosis, but when combined with endothelial cell (EC) glucocorticoid receptor activation, it led to epithelial sodium channel (EnNaC) activation, thereby increasing EC and arterial stiffness and impairing vascular relaxation[63]. In our previous study, we also found that HS-induced nuclear factor of activated T-cell 5 (NFAT5) controls endothelium-dependent fibrinolysis activity and the expression of inflammatory adhesion-related molecules by regulating plasminogen activator inhibitor-1 (PAI-1)[64]. The results of this study further demonstrated that a high concentration of NaCl could diminish the angiogenic ability of HUVECs in a concentration-dependent manner. Compared to MC3T3-E1 cells, the effect of a HS microenvironment on HUVECs was more significant. Therefore, MC3T3-E1 cells from generations 1, 5, and 10 were chosen for analysis. In the first generation, the cell viability, proliferation, and early osteogenic ability of MC3T3-E1 cells cultured in a HS environment did not show significant differences. This suggests that MC3T3-E1 cells are not immediately affected by a HS environment. However, when generations 5 and 10 were cultured under HS concentrations, there were noticeable effects on biological behaviours like proliferation and early osteogenesis. The longer and higher the concentration of Na+ exposure, the more significant the impact on MC3T3-E1 cell behaviors, indicating a concentration-dependent effect. In subsequent experiments, we plan to further investigate the interaction between HUVECs and MC3T3-E1 cells in HS microenvironment, and explore their potential molecular mechanisms in detail.
-
The purpose of this study was to investigate the effects of varying concentrations and exposure durations of NaCl on vascularized bone regeneration around titanium implants, particularly in the context of a high-sodium diet. It was found that in animal models, both microvascular regeneration and new bone formation around the implant were diminished in rats with elevated blood sodium levels, and these effects were exacerbated with increasing NaCl concentrations. Similarly, in vitro cell experiments demonstrated that the proliferation, migration and angiogenesis of HUVECs were adversely affected by higher Na+ concentrations. Furthermore, the proliferation and osteogenic differentiation of MC3TC-E1 cells also declined with prolonged exposure to elevated Na+ concentrations. Based on these results, we believe that a HS diet could potentially hinder the healing and restoration process of titanium implants. This study not only highlights the possible detrimental effects of excessive sodium intake on implant recovery but also provides a theoretical framework for dietary recommendations post-implant surgery, particularly for patients with edentulous conditions.
Impact of High Sodium Diet on Neovascularization and Osseointegration around Titanium Implant: An in Vivo and in Vitro Study
doi: 10.3967/bes2024.077
- Received Date: 2023-10-08
- Accepted Date: 2023-12-11
-
Key words:
- High-sodium /
- Implants /
- Vascularization /
- Osseointegration.
Abstract:
We have no conflicts of interest to declare.
| Citation: | XU Ke Yuan, TANG XiaoTing, XIANG Yun, SHEN YiDing, DENG ZhenNan, MA PingPing, SHEN Xin Kun. Impact of High Sodium Diet on Neovascularization and Osseointegration around Titanium Implant: An in Vivo and in Vitro Study[J]. Biomedical and Environmental Sciences. doi: 10.3967/bes2024.077 |








 Quick Links
Quick Links
 DownLoad:
DownLoad: