-
Vibrio parahaemolyticus is a Gram-negative halophilic bacterium that primarily causes seafood-associated gastrointestinal illness in humans [1]. V. parahaemolyticus produces multiple virulence determinants, such as two type-III secretion systems (T3SSs) and thermostable direct hemolysin (TDH) or TDH-related hemolysin, and forms robust biofilms on surfaces that support bacteria to survive in adverse conditions [1, 2]. V. parahaemolyticus employs various specific structures, such as flagella, type-IV pili, and exopolysaccharide (EPS), to form mature biofilms [2]. EPS is the main component of the biofilm matrix [2]. In V. parahaemolyticus, the cpsA-J operon is responsible for the synthesis of EPS [3]. The deletion of cpsA-J led to smooth colonies on plates, whereas the wild-type (WT) strain formed wrinkled colonies [3]. Previous studies showed that the variations between smooth and wrinkled colonies are strictly regulated by intracellular cyclic diguanylate (c-di-GMP) levels, CpsS-CpsR-CpsQ cascade, quorum sensing (QS), and histone-like nucleoid structuring protein (H-NS) in V. parahaemolyticus [4-10].
C-di-GMP is a ubiquitous secondary messenger involved in regulating multiple behaviors of bacteria, including biofilm formation, motility, and virulence [11]. Increased intracellular c-di-GMP promotes biofilm formation and motility inhibition [12]. The synthesis of c-di-GMP is catalyzed by diguanylate cyclase containing a GGDEF domain, whereas its degradation is associated with phosphodiesterases (PDE) possessing either an HD-GYP or EAL domain [12]. In V. parahaemolyticus, two proteins, namely, ScrG and ScrC, contain both GGDEF and EAL domains and act as PDEs to degrade c-di-GMP [4, 6]. In addition, the GGDEF-type proteins ScrO, GefA, ScrJ, and ScrL and the EAL type protein LafV are required for the regulation of motility and biofilm formation in V. parahaemolyticus [13-15]. We recently demonstrated that a group of genes encoding the proteins with GGDEF and/or EAL domains, including scrABC and scrG, were under the direct regulation of the master QS regulator OpaR, which acts as a repressor of biofilm formation and c-di-GMP production in V. parahaemolyticus RIMD2210633 [9].
H-NS is an abundant nucleoid-associated DNA-binding protein involved in chromosome folding and gene regulation in Gram-negative bacteria [16]. H-NS prefers to bind AT-rich DNA sequences and has been generally described as a negative regulator of gene expression [16]. In V. parahaemolyticus, H-NS is a negative regulator of the major virulence determinants, including T3SS1, T3SS2, TDH, T6SS1, and T6SS2 [17, 18]. The deletion of hns led to enhanced swarming motility and laf expression in V. parahaemolyticus [19]. In addition, previous studies showed that H-NS acts as a positive regulator of biofilm formation by V. parahaemolyticus RIMD2210633 because the hns mutant formed smooth colonies and manifested lower expression of cpsA-J [8, 20]. However, the regulatory mechanisms remain not fully understood. The data on crystal violet (CV) staining presented here showed that H-NS acted as a repressor of biofilm formation by V. parahaemolyticus RIMD2210633. H-NS was a negative regulator of intracellular c-di-GMP synthesis and transcription of scrA, scrG, VP0117, VPA0198, VPA1176, VP0699, and VP2979, encoding a set of proteins containing GGDEF and/or EAL domains.
-
V. parahaemolyticus strain RIMD2210633 was used as the WT [21]. Non-polar hns deletion mutant (Δhns) and complementary Δhns (C-Δhns) were constructed in our previous study [17]. Control strains were also constructed by transferring the empty pBAD33 into the WT and Δhns strains to counteract the effects of arabinose and chloramphenicol on the results [17]. Table 1 lists all the primers used in the current study.
Table 1. Oligonucleotide primers used in this study
Target Primers (forward/reverse, 5'–3') Reference Protein expression hns AGCGGGATCCATGTCAGAGCTGACTAAAACAC/AGCGAAGCTTTTAGATTAGGAAATCGTCTAG [17] qPCR scrA CACACCACGAACACATTGC/TCAATAGCGTCACGGAATGC [9] scrG AAGCCGTGGTGGAAGAAGG/GCGTGTTGAGTGCGTTGG VP0117 GACCACCTCAATAGTTATCTG/TAAGTAGGCTTGGACATCTC VPA0198 GCATCAGAATCAGCAAGAC/ATGCTTAGCTCCTCTTCTTC VPA1176 GCCATATTCCAAACTCGTTGTG/TGCGTAAGCCAAGTTGATGAG VP0699 CTGACACATCGTGATACTTC/TTGATGTTGCAGCTCTTG VP2979 GCAACTCTCAAGTCATCATC/CAACAACCGTCTTCTATGG LacZ fusion scrA GCGCGTCGACCATCAAGCCATTTTATGAAAC/GCGCGAATTCGTCGGCTGCGATTAGTCTG [9] scrG GCGCGTCGACTAGCACGCTTGTGTTGGAC/GCGCGAATTCCAGGGAAATGAAGTAATCATGC VP0117 GCGCTCTAGACTCACACAACACTTTCTG/GCGCGAATTCAGACAATCACACCGATAG VPA0198 GCGCGTCGACCTCTGGTTCATTGTCTTG/GCGCGAATTCGTCTTGCTGATTCTGATG VPA1176 AAAGTCGACTCAGGTACGCTTGCTTCAC/GGGGAATTCCGTTGCTTGGTAGTGGTAATAG VP0699 GCGCGTCGACGGAGAATACCTAGCAGAG/GCGCGAATTCAGTATCACGATGTGTCAG VP2979 GCGCTCTAGATTTCTATCCGTTGGCTAC/GCGCGAATTCCTGACTTACATCGTGGAC EMSA scrA GAGCGTATATCCAAGTGGTTTG/GTCGGCTGCGATTAGTCTG [9] scrG TAGCACGCTTGTGTTGGAC/CAGGGAAATGAAGTAATCATGC VP0117 CTCACACAACACTTTCTG/GCGCGAATTCAGACAATCACACCGATAG VPA0198 CTCTGGTTCATTGTCTTG/GTCTTGCTGATTCTGATG VPA1176 TCAGGTACGCTTGCTTCAC/CGTTGCTTGGTAGTGGTAATAG VP0699 GGAGAATACCTAGCAGAG/AGTATCACGATGTGTCAG VP2979 TTTCTATCCGTTGGCTAC/CTGACTTACATCGTGGAC 16S rDNA GACACGGTCCAGACTCCTAC/GGTGCTTCTTCTGTCGCTAAC Note. EMSA: electrophoretic mobility shift assay. -
The V. parahaemolyticus strains were similarly cultured as described previously [9]. Briefly, V. parahaemolyticus was cultivated overnight (about 12 h) in 2.5% (w/v) Bacto heart infusion (HI) broth (BD Biosciences, USA) at 37 °C with shaking at 200 rpm. The resultant cultures were diluted 40-fold into the phosphate-buffered saline buffer (PBS, pH 7.2). Then, 150 µL of the diluted cultures were spread onto the HI plate with a diameter of 5 cm. After incubation at 37 °C for 6 h, the bacterial cells were harvested by adding 2 mL pre-cold PBS. When necessary, the media were supplemented with 100 µg/mL gentamicin, 5 µg/mL chloramphenicol, or 0.1% arabinose.
-
The CV staining was carried out as previously described [9, 10]. Briefly, overnight cell cultures were diluted 50-fold into 5 mL fresh HI broth and cultured at 37 °C with shaking at 200 rpm and OD600 of approximately 1.4. Thereafter, the resultant cultures were 50-fold diluted into 2 mL fresh Difco marine (M) broth 2216 (BD Biosciences, USA) containing 0.1% arabinose and 5 µg/mL chloramphenicol in glass tubes and cultured at 30 °C with shaking at 150 rpm for 48 h. Media with planktonic cells were collected for the measurement of OD600 values. The surface-attached cells were stained with 0.1% CV. The bound CV was dissolved with 20% ethanol, and the OD570 values were measured. The relative biofilm formation was calculated with the formula: 100 × OD570/OD600.
-
The intracellular c-di-GMP levels were measured similarly as previously described [22]. Briefly, V. parahaemolyticus was grown on HI plates at 37 °C for 6 h, and the bacterial cells were harvested by adding 2 mL pre-chilled PBS. Two OD600 bacterial suspensions were centrifuged at 10,000 ×g for 3 min at 4 °C. The cell pellet was washed twice with ice-cold PBS, resuspended in 2 mL ice-cold PBS, and then sonicated for 30 min (power 100%, frequency 37 kHz) in an ice-water bath. After centrifugation at 10,000 ×g for 10 min at 4 °C, the supernatant containing extracted c-di-GMP was collected to determine the intracellular c-di-GMP concentration with a c-di-GMP enzyme-linked immunosorbent assay kit (Mskbio, Beijing, China). Total proteins in the cell extracts were also determined by a Micro BCA Protein Assay Kit (ThermoFisher Scientific, USA), in accordance with the manufacturer’s instructions. The intracellular c-di-GMP concentration was expressed as pmol/mg of protein.
-
The qPCR assay was performed as previously described [23]. Briefly, the total RNAs were extracted from the WT and Δhns strains, and the cDNAs were generated from 1 µg total RNAs using the FastKing First-Strand cDNA Synthesis Kit (Tiangen Biotech, Beijing, China) following the manufacturer’s instructions. The relative mRNA levels of each target gene were determined using the classic 2−ΔΔCt method.
-
The promoter–proximal DNA region of each target gene was amplified and cloned into the pHRP309 vector containing a promoterless lacZ gene and a gentamicin resistance gene [24]. After verification by DNA sequencing, the recombinant pHRP309 was transferred to the WT and Δhns strains, respectively. Thereafter, the transformants were cultured and lysed to measure the β-galactosidase activity in the cell extracts using the β-Galactosidase Enzyme Assay System (Promega, USA), in accordance with the manufacturer’s instructions.
The two-plasmid reporter assay in E. coli was performed as previously described [25]. Briefly, the E. coli 100 λpir (Epicenter) bearing the indicated hns expression plasmid pBAD33-hns or the empty pBAD33 vector and a recombinant lacZ plasmid was cultivated overnight in Luria–Bertani (LB) broth at 37 °C with shaking at 200 rpm. The overnight cultures were diluted 1:100 into 5 mL fresh LB broth containing 0.1% arabinose and 20 µg/mL chloramphenicol and incubated at 37 °C with shaking at 200 rpm to reach an OD600 value of about 1.2. The E. coli cells were harvested and lysed to measure the β-galactosidase activity in the cell extracts.
-
The coding region of hns was amplified and cloned into the pET28a vector (Novagen, USA) and then transferred to E. coli BL21λDE3 to express 6 × His-tagged H-NS (His-H-NS). The expression and purification of His-H-NS were performed as previously described [17]. The dialyzed His-H-NS was concentrated to a final concentration of 1.4 mg/mL. EMSA was performed exactly as previously described [26]. The results of EMSA were detected by ethidium bromide (EB) staining, and the gel images were displayed with an ultraviolet (UV) transilluminator.
-
The EMSA was performed at least twice with similar results. The CV staining, c-di-GMP quantification, LacZ fusion, and qPCR were performed at least thrice with three replications each, with values expressed as the means ± standard deviation (SD). Analysis of variance with the Tukey-Kramer test was applied to calculate statistical significance, with P < 0.01 considered significant.
-
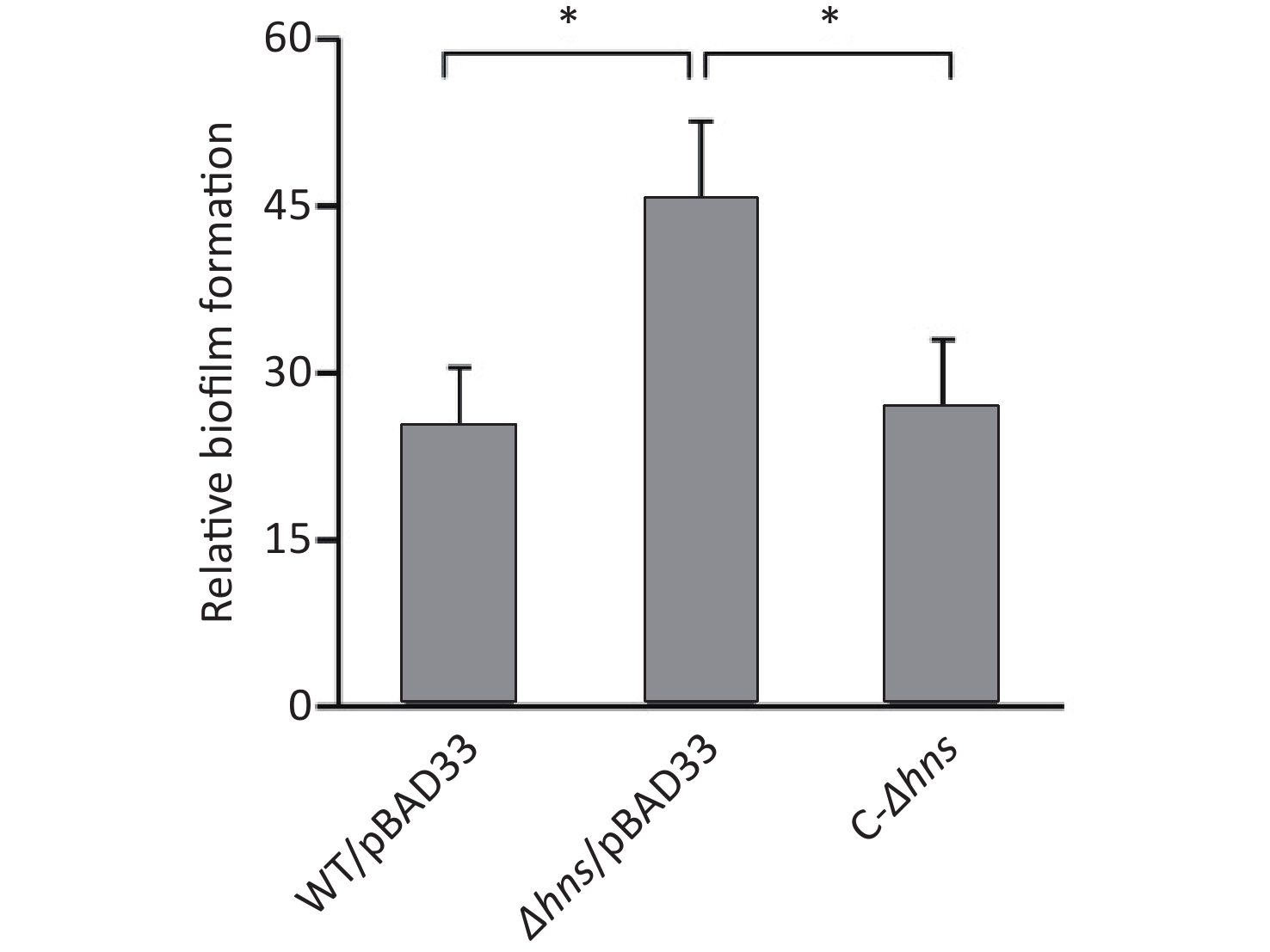
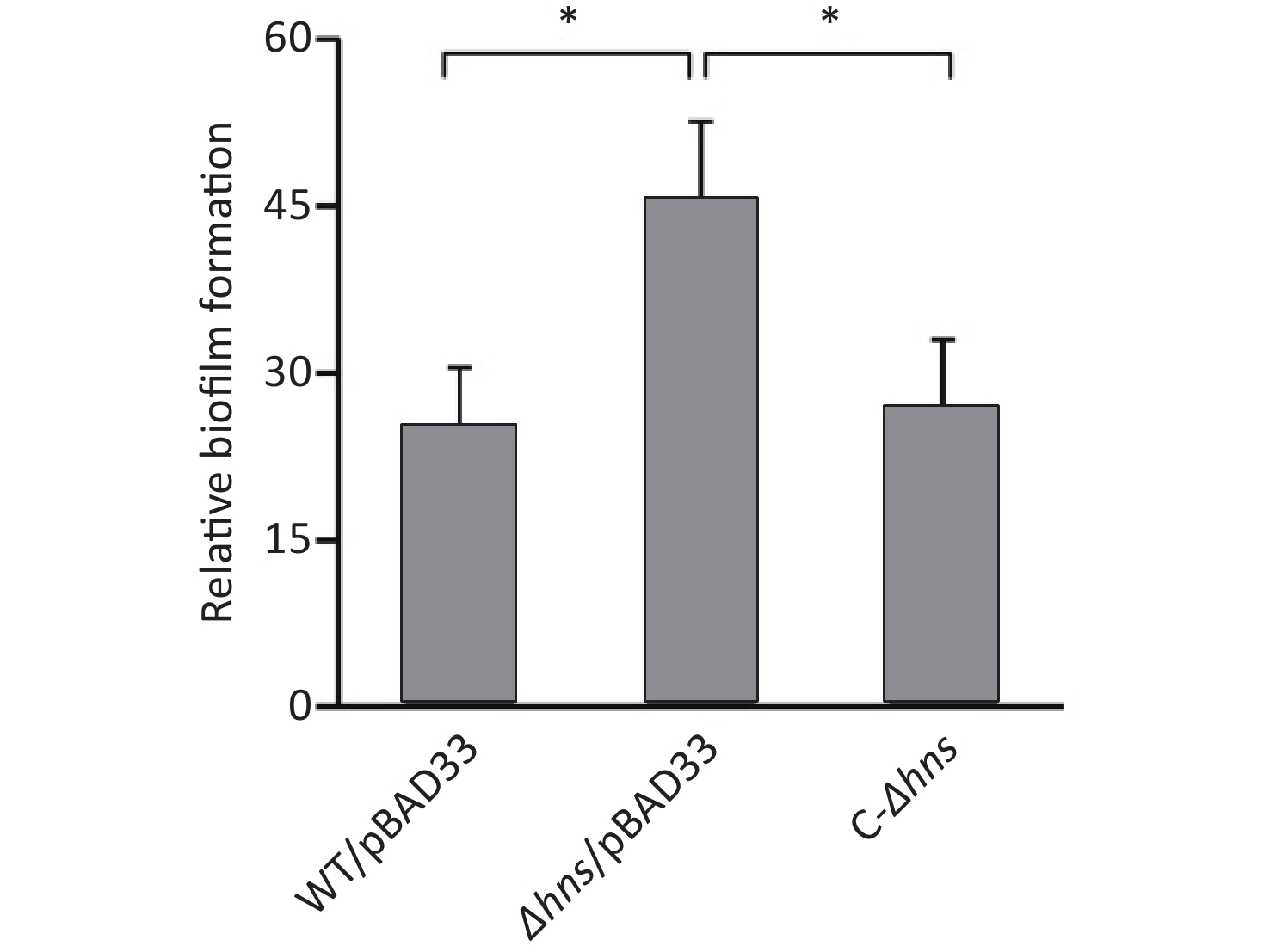
We previously reported that H-NS activated the EPS-dependent bacterial colony morphology and the transcription cpsA-J in V. parahaemolyticus RIMD2210633 [8]. In this work, we investigated whether H-NS regulated biofilm formation by V. parahaemolyticus RIMD2210633 using the CV staining assay. As shown in Figure 1, the normalized CV staining by Δhns/pBAD33 significantly increased relative to that by WT/pBAD33, and C-Δhns produced a restored CV staining. This result indicated that H-NS repressed biofilm formation by V. parahaemolyticus RIMD2210633, as assessed by CV staining.
-
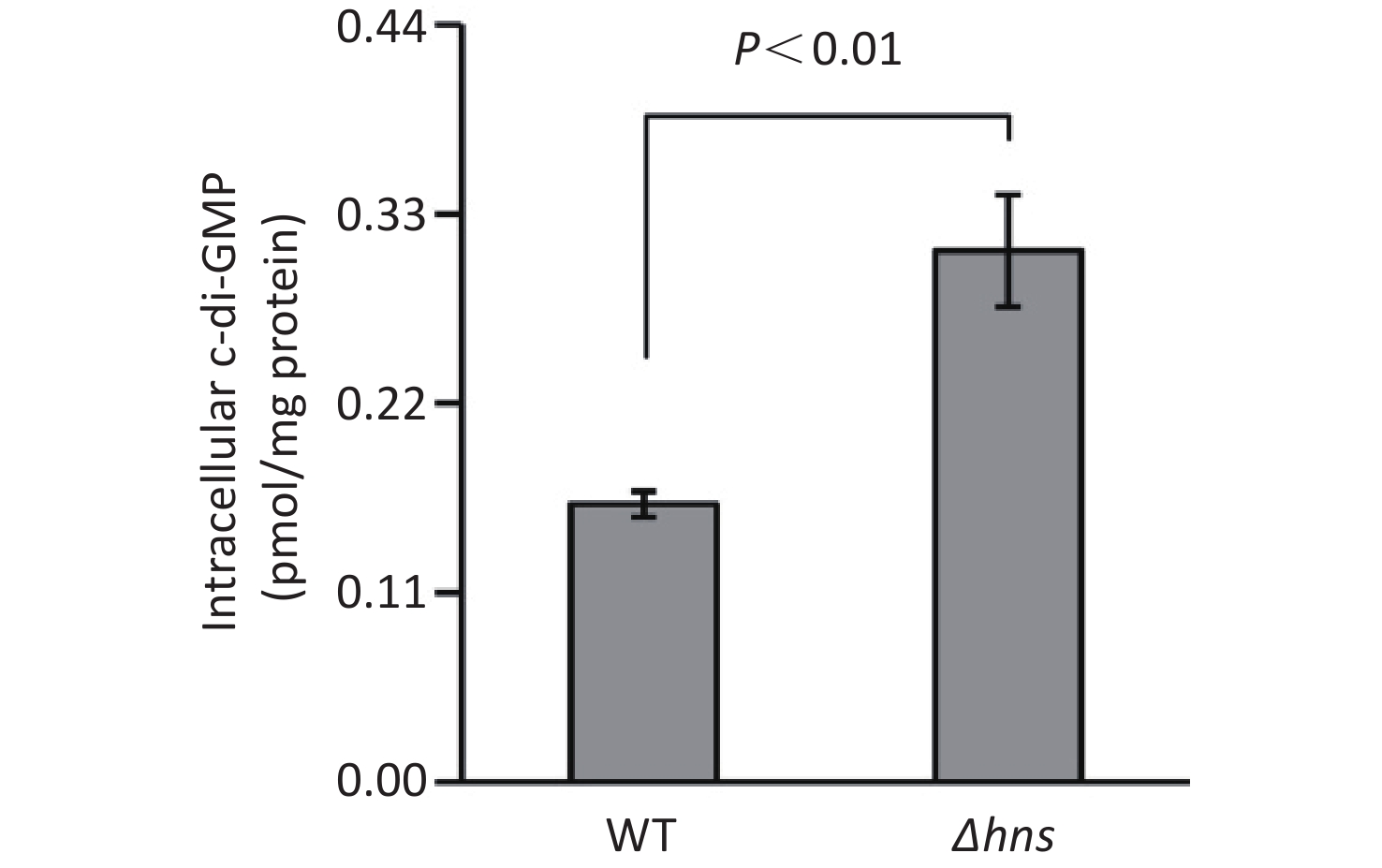
An increased c-di-GMP level leads to enhanced biofilm formation in bacteria [2]. Thus, the increased CV staining by Δhns prompted us to investigate whether H-NS affects the production of c-di-GMP in V. parahaemolyticus RIMD2210633. As shown in Figure 2, the intracellular c-di-GMP level was significantly enhanced in Δhns relative to that in WT, indicating that H-NS inhibited the synthesis of c-di-GMP in V. parahaemolyticus RIMD2210633.
-
The transcription of a group of genes encoding proteins with GGDEF and/or EAL domains, namely, scrABC, scrG, VP0117, VPA0198, VPA1176, VP0699, and VP2979, were directly regulated by the QS master regulator OpaR [9]. H-NS is also a transcriptional repressor of scrABC [8], but whether H-NS can directly bind scrABC promoters is still unknown. In this study, we investigated the regulatory mechanisms of H-NS on the transcription of these seven genes. The results of qPCR demonstrated that the mRNA levels of all of the seven genes significantly increased in Δhns relative to those in WT (Figure 3A), suggesting that their transcription was repressed by H-NS. The upstream DNA region of each of the seven genes was cloned into the pHRP309 plasmid containing a gentamicin resistance gene and a promoterless lacZ gene and then transferred into Δhns and WT to investigate the β-galactosidase activities in the cell extracts. As shown in Figure 3B, the promoter activities of all the seven genes were considerably higher in Δhns than those in WT, suggesting that the expressions of these genes were under the negative control of H-NS. Altogether, H-NS acted as a repressor of scrABC, scrG, VP0117, VPA0198, VPA1176, VP0699, and VP2979 in V. parahaemolyticus RIMD2210633.
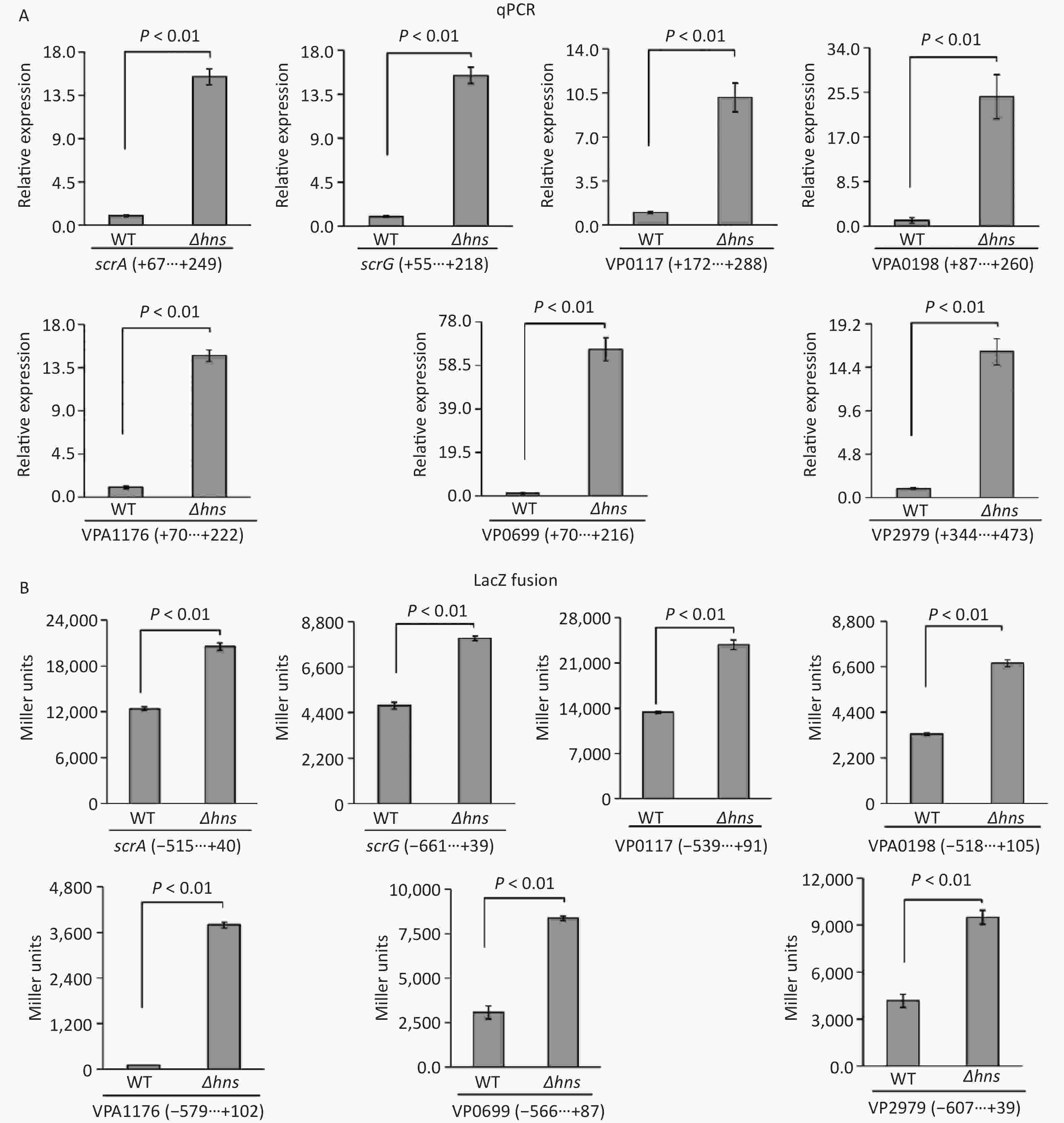
Figure 3. H-NS repressed the transcription of the seven genes associated with c-di-GMP metabolism in V. parahaemolyticus. The negative and positive numbers indicate the nucleotide positions upstream and downstream of indicated genes, respectively. (A) qPCR. After incubation at 37 °C for 6 h on HI plates, the bacterial cells were harvested, and total RNA was extracted using TRIzol Reagent. qPCR assay was used to determine the relative mRNA levels for target genes in Δhns and WT using the classical 2−ΔΔCt method. (B) LacZ fusion. The promoter DNA region of each target gene was cloned into the pHRP309 and then transferred into Δhns and WT to test the promoter activity in cellular extracts.
-
To test whether H-NS can control the expressions of the seven target genes in a heterologous host, we expressed H-NS from the pBAD33 vector in E. coli 100 λpir, which also contained a lacZ fusion reporter plasmid [25]. As shown in Figure 4, after induction by the addition of 0.2% arabinose in the culture media, the promoter activities for all of the seven target genes in the E. coli 100λpir containing the recombinant pBAD33-hns were significantly lower than those in E. coli 100λpir containing the empty pBAD33 vector. These results suggested that H-NS can bind the promoter DNA region of each of the seven target genes to repress their expression.

Figure 4. Two-plasmid reporter assay in E. coli. The E. coli 100λpir containing the pBAD33 plasmid or recombinant pBAD33-hns plasmid and a LacZ fusion reporter plasmid were grown in LB broth containing 0.2% arabinose until the mid-log phase (OD600 = 1.2). The bacterial cells were collected and assayed for lacZ expression using the β-galactosidase assay. The negative and positive numbers indicate the nucleotide positions upstream and downstream of indicated genes, respectively.
-
The EMSA was employed to detect whether His-H-NS had binding activities to the regulatory DNA regions of the seven target genes in vitro. As shown in Figure 5, His-H-NS can bind the regulatory DNA regions of all of the seven target genes in dose-dependent manners, but it cannot bind the DNA fragment of 16S rDNA, which was used as the negative control. These results further suggested that H-NS directly inhibited the transcription of scrABC, scrG, VP0117, VPA0198, VPA1176, VP0699, and VP2979 in V. parahaemolyticus RIMD2210633.
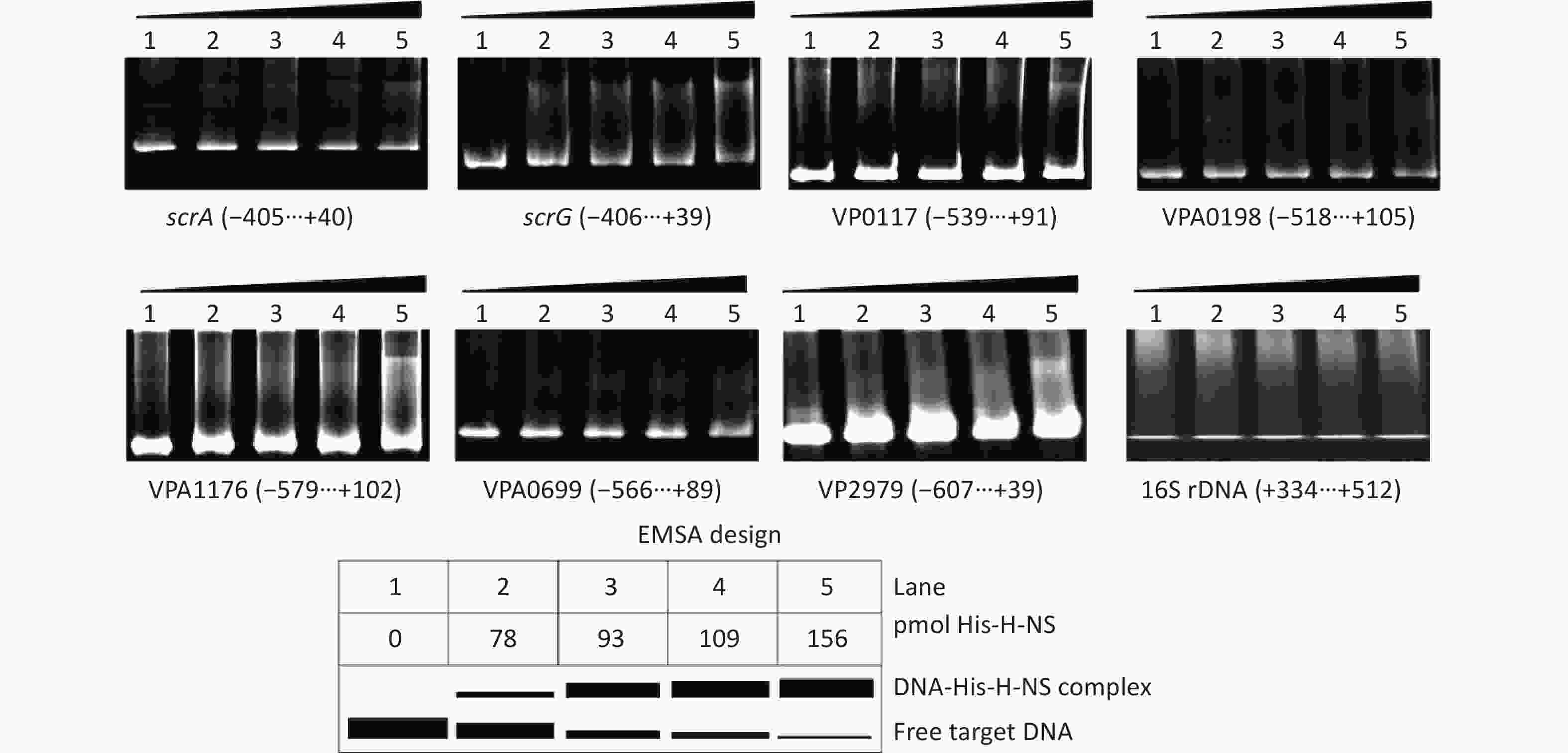
Figure 5. Binding of His-H-NS to the target DNA fragments. The regulatory DNA region of each target gene was incubated with increased amounts of His-H-NS and then subjected to 6% (w/v) polyacrylamide gel electrophoresis. The results were detected by EB staining and a UV transilluminator. A schematic representation of the EMSA design is shown below. The negative and positive numbers denote the nucleotide positions upstream and downstream of indicated genes, respectively.
-
H-NS has been shown to be a positive regulator of biofilm formation by V. parahaemolyticus BB22 because the deletion of hns forms defective submerged biofilms relative to the WT strain [20]. The hns mutant of V. parahaemolyticus RIMD2210633 formed smooth colonies and expressed low cpsA-J, indicating that H-NS may also act as a positive regulator of biofilm formation by V. parahaemolyticus RIMD2210633 [8]. However, the data on CV staining presented here showed that H-NS repressed biofilm formation by V. parahaemolyticus RIMD2210633 (Figure 1). This result is not only contrary to the previous observations on V. parahaemolyticus BB22 but also V. parahaemolyticus RIMD2210633 [8, 20]. The genetic information contained in the genomes of BB22 and RIMD2210633 are different, and RIMD2210633 contains many unique genes relative to BB22 [27]. Thus, the discrepancy between the data presented here and the previous conclusion regarding BB22 may be explained by these genetic differences. However, why the results of CV staining and colony morphology are contrary to each other in RIMD2210633 is difficult to understand. Colony morphology is mainly attributed to the EPS content, which is associated with the expression of cpsA-J [3]. However, a previous study showed that the biofilm formation by the V. parahaemolyticus trh positive strain ATCC17802 was strongly correlated with extracellular DNA and protein content in the extracellular polymeric substances but not with the EPS content [28]. Thus, similar observations may be obtained with V. parahaemolyticus RIMD2210633. Such a condition may be also due to the global influence of H-NS [16]. Specifically, studies showed that H-NS induced opaR expression and polar flagellum production but repressed the transcription of lateral flagellar genes in V. parahaemolyticus RIMD2210633 [19, 29, 30]. Flagella-mediated motilities promote not only the initial adhesion of biofilm formation but also the active dispersal of biofilms [31]; OpaR is an inhibitor of biofilm formation by V. parahaemolyticus TIMD2210633 [9]. OpaR may have some effects on the H-NS-mediated inhibition of biofilm formation in V. parahaemolyticus RIMD2210633. Although regulatory mechanisms may be different, the negative regulation of H-NS on biofilm formation has been confirmed in V. cholerae [32, 33].
The increased c-di-GMP content results in enhanced biofilm formation and motility inhibition [2]. The data presented here showed that the deletion of hns significantly increased the intracellular c-di-GMP level relative to that in the WT strain (Figure 2). The genome of V. parahaemolyticus RIMD2210633 contains more than 50 genes encoding proteins that may be associated with the c-di-GMP metabolism [21], among which the transcriptions of scrABC, scrG, VP0117, VPA0198, VPA1176, VP0699, and VP2979 are directly regulated by the master QS regulator OpaR [9]. Here, we showed that H-NS can bind the promoter–proximal DNA regions of scrA, scrG, VP0117, VPA0198, VPA1176, VP0699, and VP2979 to repress their transcription (Figures 3, 4, and 5). Notably, OpaR showed the reverse regulation of scrG and VP2979 compared with those of H-NS regulation [9]; OpaR and H-NS exerted the same effects on c-di-GMP production. However, such findings should be easy to understand because the effect degrees of dozens of putative enzymes on intracellular c-di-GMP metabolism should be inconsistent under various genetic backgrounds (or growth conditions). Except for scrA and scrG, the functions of the other 5 target genes are completely unknown, and the data presented here showed that H-NS repression of the intracellular c-di-GMP levels in V. parahaemolyticus may occur via the direct regulation of the transcription of enzyme genes involved in the synthesis of c-di-GMP. The H-NS binding sites are usually long and overlap with the core −10 and −35 elements [17, 19]. Thus, H-NS is usually thought to repress gene transcription by directly interfering with the action of RNA polymerase.
Altogether, this work reported that H-NS was a negative regulator of the biofilm formation by V. parahaemolyticus RIMD2210633. H-NS directly repressed the transcription of scrA, scrG, VP0117, VPA0198, VPA1176, VP0699, and VP2979, which encode a group of GGDEF and/or EAL type proteins and thus decrease the intracellular c-di-GMP level. Therefore, H-NS repression of the biofilm formation by V. parahaemolyticus RIMD2210633 may occur via the repression of the production of intracellular c-di-GMP. However, the regulation mechanisms of H-NS on the biofilm formation by V. parahaemolyticus still need to be further studied.
doi: 10.3967/bes2022.106
-
Abstract:
Objective This study aimed to investigate the regulation of histone-like nucleoid structuring protein (H-NS) on biofilm formation and cyclic diguanylate (c-di-GMP) synthesis in Vibrio parahaemolyticus RIMD2210633. Methods Regulatory mechanisms were analyzed by the combined utilization of crystal violet staining, quantification of c-di-GMP, quantitative real-time polymerase chain reaction, LacZ fusion, and electrophoretic-mobility shift assay. Results The deletion of hns enhanced the biofilm formation and intracellular c-di-GMP levels in V. parahaemolyticus RIMD2210633. H-NS can bind the upstream promoter–proximal DNA regions of scrA, scrG, VP0117, VPA0198, VPA1176, VP0699, and VP2979 to repress their transcription. These genes encode a group of proteins with GGDEF and/or EAL domains associated with c-di-GMP metabolism. Conclusion One of the mechanisms by which H-NS represses the biofilm formation by V. parahaemolyticus RIMD2210633 may be via repression of the production of intracellular c-di-GMP. -
Key words:
- Vibrio parahaemolyticus /
- Biofilm /
- H-NS /
- C-di-GMP
注释: -
Figure 3. H-NS repressed the transcription of the seven genes associated with c-di-GMP metabolism in V. parahaemolyticus. The negative and positive numbers indicate the nucleotide positions upstream and downstream of indicated genes, respectively. (A) qPCR. After incubation at 37 °C for 6 h on HI plates, the bacterial cells were harvested, and total RNA was extracted using TRIzol Reagent. qPCR assay was used to determine the relative mRNA levels for target genes in Δhns and WT using the classical 2−ΔΔCt method. (B) LacZ fusion. The promoter DNA region of each target gene was cloned into the pHRP309 and then transferred into Δhns and WT to test the promoter activity in cellular extracts.
Figure 4. Two-plasmid reporter assay in E. coli. The E. coli 100λpir containing the pBAD33 plasmid or recombinant pBAD33-hns plasmid and a LacZ fusion reporter plasmid were grown in LB broth containing 0.2% arabinose until the mid-log phase (OD600 = 1.2). The bacterial cells were collected and assayed for lacZ expression using the β-galactosidase assay. The negative and positive numbers indicate the nucleotide positions upstream and downstream of indicated genes, respectively.
Figure 5. Binding of His-H-NS to the target DNA fragments. The regulatory DNA region of each target gene was incubated with increased amounts of His-H-NS and then subjected to 6% (w/v) polyacrylamide gel electrophoresis. The results were detected by EB staining and a UV transilluminator. A schematic representation of the EMSA design is shown below. The negative and positive numbers denote the nucleotide positions upstream and downstream of indicated genes, respectively.
Table 1. Oligonucleotide primers used in this study
Target Primers (forward/reverse, 5'–3') Reference Protein expression hns AGCGGGATCCATGTCAGAGCTGACTAAAACAC/AGCGAAGCTTTTAGATTAGGAAATCGTCTAG [17] qPCR scrA CACACCACGAACACATTGC/TCAATAGCGTCACGGAATGC [9] scrG AAGCCGTGGTGGAAGAAGG/GCGTGTTGAGTGCGTTGG VP0117 GACCACCTCAATAGTTATCTG/TAAGTAGGCTTGGACATCTC VPA0198 GCATCAGAATCAGCAAGAC/ATGCTTAGCTCCTCTTCTTC VPA1176 GCCATATTCCAAACTCGTTGTG/TGCGTAAGCCAAGTTGATGAG VP0699 CTGACACATCGTGATACTTC/TTGATGTTGCAGCTCTTG VP2979 GCAACTCTCAAGTCATCATC/CAACAACCGTCTTCTATGG LacZ fusion scrA GCGCGTCGACCATCAAGCCATTTTATGAAAC/GCGCGAATTCGTCGGCTGCGATTAGTCTG [9] scrG GCGCGTCGACTAGCACGCTTGTGTTGGAC/GCGCGAATTCCAGGGAAATGAAGTAATCATGC VP0117 GCGCTCTAGACTCACACAACACTTTCTG/GCGCGAATTCAGACAATCACACCGATAG VPA0198 GCGCGTCGACCTCTGGTTCATTGTCTTG/GCGCGAATTCGTCTTGCTGATTCTGATG VPA1176 AAAGTCGACTCAGGTACGCTTGCTTCAC/GGGGAATTCCGTTGCTTGGTAGTGGTAATAG VP0699 GCGCGTCGACGGAGAATACCTAGCAGAG/GCGCGAATTCAGTATCACGATGTGTCAG VP2979 GCGCTCTAGATTTCTATCCGTTGGCTAC/GCGCGAATTCCTGACTTACATCGTGGAC EMSA scrA GAGCGTATATCCAAGTGGTTTG/GTCGGCTGCGATTAGTCTG [9] scrG TAGCACGCTTGTGTTGGAC/CAGGGAAATGAAGTAATCATGC VP0117 CTCACACAACACTTTCTG/GCGCGAATTCAGACAATCACACCGATAG VPA0198 CTCTGGTTCATTGTCTTG/GTCTTGCTGATTCTGATG VPA1176 TCAGGTACGCTTGCTTCAC/CGTTGCTTGGTAGTGGTAATAG VP0699 GGAGAATACCTAGCAGAG/AGTATCACGATGTGTCAG VP2979 TTTCTATCCGTTGGCTAC/CTGACTTACATCGTGGAC 16S rDNA GACACGGTCCAGACTCCTAC/GGTGCTTCTTCTGTCGCTAAC Note. EMSA: electrophoretic mobility shift assay. -
[1] Broberg CA, Calder TJ, Orth K. Vibrio parahaemolyticus cell biology and pathogenicity determinants. Microbes Infect, 2011; 13, 992−1001. doi: 10.1016/j.micinf.2011.06.013 [2] Yildiz FH, Visick KL. Vibrio biofilms: so much the same yet so different. Trends Microbiol, 2009; 17, 109−18. doi: 10.1016/j.tim.2008.12.004 [3] Chen YS, Dai JL, Morris JG Jr, et al. Genetic analysis of the capsule polysaccharide (K antigen) and exopolysaccharide genes in pandemic Vibrio parahaemolyticus O3: K6. BMC Microbiol, 2010; 10, 274. doi: 10.1186/1471-2180-10-274 [4] Ferreira RB, Antunes LCM, Greenberg EP, et al. Vibrio parahaemolyticus ScrC modulates cyclic dimeric GMP regulation of gene expression relevant to growth on surfaces. J Bacteriol, 2008; 190, 851−60. doi: 10.1128/JB.01462-07 [5] Ferreira RB, Chodur DM, Antunes LCM, et al. Output targets and transcriptional regulation by a cyclic dimeric GMP-responsive circuit in the Vibrio parahaemolyticus scr network. J Bacteriol, 2012; 194, 914−24. doi: 10.1128/JB.05807-11 [6] Kim YK, McCarter LL. ScrG, a GGDEF-EAL protein, participates in regulating swarming and sticking in Vibrio parahaemolyticus. J Bacteriol, 2007; 189, 4094−107. doi: 10.1128/JB.01510-06 [7] Güvener ZT, McCarter LL. Multiple regulators control capsular polysaccharide production in Vibrio parahaemolyticus. J Bacteriol, 2003; 185, 5431−41. doi: 10.1128/JB.185.18.5431-5441.2003 [8] Zhang LL, Weng YW, Wu Y, et al. H-NS is an activator of exopolysaccharide biosynthesis genes transcription in Vibrio parahaemolyticus. Microb Pathog, 2018; 116, 164−7. doi: 10.1016/j.micpath.2018.01.025 [9] Zhang YQ, Qiu Y, Gao H, et al. OpaR controls the metabolism of c-di-GMP in Vibrio parahaemolyticus. Front Microbiol, 2021; 12, 676436. doi: 10.3389/fmicb.2021.676436 [10] Wang L, Ling Y, Jiang HW, et al. AphA is required for biofilm formation, motility, and virulence in pandemic Vibrio parahaemolyticus. Int J Food Microbiol, 2013; 160, 245−51. doi: 10.1016/j.ijfoodmicro.2012.11.004 [11] Biswas S, Chouhan OP, Bandekar D. Diguanylate cyclases in Vibrio cholerae: essential regulators of lifestyle switching. Front Cell Infect Microbiol, 2020; 10, 582947. doi: 10.3389/fcimb.2020.582947 [12] Jenal U, Reinders A, Lori C. Cyclic di-GMP: second messenger extraordinaire. Nat Rev Microbiol, 2017; 15, 271−84. doi: 10.1038/nrmicro.2016.190 [13] Kimbrough JH, McCarter LL. Identification of three new GGDEF and EAL domain-containing proteins participating in the scr surface colonization regulatory network in Vibrio parahaemolyticus. J Bacteriol, 2021; 203, e00409−20. [14] Kimbrough JH, Cribbs JT, McCarter LL. Homologous c-di-GMP-binding scr transcription factors orchestrate biofilm development in Vibrio parahaemolyticus. J Bacteriol, 2020; 202, e00723−19. [15] Zhong XJ, Lu Z, Wang F, et al. Characterization of GefA, a GGEEF domain-containing protein that modulates Vibrio parahaemolyticus motility, biofilm formation, and virulence. Appl Environ Microbiol, 2022; 88, e0223921. doi: 10.1128/aem.02239-21 [16] Grainger DC. Structure and function of bacterial H-NS protein. Biochem Soc Trans, 2016; 44, 1561−9. doi: 10.1042/BST20160190 [17] Sun FJ, Zhang YQ, Qiu YF, et al. H-NS is a repressor of major virulence gene loci in Vibrio parahaemolyticus. Front Microbiol, 2014; 5, 675. [18] Salomon D, Klimko JA, Orth K. H-NS regulates the Vibrio parahaemolyticus type VI secretion system 1. Microbiology (Reading), 2014; 160, 1867−73. doi: 10.1099/mic.0.080028-0 [19] Wang Y, Zhang YQ, Yin Z, et al. H-NS represses transcription of the flagellin gene lafA of lateral flagella in Vibrio parahaemolyticus. Can J Microbiol, 2018; 64, 69−74. doi: 10.1139/cjm-2017-0315 [20] Enos-Berlage JL, Guvener ZT, Keenan CE, et al. Genetic determinants of biofilm development of opaque and translucent Vibrio parahaemolyticus. Mol Microbiol, 2005; 55, 1160−82. [21] Makino K, Oshima K, Kurokawa K, et al. Genome sequence of Vibrio parahaemolyticus: a pathogenic mechanism distinct from that of V cholerae. Lancet, 2003; 361, 743−9. doi: 10.1016/S0140-6736(03)12659-1 [22] Gao H, Ma LZ, Qin Q, et al. Fur represses Vibrio cholerae biofilm formation via direct regulation of vieSAB, cdgD, vpsU, and vpsA-K transcription. Front Microbiol, 2020; 11, 587159. doi: 10.3389/fmicb.2020.587159 [23] Gao H, Zhang YQ, Yang L, et al. Regulatory effects of cAMP receptor protein (CRP) on porin genes and its own gene in Yersinia pestis. BMC Microbiol, 2011; 11, 40. doi: 10.1186/1471-2180-11-40 [24] Parales RE, Harwood CS. Construction and use of a new broad-host-range lacZ transcriptional fusion vector, pHRP309, for gram- bacteria. Gene, 1993; 133, 23−30. doi: 10.1016/0378-1119(93)90220-W [25] Qiu Y, Hu LF, Yang WH, et al. The type VI secretion system 2 of Vibrio parahaemolyticus is regulated by QsvR. Microb Pathog, 2020; 149, 104579. doi: 10.1016/j.micpath.2020.104579 [26] Zhang LY, Osei-Adjei G, Zhang Y, et al. CalR is required for the expression of T6SS2 and the adhesion of Vibrio parahaemolyticus to HeLa cells. Arch Microbiol, 2017; 199, 931−8. doi: 10.1007/s00203-017-1361-6 [27] Jensen RV, DePasquale SM, Harbolick EA, et al. Complete genome sequence of prepandemic Vibrio parahaemolyticus BB22OP. Genome Announc, 2013; 1, e00002−12. [28] Li W, Wang JJ, Qian H, et al. Insights into the role of extracellular DNA and extracellular proteins in biofilm formation of Vibrio parahaemolyticus. Front Microbiol, 2020; 11, 813. doi: 10.3389/fmicb.2020.00813 [29] Zhang YQ, Zhang LY, Hou SN, et al. The master quorum-sensing regulator OpaR is activated indirectly by H-NS in Vibrio parahaemolyticus. Curr Microbiol, 2016; 73, 71−6. doi: 10.1007/s00284-016-1018-8 [30] Wang J, Lin L, Sun FJ, et al. Regulation of swimming motility by H-NS in Vibrio parahaemolyticus. Mil Med Sci, 2015; 9, 694−7. (In Chinese [31] Ashrafudoulla M, Mizan FR, Park SH, et al. Current and future perspectives for controlling Vibrio biofilms in the seafood industry: a comprehensive review. Crit Rev Food Sci Nutr, 2021; 61, 1827−51. doi: 10.1080/10408398.2020.1767031 [32] Wang HX, Ayala JC, Silva AJ, et al. The histone-like nucleoid structuring protein (H-NS) is a repressor of Vibrio cholerae exopolysaccharide biosynthesis (vps) genes. Appl Environ Microbiol, 2012; 78, 2482−8. doi: 10.1128/AEM.07629-11 [33] Ayala JC, Wang HX, Silva AJ, et al. Repression by H-NS of genes required for the biosynthesis of the Vibrio cholerae biofilm matrix is modulated by the second messenger cyclic diguanylic acid. Mol Microbiol, 2015; 97, 630−45. doi: 10.1111/mmi.13058 -




 下载:
下载:



 Quick Links
Quick Links