-
Particulate matter (PM), which is a great environmental concern, has been classified as a Group 1 human carcinogen by the International Agency for Research on Cancer (IARC)[1]. Epidemiological and experimental studies have indicated that chronic exposure to PM, especially PM2.5 (particles with an aerodynamic diameter less than 2.5 μm) may lead to an increase in lung cancer incidence[1-2]. PM2.5 entering airway epithelial cells and alveolar macrophages contribute to the production of reactive oxygen species (ROS)[3], a major cause of many adverse effects induced by PM2.5[4]. Such increases in the level of ROS, induced by PM2.5, may lead to various deleterious effects including inflammation, DNA damage, cell apoptosis, and abnormalities in the epigenetics machinery. Recent reports have indicated that DNA methylation and histone modifications may be involved in PM2.5-induced gene modification at specific loci such as p53, AHR, and SATɑ[4-5]. This supports the notion that epigenetic changes may contribute to PM2.5-induced cellular toxicity and tumorigenicity. However, some studies have shown that long non-coding RNAs (lncRNAs) may act as an important epigenetic factor involved in PM2.5-induced adverse reactions. Long non-coding RNAs (lncRNAs), a class of transcripts (> 200 nt in length) which, although similar in structure to that of mRNAs, do not encode proteins, may play crucial roles in many normal cellular processes as well as in the multistep processes of carcinogenesis[6]. The lncRNAs, which are mostly transcribed by RNA polymerase Ⅱ, interact with various target molecules, including proteins, microRNAs, and mRNAs, and participate in processes such as transcriptional regulation, epigenetic gene regulation, and disease induction[7]. It is acknowledged that lncRNAs are subjected to the same regulatory mechanisms that other protein-coding genes are subjected to, including that of epigenetic regulation[8]. DNA methylation is an epigenetic modification that plays a pivotal role in gene regulation at the transcription level. Several studies have reported that CpG island methylation-associated silencing of lncRNA may induce the function of tumor suppressors such as lnc-WT1 and lnc-POU3F3 in human cancer cells[9]. However, high CpG methylation of promoters such as lncRNA MEG3, may cause downregulation of some tumor-suppressive lncRNAs in many cancers[10]. Therefore, aberrant hypermethylation in the regulatory regions of lncRNAs may play a role in cancer development as well as in other biological processes. However, to our knowledge, an association between lncRNAs and DNA methyltransferases (DNMTs) has not been reported so far in PM2.5-treated cells.
Several reports have indicated that aberrant expression of DNMTs and lncRNA may be associated with several cancers, including lung cancer, which mainly results from exposure to PM[11]. Therefore, we hypothesized that regulation of lncRNA by DNMTs may be involved in cellular apoptosis induced by PM2.5.
To test this hypothesis and to investigate the possible molecular mechanisms underlying this process, we examined cellular apoptosis and the expression of a group of lung cancer-related lncRNAs, namely LUCAT1, MALAT1, and ARINL, in the human bronchial epithelial cell line, BEAS-2B treated with PM2.5.
The BEAS-2B cell line, kindly provided by Professor LYU Jia Chun (Guangzhou Medical University), was cultured in RPMI-1640 medium containing 10% fetal bovine serum and maintained in a 100% humidified atmosphere of 5% CO2 at 37 ℃. PM2.5 (National Institute of Standards and Technology, Cat. no. 1650b), was dissolved in phosphate-buffered saline (PBS) via ultrasound for 30 min on ice, and sterilized by filtration[37]. Cell media containing BEAS-2B cells were treated with 300 μg/mL of Sterilized PM2.5 for 48 h, with phosphate-buffered saline (PBS) as the control.
The data obtained from three independent experiments were expressed as mean ± SD. One-way ANOVA or a double-sided Student's t-test was used to test differences among the samples, and least significant difference testing was used for post-hoc multiple comparisons. Statistical analysis was performed using SPSS software (version 19.0). The level of significance was set at P < 0.05.
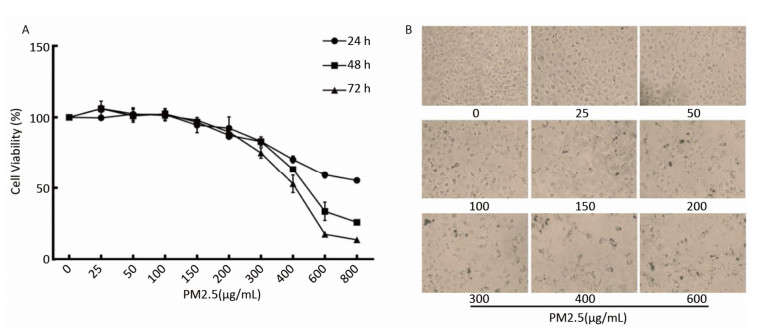
PM2.5 reportedly induces many biological processes including apoptosis, cell cycle, autophagy, DNA damage, and oxidative stress[7] associated with lung tumorigenesis. To determine the effects of PM2.5 on cell viability, BEAS-2B cells were treated with serial PM2.5 concentrations (PBS, 25, 50, 100, 200, 300, 400, 500, 600 μg/mL) for 24 h, 48 h, and 72 h, respectively. Results showed that cell viability decreased with increasing PM2.5 concentration, and that longer periods of exposure caused more significant changes in cell viability (Supplementary Figures S1 available in www.besjournal.com). Based on the above results and the principle that cell viability should be maintained above 75%, so that modest toxicity could be observed, 300 μg/mL of PM2.5 was selected for the study. As the objective was to explore the effect of PM2.5 on DNA methylation, the 48-h time point was selected.
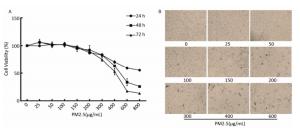
Figure Supplementary Figure S1. Changes of cell viability of BEAS-2B cells induced by PM2.5. (A) Cells were incubated in serial concentrations of PM2.5 exposure for 24, 48 and 72 h respectively, and then cell viability was tested by CCK-8 assay kit. Data were expressed with the mean of three independent experiments. (B) Morphological change of BEAS-2B cells treated with serial concentrations of PM2.5.
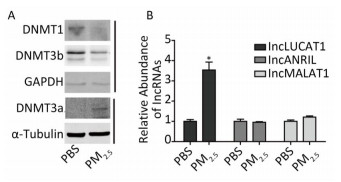
Aberrant DNA methylation patterns, mainly resulting from abnormal expression of DNMTs, are commonly observed in various cancers, including lung cancer. PM2.5, a main cause of lung cancer, may also alter DNMTs expression, as increasingly reported. Western blot analyses showed decreased DNMT1 and DNMT3b protein levels, and increased DNMT3a protein levels in the PM2.5-treated cells (Figure 1A). Previous reports showed that expression of LUCAT1, MALAT1, and ARINL was significantly changed in lung cancer. However, the mechanisms underlying such aberrations remain unclear. To determine the effect of aberrant DNMTs on gene expression, the expression of the three lncRNAs stated above was evaluated via qRT-PCR (Supplementray Table S1 available in www.besjournal.com). LUCAT1 expression was increased in PM2.5-treated cells, compared to that in PBS-treated cells. The expression of MALAT1 and ANRIL remained unchanged (Figure 1B). These results indicate that a decrease in DNMT1 and/or DNMT3b, induced by PM2.5, may play a role in the activation of LUCAT1, which, in turn, may be based on the negative effect of DNA methylation on gene expression.
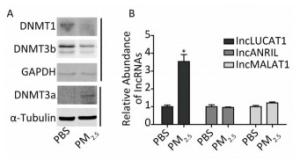
Figure 1. Altered expression of lung cancer-related lncRNA, and DNMTs in BEAS-2B cells treated with PM2.5. (A) DNMTs were detected using western blot. Protein expression was normalized against GAPDH or α-tubulin protein. (B) The expression of lncLUCAT1 was increased in malignant cells, while that of lncANRIL and lncMALAT1 remained unchanged. lncRNA expression was analyzed using RT-PCR. The lncRNA levels were normalized against GAPDH mRNA and the data were expressed as fold change relative to PBS-treated control cells. All assays were performed in triplicate; representative western blot images obtained from three similar experiments are shown. Each bar represents the mean ± SD of three independent experiments. *P < 0.05 vs. PBS group for the same gene.
Primer Primer Sequence (5'-3') lncLUCAT1-M-F TTTTAGAAATAAGGTTGAGTGTCGT lncLUCAT1-M-R TTACTTCAACTTCCTAAATACCGAA lncLUCAT1-U-F TTAGAAATAAGGTTGAGTGTTGT lncLUCAT1-U-R TTACTTCAACTTCCTAAATACCAAA lncMALAT1-F TTTGTTCATTTCTGGTGGTGGG lncMALAT1-R TAAGACCAAGGGAGGGGAGAG lncLUCAT1-F TGTCAAGCTCGGATTGCCTT lncLUCAT1-R GCTGGGTGAGCTTCTTGTGA lncANRIL-F TTGTGAAGCCCAAGTACTGC lncANRIL-R TTCACTGTGGAGACGTTGGT GAPDH-F GGAGTCAACGGATTTGGTCGTATTG GAPDH-R TCTCGCTCCTGGAAGATGGTGAT Table Supplementary Table S1. Sequence of PCR Primers
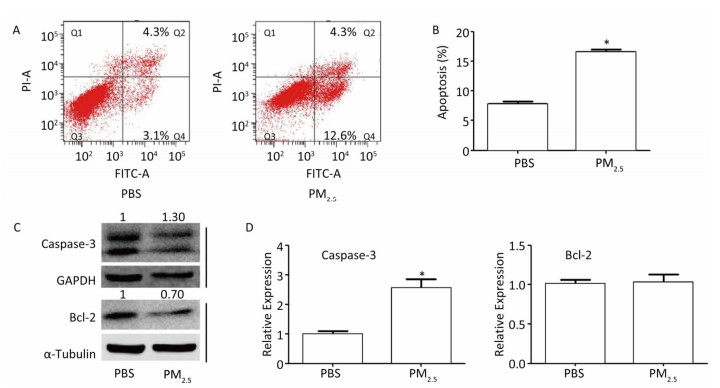
PM2.5 may induce cell apoptosis, which is common in lung cancer[7]. In order to clarify the role of LUCAT1 in cell apoptosis, cell apoptosis was assessed using a cytometer assay. The results of the assay showed that in cells treated with 300 μg/mL of PM2.5, the percentage of cells that underwent apoptosis increased from -7.8% to -16.9%, compared to that in the PBS-treated cells (Figure 2A, 2B). Caspase-3, a pro-apoptosis gene, and Bcl-2, an anti-apoptosis gene, were used as indicators of cell apoptosis. Expression of caspase-3 increased at both mRNA and protein levels, whereas that of Bcl-2 decreased at the protein level with no change at the mRNA level (Figure 2C, 2D). Alternation of caspase-3 and Bcl-2 proteins was consistent with the apoptosis results obtained from the cytometer assay. These results may confirm the contention that PM2.5 may induce cell apoptosis, which, according to our hypothesis, involves de-regulated DNMT1 and/or DNMT3b-mediated activation of LUCAT1.
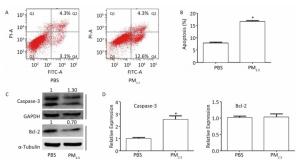
Figure 2. PM2.5 plays a role in the apoptosis of BEAS-2B cells. (A & B) Apoptosis was measured using the PI/Annexin V apoptosis detection kit (n = 3). (C) Levels of the Bcl-2, and Caspase-3 proteins were detected using western blot. All assays were performed in triplicate; representative western blot images obtained from three similar experiments are shown. (D) Expression of Caspase-3 and Bcl-2 was analyzed using qRT-PCR. Each bar represents the mean ± SD of three independent experiments. *P < 0.05 vs. PBS group for the same indicator.
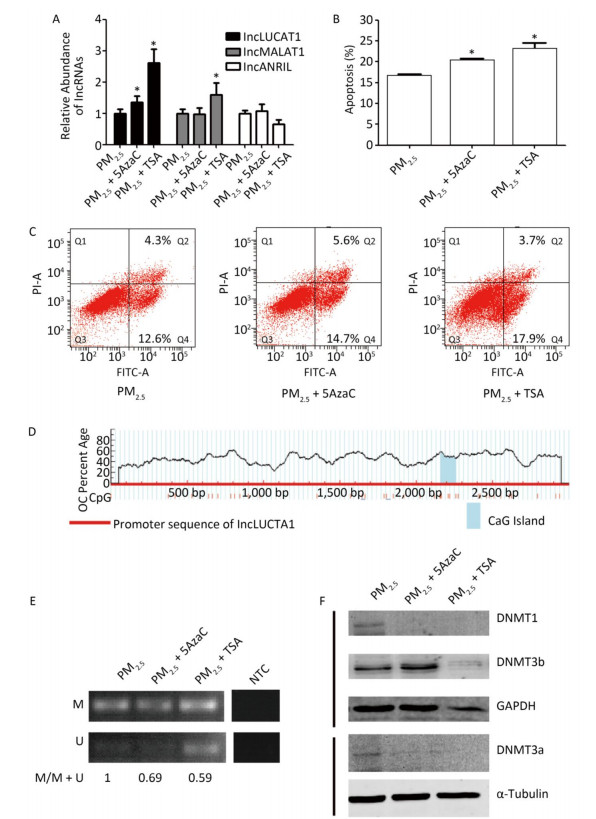
In order to further confirm our hypothesis, we used 5-Aza-2'-deoxycytidine (5-AzaC), a known DNMT inhibitor, to inhibit DNMTs, and TSA, a histone deacetylase (HDAC) inhibitor, to repress HDACs, which are co-repressors of transcription regulated by DNA methylation. The expression of LUCAT1, but not that of MALAT1 or ANRIL, was increased in cells treated with PM2.5 in combination with 5-AzaC or TSA, compared to that in PM2.5-treated cells (Figure 3A). Furthermore, the percentage of cells that underwent apoptosis increased from -16.9% to 20.5% and -22.1%, respectively (Figure 3B, 3C). These results indicate that de-regulated DNMT1 and/or DNMT3b-mediated activation of LUCAT1 may play a role in cell apoptosis induced by PM2.5. To further elucidate the mechanism underlying the activation of LUCAT1 transcription by DNMT1 and/or DNMT3b, MSP was used to detect DNA methylation of the LUCAT1 promoter (Figure 3D). The percentage of 5'-methylcytosine in the LUCAT1 promoter decreased in cells treated with PM2.5 in combination with 5-AzaC or TSA (Figure 3E, Supplementary Table S1), which was consistent with LUCAT1 expression. DNMT1 expression decreased in cells treated with PM2.5 in combination with 5-AzaC, but DNMT3b expression was slightly increased (Figure 3F). Expression of both DNMTs decreased in cells treated with PM2.5 in combination with TSA. The results for both DNMTs showed that DNMT1 may be involved in the regulation of LUCAT1 via demethylation.
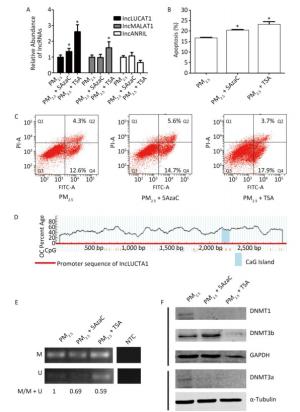
Figure 3. 5-AzaC or TSA enhances LUCAT1 expression and apoptosis via demethylation of LUCAT1 promoter. (A) qRT-PCR was used to detect the levels of lncRNA in BEAS-2B cells incubated with PM2.5 for 24 h, and then incubated with 5-AZA (PM2.5 + 5AzaC) or TSA (PM2.5 + TSA) for 24 h. RNA levels were normalized against GAPDH, and the data were expressed as the fold change relative to the PM2.5-treated control cells. 5-AzaC or TSA treatment promoted apoptosis (B & C) and DNA demethylation of LUCAT1 (D & E). (D) CpG island and structure of LUCAT1 promoter. (E) Representative image of MSP obtained using the methylation and non-methylation PCR primers. Apoptosis was assessed using the PI/Annexin V apoptosis detection kit. (F) DNMT1 and DNMT3a were repressed using 5AzaC or TSA. DNMTs were detected via western blot. Protein expression was normalized against GAPDH or α-tubulin protein. The data are expressed as the mean ± SD. *P < 0.05 vs. PM2.5 group for the same indicator.
In summation, these results may be useful in clarifying the relationship between methylation and biological functions, thereby providing a potential target for treatment of lung cancer as well as a powerful tool for early diagnosis.
LncRNA LUCAT1 Activation Mediated by the Down-regulation of DNMT1 Is Involved in Cell Apoptosis Induced by PM2.5
doi: 10.3967/bes2018.082
- Received Date: 2018-06-05
- Accepted Date: 2018-08-03
| Citation: | LING Xiao Xuan, ZHANG Hai Qiao, LIU Jia Xian, LIU Zhi Dong, PENG Jian Ming, ZHANG Xiao Qing, SHAO Jun Li, CHEN Zhen Fa, LIU Lin Hua. LncRNA LUCAT1 Activation Mediated by the Down-regulation of DNMT1 Is Involved in Cell Apoptosis Induced by PM2.5[J]. Biomedical and Environmental Sciences, 2018, 31(8): 608-612. doi: 10.3967/bes2018.082 |







 Quick Links
Quick Links
 DownLoad:
DownLoad: