-
Osteosarcoma, which invades and destroys bone and adjacent soft tissues, is the most common malignant bone tumor diagnosed in children and adolescents[1, 2]. Despite significant advances made in surgery-and multi-agent chemotherapy-based treatments, nearly 30% of patients still die from osteosarcoma. The international incidence of osteosarcoma has been rising and the 5-year patient survival rate of 65%-70% has already plateaued[3, 4]. Therefore, investigations on osteosarcoma pathogenesis, including mechanisms of cell proliferation, are required to aid in the development of novel therapeutic approaches.
Oncogenes and tumor suppressors are typically regulated through post-translational modifications, such as those mediated by the ubiquitin-proteasome sys tem[5]. Currently, E3 ligases involved in the regulation of oncogenes and tumor suppressor proteins are a focus of tumor research[6]. The evolutionarily conserved tripartite motif (TRIM) family of proteins are composed of more than 70 known members in humans and mice, and members are involved in a wide range of biological processes and diverse pathological conditions, including developmental disorders, neurodegenerative diseases, viral infections, and cancers[7]. Most TRIM members contain a RING-finger domain and, thus, function as E3 ubiquitin ligases[6]. In addition, several TRIM members are involved in oncogenic processes, such as transcriptional regulation, cell proliferation, and apoptosis[6]. Unlike other ubiquitin ligases that function through the formation of multisubunit complexes, TRIM proteins function as E3 ubiquitin ligases in the form of single component[8]. In addition to the presence of a RING-finger domain in the N-terminus of nearly all TRIM proteins, one or two B-box domains and a coiled-coil domain are also found in the majority of TRIM family members[6].
Tripartite motif 21 (TRIM21), i.e., Ro52 or RNF81, is a member of the TRIM family with E3 ligase activity[9]. Previous studies on TRIM21 have mainly focused on its role in the innate immune system, including in neutralizing viral infection[10] and regulating inflammatory signaling pathways[11]. The functions of TRIM21 in tumor biology have gradually been gaining attention. Recent studies identified TRIM21 as a novel interacting partner of the tumor-suppressor prostate apoptosis response protein 4 (Par-4) in colon and pancreatic cancer cells[12]. TRIM21 has been shown to downregulate Par-4 levels in response to cisplatin and protect colon cancer cells from cisplatin-induced apoptosis[12]. Along these lines, high TRIM21 expression levels correlate with reduced overall and disease-free survival in patients with pancreatic cancer[12]. Conversely, TRIM21 has also been reported to interact with and negatively regulate the protein levels of Lifeguard, which protects neurons against Fas-induced apoptosis[13]. Furthermore, TRIM21 regulates the oncogenic NF-κB signaling pathway by mono-ubiquitinating IKKβ[14], as well as positively regulates apoptosis by promoting degradation of Bcl-2 and c-FLIP[15-17]. TRIM21 reduces B cell proliferation[18] and knockdown of TRIM21 increases proliferation of the hepatocellular carcinoma (HCC) cell lines HepG2 and LM3[19]. In addition, reduced TRIM21 expression is correlated with a poor prognosis in patients with HCC or diffuse large B-cell lymphoma[19, 20]. Therefore, TRIM21 protein plays various roles in tumorigenesis, but its functions in osteosarcoma cell proliferation have not yet been reported.
YWHAZ, i.e., 14-3-3ζ, belongs to the 14-3-3 family and functions as an oncogene in tumors by interacting with multiple proteins involved in diverse biological processes, including regulation of cell proliferation, apoptosis, cell cycle, and DNA damage[21, 22]. YWHAZ is overexpressed in various tumors and is a promising prognostic marker for breast, prostate, gastric, head and neck cancers, HCC, lung adenocarcinoma, and adenocarcinoma of the esophagogastric junction[21, 23-27]. However, there have been few reports on the role of YWHAZ in osteosarcoma.
In the present study, TRIM21 was found to enhance osteosarcoma cell viability. By using co-immunoprecipitation (co-IP) in combination with mass-spectrometry, YWHAZ protein was identified as a novel interacting partner of TRIM21. Furthermore, TRIM21 E3 ligase was determined to be a negative regulator of YWHAZ expression levels. Surprisingly, YWHAZ protein overexpression did not dampen the effect of TRIM21 on osteosarcoma cell proliferation.
-
Human osteosarcoma U2-OS and Saos-2 cells (Institute of Life Science Chinese Academy of Sciences, Shanghai, China) were cultured in McCoy's 5A medium (Sigma-Aldrich, St Louis, MO, USA) and Dulbecco's modified Eagle medium (Gibco BRL, Grand island, NY, USA) containing 10% fetal bovine serum (PAN-Biotech, Germany) at 37 ℃ in a humidified atmosphere containing 5% CO2.
-
The sense strand sequences of the short interfering RNAs (siRNAs) used in this study for TRIM21 and the control (NC) were 5′-GCAGGAGUUGGCUGAGAAG-3′ and 5′-UUCUCCGAACGUGUCACGUTT-3′, respectively. At 48 h post-transfection of siRNA using Lipofectamine™ 2000 (Invitrogen, Carlsbad, CA) according to the manufacturer's instructions, the cells were harvested.
-
HA-TRIM21, which has been previously described by us[15], was used as a template to construct the HA-TRIM21-ΔRING mutant plasmid. The primers used were F1: 5′-ATAGAATTCATGGCTTCAGCAGCACGC-3′, R1: 5′-TTGAGCAGAAAGCGCTGGCATGTGACCTCCTCCCACATC-3′, F2: 5′-CAGCGCTTTCTGCTCAAGAAT-3′, and R2: 5′-AGACTCGAGTCAATAGTCAGTGGATCCTTGTGA-3′. The F1 and R1 primers amplify the DNA fragment upstream of the RING domain, whereas the F2 and R2 as primers amplify the DNA fragment downstream of the RING domain. A second round of PCR was then performed using the two fragments at a molar ratio of 1:1 as templates to generate TRIM21-ΔRING. The purified TRIM21-ΔRING-encoding complementary DNA encoding TRIM21-ΔRING was inserted into HA-vector to generate HA-TRIM21-ΔRING mutant. The sequence was confirmed by DNA sequencing.
-
H125 pLenti-TRE-EGFP-EF1-rtTA3-IRES-Puro [(H125-V); Obio Technology (Shanghai) Corp., Ltd., China] was used to generate the recombinant lentivirus-based H125-TRIM21 expression system. Human TRIM21-encoding cDNA was amplified by PCR from HA-TRIM21[15] and inserted into the H125-V vector to generate H125-TRIM21 using EcoRI and BamHI endonucleases. EGFP protein was replaced with TRIM21 protein in the H125-V vector.
-
First, 293T cells were cultured in 10-cm dishes until 80%-90% confluent and then were co-transfected with pLP1, pLP2, pLP-VSVG, and H125-vector or H125-TRIM21 plasmids at a ratio of 1:1:1:2 using lipofectamine 2000. After 72 hours, the culture medium of the transfected cells was harvested and centrifuged in a 50 mL centrifuge tube for 10 min at 3, 500 rpm. To collect the lentivirus, the resulting supernatant was transferred to a new centrifuge tube after filtering through a 0.45 μm filter membrane.
U2-OS cells were cultured in 6-well plates overnight until 70%-80% confluent. Then, 2 mL of lentivirus-containing culture medium and 10 μL polybrene (1 mg/mL) were added to each well, including the control uninfected, H125-V-infected, and H125-TRIM21-infected groups. After 24 hours, 1 mL culture medium was replaced with 1 mL McCoy's 5A culture medium.
After 72 h, 10 μL puromycin (1 mg/mL) were added to each well for screening. Subsequently, McCoy's 5A medium containing 5 μg/mL puromycin was changed once every 2-3 days until the cells in the control uninfected group were all dead. Once the drug screening was complete, the cells were passaged and cryopreserved.
-
MTT, 3-(4, 5-dimethylthiazol-2-yl)-2, 5-diphenyltetrazolium bromide (Beyotime Biotechnology, Shanghai, China), assays were employed to evaluate changes in cell viability. U2-OS or Saos-2 cells were transfected with HA-Vector or HA-TRIM21 alone or a combination of HA-TRIM21 and Flag-YWHAZ plasmids in 6-well plates as indicated and then incubated for 12 hours. The cells were then resuspended in Dulbecco's modified Eagle medium containing fetal bovine serum and seeded into 96-well plates (TCP011096, JET, Guangzhou, China) at a density of 2 × 105 cells/well. Then, cells were treated with CDDP (cisplatin, 30 μg/mL, Sigma-Aldrich, America), TRAIL (tumor necrosis factor related apoptosis-inducing ligand, 50 ng/mL, Sino Biological Inc. Beijing, China), or serum starvation for 24 hours. Next, the cells were incubated with MTT solution (20 μg/mL) in a 37 ℃ incubator for 4 h and the generated formazan crystals were dissolved in 150 μL DMSO/well as described previously[15, 28]. Finally, the absorbance of the formazan crystals at 570 nm was measured using a spectrophotometric microplate reader (ELX800, BioTek Instrument, USA) and cellular viability was evaluated compared to the control samples (set at 100%).
-
U2-OS cells stably infected with H125-V or H125-TRIM21 lentivirus were incubated with tetracycline (TET) for 18 hours. The cells were then lysed in lysis buffer (50 mmol/L Tris at pH 7.5, 150 mmol/L NaCl, 1% NP-40, 0.5% deoxycholate, 0.1% SDS, 5 mmol/L NaF, 2 mmol/L Na3VO4, 1 mmol/L PMSF, and 1% protease inhibitor cocktail from Roche) on ice for 30 min. Co-IPs were performed using previously described protocols[28, 29] with TRIM21 or control IgG primary antibody. Approximately 1 mg of total protein from the lysate was mixed with 2 μg of TRIM21 antibody. The immune complexes were eluted from beads using 500 μL 200 mmol/L NH2CH2COOH (pH 2.5) and promptly neutralized with 160 μL 1 mol/L NH4HCO3 (pH 8.8)[30]. The co-IPs were repeated three times for subsequent in-solution digestion and liquid chromatography-tandem mass spectroscopy (LC-MS/MS).
The standard co-IP assays were performed with HA antibody to validate the U2-OS cells were transfected with HA-vector or HA-TRIM21 plasmid. The immune complexes were separated by western blotting using antibodies against TRIM21 and YWHAZ.
-
Freeze-dried samples were dissolved in 100 μL of 8 mol/L urea. Dithiothreitol was added at a final concentration of 10 mmol/L and the samples were incubated for 1 h at 57 ℃. Then, Iodoacetamide was added to the solution at a final concentration of 55 mmol/L and the samples were incubated at room temperature in the dark for 0.5 h. For in-solution digestion, trypsin was added to the eluate at a trypsin:protein ratio of 1:50 and the samples were incubated at 37 ℃ overnight. The tryptic digest was acidified to pH 3 using trifluoroacetic acid and purified using C18 Microspin columns (Harvard Apparatus). Dried peptides were dissolved in 0.1% formic acid containing 2% acetonitrile before analysis by mass spectrometry.
-
The peptide mixtures were loaded onto a C18 reverse phase column with an auto-sampler and then eluted with a 60-min gradient at a flow rate of 280 nL/min. The effluent was directly electrosprayed into an Orbitrap Fusion Lumos (Thermo Scientific, Waltham, MA, USA) mass spectrometer operated in data-dependent mode. The typical mass spectrometric parameters were a spray voltage of 2 kV, heated capillary temperature of 320 ℃, scan range of 350-1, 500 m/z, resolution of 60, 000 at m/z 400, repeat duration of 30 s, and exclusion duration of 90 s[29].
Raw data were automatically processed by the Sequest HT engine of the Proteome Discoverer v2.1.1.21 (Thermo) against a database of Uniport-Humo (20171002) using the default settings, including a precursor-ion mass tolerance of 10 ppm and fragment-ion mass tolerance of 0.02 Da. The false discovery rate for protein and peptide determination was set to 0.01. At least two unique peptide identifications were required for each protein.
-
IF assays were performed for U2-OS cells using previously described methods[31, 32]. TRIM21 (1:200 dilution) and YWHAZ primary antibodies (1:200 dilution) and Alexa Fluor 488-and Alexa Fluor 594-conjugated secondary antibodies (ZSGB-BIO, China) were used for examination. DAPI staining was used to assess the morphology of the cell nuclei.
Imaging was carried out using a laser scanning confocal microscope (LSM700, Zeiss, Jena, Germany) equipped with a Zeiss Plan-Neofluar 40X/1.3 NA Oil Dic objective as described previously[31, 32].
-
Bimolecular fluorescence complementation (BiFC) is used to observe protein-protein interactions in live cells[33]. The fundamental principles of BiFC are reconstitution of an intact fluorescent protein (such as Venus fluorescent protein) by a pair of interact ing proteins, which are linked to two complementary non-fluorescent fragments[34]. BiFC vector plasmids, including pHA-VC155 and pMyc-VN155, were kindly provided by Prof. Hu of Purdue University. The cDNA of TRIM21 was amplified by PCR from human HA-TRIM21 and inserted into the pHA-VC155 plasmid to generate VC-TRIM21. YWHAZ cDNA was amplified by PCR from human Flag-YWHAZ and inserted into pMyc-VN155 to produce VN-YWHAZ. U2-OS cells were co-transfected with pHA-VC155 (VC-V), pMyc-VN155 (VN-V), VC-TRIM21, or VN-YWHAZ as indicated for 24 hours and then observed using an inverted fluorescence microscope (Olympus, Japan) with a 20× objective.
-
U2-OS cells were transfected with the indicated plasmids or siRNAs and left untreated or were treated with MG132 (100 ng/mL) as indicated. Total cell lysates were harvested and prepared with buffer containing 20 mmol/L pH 7.5 Tris, 150 mmol/L NaCl, 1% Triton X-100, sodium pyrophosphate, β-glycerophosphate, EDTA, Na3VO4, leupeptin, and 1% protease inhibitor cocktail from Roche[15, 28, 32]. Then, 20-50 μg of protein were separated by 10% or 12% SDS-PAGE.
Blots were probed with antibodies against TRIM21 (Santa Cruz, USA), YWHAZ (Proteintech, Wuhan, China), HA (H9658, Sigma, USA), Flag (Sigma, USA), proliferating cell nuclear antigen (PCNA; Santa Cruz, PC10: sc-56), β-Actin (ZSGB-BIO, China), and GAPDH (ZSGB-BIO, China) according to a standard protocol. The images are representative of at least 3 independent experiments.
-
Statistical analysis using Student's t-test was performed and statistical significance was defined as a P < 0.05.
-
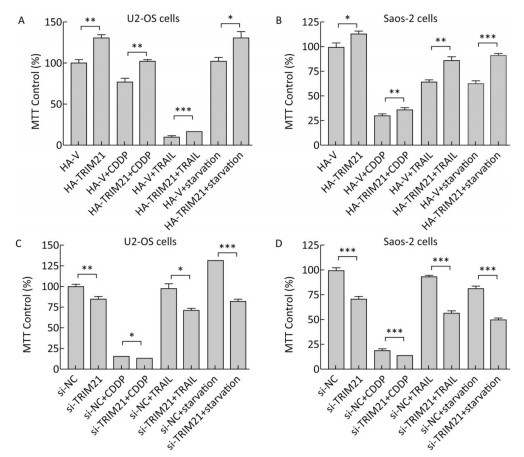
TRIM21 is involved in the tumorigenesis of several types of tumors[12, 13, 19, 20]. In the present study, we found overexpression of HA-TRIM21 significantly increased U2-OS and Saos-2 osteosarcoma cell viability using MTT assays (Figure 1A-B). Meanwhile, overexpression of HA-TRIM21 protected U2-OS and Saos-2 cells from cytotoxicity in response to several stresses, including exposure to CDDP (30 μg/mL), TRAIL (50 ng/mL), and serum starvation for 24 hours (Figure 1A-B). By contrast, TRIM21 knockdown with siRNA significantly decreased U2-OS and Saos-2 cell viability (Figure 1C-D), as well as amplified cytotoxicity from the aforementioned stresses (Figure 1C-D). These results suggest TRIM21 was a positive regulator of U2-OS and Saos-2 cell proliferation and increased U2-OS and Saos-2 cell tolerance to different stresses.
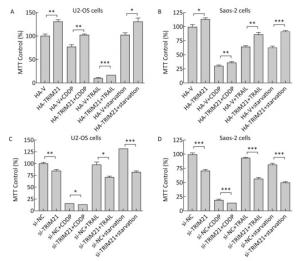
Figure 1. The effects of TRIM21 on osteosarcoma cell proliferation. (A) U2-OS and (B) Saos-2 cells were transfected with HA-TRIM21 or HA-vector plasmid in 6-well plates and incubated overnight. The cells were then resuspended in McCoy's 5A medium containing fetal bovine serum and seeded into 96-well plates. The cells were then treated with CDDP (30 μg/mL), TRAIL (50 ng/mL), or serum starvation for 24 hours as indicated. MTT assays were performed to assess cellular viability through monitoring the absorbance of formazan crystals at 570 nm. (C) U2-OS and (D) Saos-2 cells were transfected with control (si-NC) or TRIM21-targeting siRNA in 6-well plates overnight. The cells were treated as described in A and B. The absorbance of the untreated control group was set to 100%. *, P < 0.05; **, P < 0.01; ***, P < 0.001.
-
Next, we identified TRIM21-interacting partners in U2-OS cells using co-IP coupled with LC-MS/MS analysis. To this end, U2-OS cells stably expressing H125-TRIM21 were employed, where the H125-TRIM21 expression system was recombinant lentivirus-based and regulated by a tetracycline (TET) response element (Figure 2A). After induction with TET (10 mg/mL) for 18 h, control U2-OS cells stably expressing the H125-Vector displayed strong green fluorescence (Figure 2B), whereas U2-OS cells stably expressing H125-TRIM21 produced large amounts of TRIM21 protein (Figure 2C).
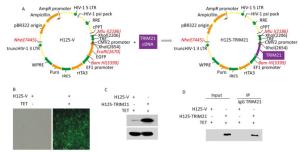
Figure 2. Introduction of recombinant lentivirus vector expressing TRIM21 (H125-TRIM21) and co-IP assay prepared for LC-MS/MS analysis. (A) Schematic of the construction of recombinant lentivirus vector expressing TRIM21 (H125-TRIM21). (B) Expression of GFP by H125-Vector (H125-V)-transfected cells. U2-OS cells stably expressing H125-V were incubated with tetracycline (TET; 10 mg/mL) for 18 hours. The cells were then visualized using an inverted fluorescence microscope (Olympus, Japan) with a 20 × objective. (C) TRIM21 protein levels in U2-OS cells stably expressing H125-TRIM21. U2-OS cells stably expressing H125-TRIM21 were incubated with TET (10 mg/mL) for 18 hours and then analyzed by Western Blotting with TRIM21 and GAPDH antibodies. (D) Co-IP assay was used to identify the interacting partners of TRIM21. U2-OS cells stably expressing H125-V or H125-TRIM21 incubated with TET (10 mg/mL) for 18 hours were used in standard co-IP assay with control IgG or TRIM21 antibody. The resulting immune complexes containing IgG or TRIM21 antibody were subsequently subjected to in-solution digestion and analyzed by LC-MS/MS.
U2-OS cells stably expressing H125-TRIM21 were used in co-IP with TRIM21 antibody (Figure 2D). Proteins that co-precipitated with TRIM21 were identified by LC-MS/MS. The top three identified proteins are listed in Table 1. The top TRIM21-interacting partner was YWHAZ, which is overexpressed and acts as oncogene to promote proliferation in various tumors[21, 22]. However, there have been few reports on the role of YWHAZ in osteosarcoma. In the present study, to investigate the mechanism of TRIM21 as a positive regulator of U2-OS and Saos-2 cell proliferation, YWHAZ protein was selected for subsequent experiments.
Table 1. Top Three TRIM21 Protein-interacting Partners Identified in this Study
Accession Gene Names Protein Names Molecular Weight (kD) Total Number of Peptides Assigned Sequence Coverage (%) Protein False Discovery Rate Confidence P63104 YWHAZ 14-3-3 protein zeta/delta 27.7 3 12.6531 High P26641 EEF1G Elongation factor 1-gamma 50.1 2 6.4073 High P17066 HSPA6 Heat-shock 70 kD protein 6 71.0 2 6.0653 High -
To confirm the interaction between TRIM21 and YWHAZ, co-IP of U2-OS cells transfected with HA-vector or HA-TRIM21 plasmid were performed. The interacting proteins bound by HA antibody were subsequently analyzed by immunoblot (IB) assay. As shown in Figure 3A, YWHAZ bound to TRIM21.
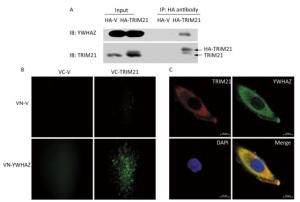
Figure 3. Interaction between TRIM21 and YWHAZ. (A) Co-IP assay was used to validate the interaction between TRIM21 and YWHAZ. U2-OS cells transfected with HA-vector or HA-TRIM21 plasmid were employed to carry out Co-IP assay with HA antibody. The immune complexes were analyzed by immunoblotting with TRIM21 and YWHAZ antibodies. (B) BIFC assay. U2-OS cells were co-transfected with pHA-VC155 (VC-V), pMyc-VN155 (VN-V), VC-TRIM21, or VN-YWHAZ as indicated in B. After 24 hours, the fluorescence was visualized using an inverted fluorescence microscope with a 20 × objective. (C) Confocal microscopy assay. TRIM21 and YWHAZ were immunostained and then visualized using a laser scanning confocal microscope. DAPI staining was used to assess the morphology of cell nuclei. In the merged panel, yellow represents co-localization of two proteins. Scale bar: 10 μm.
BiFC was then employed to further confirm the interaction between TRIM21 and YWHAZ. BiFC is a method used to observe protein-protein interactions in live cells[33]. BiFC vector plasmids, including pHA-VC155 and pMyc-VN155, were used to construct VC-TRIM21 and VN-YWHAZ, respectively. Interactions between TRIM21 and YWHAZ would promote linking of the complementary non-fluorescent fragments VC and VN, which produces Venus fluorescence. As shown in Figure 3B, 24 hours after transfection of U2-OS cells with VC-TRIM21 and VN-YWHAZ, fluorescence was only detected in cells co-transfected with VC-TRIM21 and VN-YWHAZ plasmids, suggesting binding of YWHAZ to TRIM21. Co-localization of TRIM21 with YWHAZ was then examined using confocal microscopy. As shown in Figure 3C, both TRIM21 and YWHAZ were mainly located in the cytoplasm within scattered granules, where co-localization of TRIM21 with YWHAZ is indicated in yellow in the merged panels. Taken together, these results suggest TRIM21 interacted with YWHAZ and these proteins co-localized in the cytoplasm.
-
TRIM21 is an E3 ligase; therefore, we next investigated whether YWHAZ expression is regulated by TRIM21. Knockdown of TRIM21 with its siRNA resulted in a marked increase in YWHAZ levels (Figure 4A), whereas overexpression of TRIM21 after tetracycline induction resulted in a large decrease in YWHAZ levels (Figure 4B). Accordingly, the addition of proteasome inhibitor MG132 abrogated this decrease (Figure 4B). These results hint at the possibility TRIM21 regulated YWHAZ levels through a proteasome pathway.
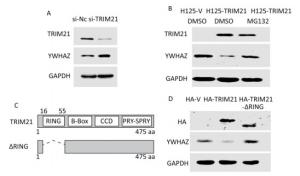
Figure 4. RING domain of TRIM21 required for negative regulation of YWHAZ expression by TRIM21. (A) Knockdown of TRIM21 led to upregulation of YWHAZ expression. U2-OS cells were transfected with control (si-NC) or TRIM21-targeting short interfering RNA and then immunoblotted with the indicated antibodies. (B) Downregulation of YWHAZ induced by TRIM21 overexpression was inhibited by the proteasome inhibitor MG132. U2-OS cells stably expressing H125-V or H125-TRIM21 after induction with TET (10 mg/mL) for 18 hours were left untreated or treated with MG132 and then immunoblotted with the indicated antibodies. (C) Diagram displaying full-length TRIM21 and the TRIM21-ΔRING mutant. (D) Effect of the TRIM21 RING domain on YWHAZ expression. U2-OS cells were transfected with control HA-V, HA-TRIM21, or HA-TRIM21-ΔRING and then immunoblotted with the indicated antibodies.
The RING-finger domain of the TRIM protein contains conserved cysteine and histidine residues in a 'cross-brace' arrangement. The RING domain is necessary for recruiting ubiquitin-conjugating enzymes (E2 Ub) and possesses ubiquitin E3 ligase activity[35]. We then determined whether this RING domain takes part in TRIM21 regulation of YWHAZ. To this end, we constructed the HA-TRIM21-ΔRING mutant as shown in Figure 4C. Unlike the plasmid expressing full-length HA-TRIM21, the HA-TRIM21-ΔRING mutant failed to reduce YWHAZ expression (Figure 4D), indicating the RING domain of TRIM21 was required for TRIM21-mediated negative regulation of YWHAZ.
-
Subsequently, we tested whether YWHAZ is involved in TRIM21 regulation of osteosarcoma cell proliferation. Based on our results showing that TRIM21 negatively regulated the expression level of YWHAZ, we investigated whether YWHAZ overexpression could antagonize the effects of TRIM21 on osteosarcoma cell proliferation. As shown in Figure 5A, overexpression of either HA-TRIM21 or Flag-YWHAZ alone, but not the HA-TRIM21-ΔRING mutant, significantly increased U2-OS cell viability. However, overexpression of Flag-YWHAZ failed to antagonize the effects of TRIM21. Moreover, co-expression of HA-TRIM21 and Flag-YWHAZ notably increased cell viability. Intriguingly, similar results were also obtained for cells co-expressing HA-TRIM21-ΔRING and Flag-YWHAZ (Figure 5A, upper panel). The protein levels in these cohorts are shown in the lower panel of Figure 5A. In agreement with the results in Figure 4, overexpression of full-length HA-TRIM21, but not the HA-TRIM21-ΔRING mutant, reduced Flag-YWHAZ levels. However, overexpression of Flag-YWHAZ had no effect on TRIM21 expression (Figure 5B).
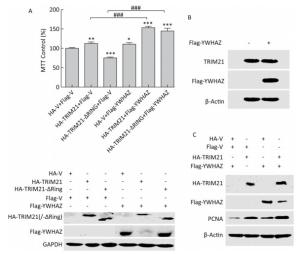
Figure 5. Effects of YWHAZ overexpression on TRIM21 overexpression-induced osteosarcoma cell proliferation. (A) U2-OS cells were co-transfected with HA-TRIM21 or HA-TRIM21-ΔRING and Flag-YWHAZ as indicated and evaluated by MTT assays (upper panel) and Western Blotting (lower panel). Results for control cells co-transfected with HA-vector and Flag-vector were set at 100%. The asterisk indicates a statistically significant difference between the indicated cells and control cells. *: P < 0.05; **: P < 0.01; ***: P < 0.001. ###: P < 0.001. (B) U2-OS cells were transfected with Flag-vector or Flag-YWHAZ as indicated and performed standard IB assay with the TRIM21 and Flag antibodies. (C) U2-OS cells were co-transfected with HA-TRIM21 combined with Flag-YWHAZ as indicated and performed standard IB assay with antibodies against HA, Flag, or PCNA.
In addition, PCNA (proliferating cell nuclear antigen), a marker of cell proliferation[36], was used to evaluated the proliferation of U2-OS cells. As shown in Figure 5C, PCNA expression was upregulated in U2-OS cells overexpressing either HA-TRIM21 or Flag-YWHAZ alone compared to the control group. Furthermore, co-expression of both HA-TRIM21 and Flag-YWHAZ caused an obvious re-regulation of PCNA (Figure 5B), which is similar to the trend in cell proliferation observed in Figure 5A. Taken together, our results suggest that YWHAZ was not involved in TRIM21 regulation of osteosarcoma cell proliferation.
-
Osteosarcoma is the most commonly occurring malignant bone tumor in children and adolescents[1, 2]. Despite major advances in surgical techniques and neoadjuvant chemotherapy, the 5-year survival rate of 65%-70% of patients with localized osteosarcoma has not increased in three decades[3, 4].
In patients with metastatic or relapsed osteosarcoma, the overall 5-year survival rate is typically < 25%. Investigations into the mechanisms underlying osteosarcoma development, including cell proliferation, are necessary for understanding the processes of tumorigenesis and developing novel therapeutic approaches. In the present study, we found TRIM21 protein was a positive regulator of osteosarcoma cell proliferation. Overexpression of TRIM21 promoted U2-OS and Saos-2 osteosarcoma cell proliferation and protected these cells from different stresses. Conversely, knockdown of TRIM21 blocked U2-OS and Saos-2 osteosarcoma cell proliferation and enhanced the toxicity of different stresses. Therefore, decreasing the expressional level or activity of TRIM21 protein might be a novel potential strategy for osteosarcoma therapy.
TRIM family proteins are involved in a wide range of biological processes and diverse pathological conditions, including developmental disorders, neurodegenerative diseases, viral infections, and cancers[7]. Due to having a RING domain, most TRIM members function as E3 ubiquitin ligases[6]. The roles of E3 ligases in the regulation of oncogenes and tumor suppressors have considerably aroused the interest of tumor researchers[6]. Through preventing E3 ligases from binding to their specific substrates, drugs or methods targeting individual E3 ligases could act as specific tumor therapeutics. TRIM21 has E3 ligase activity and functions by interacting with a variety of partners[9]. Previous studies on TRIM21 have been predominantly related to the innate immune system[10]. However, the functions of this factor in tumor biology have recently gained more attention. For example, TRIM21 was identified as a novel interacting partner of the tumor-suppressor Par-4 in colon and pancreatic cancer cells and was found to protect colon cancer cells from cisplatin-induced apoptosis by downregulating Par-4 levels[12]. There has been few reports on the role of TRIM21 in osteosarcoma; therefore, in this study, we identified the interacting partners of TRIM21 in osteosarcoma cells, such as the novel partner YWHAZ, using co-IP combined with mass-spectrometry. The biological processes related to interactions between TRIM21 and YWHAZ deserve intensive study.
The 14-3-3 protein family members are ubiquitously expressed in eukaryotes, including the seven isoforms β, τ, ε, ζ, η, γ, and σ in mammals. These 14-3-3 proteins participate in diverse biological processes, including regulation of cell proliferation, apoptosis, the cell cycle, and DNA damage[21, 22]. Despite high homology in the protein sequences of 14-3-3 family protein members, mouse models with deletions in different members have revealed different biological functions for distinct isoforms[37, 38]. YWHAZ, also called 14-3-3ζ, is overexpressed in different types of cancer and functions as an oncogene in tumors through interacting with multiple proteins[21, 24]. Recent reports have shown YWHAZ promotes proliferation, migration, and invasion of lung adenocarcinoma cells by interacting with and upregulating Mucin 1 protein[21]. The loss of YWHAZ inhibits tumor proliferation and angiogenesis and promotes apoptosis of mammary tumor cells by attenuating oncogenic signaling through Akt and ERK[39]. YWHAZ regulates hepatocellular carcinoma (HCC) cell proliferation, tumor growth, and chemosensitivity by influencing the mitogen-activated protein kinase (MAPK), c-Jun N-terminal kinase (JNK), and p38 signaling pathways[23]. However, the role of YWHAZ in osteosarcoma has been little studied. In the present study, YWHAZ protein overexpression was found to positively regulate osteosarcoma cell proliferation (Figure 5). In addition, YWHAZ was identified as a novel interacting partner of TRIM21 protein (Figure 3). TRIM21 negatively regulated YWHAZ expression and the RING domain of TRIM21, which is associated with the E3 ligase activity of TRIM21, was necessary for this negative regulation (Figure 4), suggesting YWHAZ might be a substrate of TRIM21 E3 ligase. Interestingly, YWHAZ protein overexpression did not prevent TRIM21 from promoting osteosarcoma cell proliferation. Therefore, it is possible TRIM21 and YWHAZ interactions are not involved in TRIM21-regulated osteosarcoma cell proliferation. Both TRIM21 and YWHAZ are involved in other malignant behaviors, such as migration[19, 21] and apoptosis antagonism in cancer cells[12, 23]. Therefore, it is possible TRIM21 and YWHAZ interactions mediate these processes, which should be further investigated.
In conclusion, we demonstrated TRIM21 protein positively regulated osteosarcoma cell proliferation and the expressional level of TRIM21 affected the tolerance of osteosarcoma cells to various stresses. YWHAZ protein was identified as a novel interacting partner of TRIM21 and its expression levels were found to be negatively regulated by TRIM21 E3 ligase. The functions of TRIM21 and YWHAZ interactions require further investigation as they did not have a role in TRIM21-regulated osteosarcoma cell proliferation.
-
ZENG Qing Zhong performed the majority of the experiments. LIU Wan Ting participated in LC-MS/MS analysis. LU Jun Lei and LIU Xiao Hui participated in constructing plasmids. ZHANG Yun Fang participated in discussions on osteosarcoma research progress. LIU Lang Xia and GAO Xue Juan designed the study and drafted the manuscript.
doi: 10.3967/bes2018.024
YWHAZ Binds to TRIM21 but Is Not Involved in TRIM21-stimulated Osteosarcoma Cell Proliferation
-
Abstract:
Objective Osteosarcoma is the most common type of malignant bone tumor in children and adolescents. The role of E3 ligases in tumorigenesis is currently a focus in tumor research. In the present study, we investigated the role of the E3 ligase tripartite motif 21 (TRIM21) in osteosarcoma cell proliferation. Methods 3-(4, 5-dimethylthiazol-2-yl)-2, 5-diphenyltetrazolium bromide (MTT) assays were used to assess osteosarcoma cell viability. U2-OS cells stably carrying a recombinant lentivirus expressing tetracycline-regulated TRIM21 were screened. Co-immunoprecipitation was coupled with LCMS/MS analysis to identify novel interacting partners of TRIM21. Co-immunoprecipitation and bimolecular fluorescence complementation (BIFC) were performed to validate the interactions between TRIM21 and its novel partner YWHAZ. A TRIM21-ΔRING construct was generated to test the effects of TRIM21 ligase activity on YWHAZ. Results TRIM21 positively regulated osteosarcoma cell proliferation. Overexpression of TRIM21 enhanced osteosarcoma cell tolerance toward various stresses. YWHAZ protein was identified as a novel interacting partner of TRIM21 and its expression levels were negatively regulated by TRIM21. The RING domain of TRIM21 was required for TRIM21 negative regulation of YWHAZ expression. However, overexpression of YWHAZ did not affect positive regulation of osteosarcoma cell proliferation by TRIM21. Conclusion Our results further clarify the molecular mechanisms underlying the pathogenesis of osteosarcoma. -
Key words:
- TRIM21 /
- YWHAZ /
- Osteosarcoma /
- Proliferation
-
Figure 1. The effects of TRIM21 on osteosarcoma cell proliferation. (A) U2-OS and (B) Saos-2 cells were transfected with HA-TRIM21 or HA-vector plasmid in 6-well plates and incubated overnight. The cells were then resuspended in McCoy's 5A medium containing fetal bovine serum and seeded into 96-well plates. The cells were then treated with CDDP (30 μg/mL), TRAIL (50 ng/mL), or serum starvation for 24 hours as indicated. MTT assays were performed to assess cellular viability through monitoring the absorbance of formazan crystals at 570 nm. (C) U2-OS and (D) Saos-2 cells were transfected with control (si-NC) or TRIM21-targeting siRNA in 6-well plates overnight. The cells were treated as described in A and B. The absorbance of the untreated control group was set to 100%. *, P < 0.05; **, P < 0.01; ***, P < 0.001.
Figure 2. Introduction of recombinant lentivirus vector expressing TRIM21 (H125-TRIM21) and co-IP assay prepared for LC-MS/MS analysis. (A) Schematic of the construction of recombinant lentivirus vector expressing TRIM21 (H125-TRIM21). (B) Expression of GFP by H125-Vector (H125-V)-transfected cells. U2-OS cells stably expressing H125-V were incubated with tetracycline (TET; 10 mg/mL) for 18 hours. The cells were then visualized using an inverted fluorescence microscope (Olympus, Japan) with a 20 × objective. (C) TRIM21 protein levels in U2-OS cells stably expressing H125-TRIM21. U2-OS cells stably expressing H125-TRIM21 were incubated with TET (10 mg/mL) for 18 hours and then analyzed by Western Blotting with TRIM21 and GAPDH antibodies. (D) Co-IP assay was used to identify the interacting partners of TRIM21. U2-OS cells stably expressing H125-V or H125-TRIM21 incubated with TET (10 mg/mL) for 18 hours were used in standard co-IP assay with control IgG or TRIM21 antibody. The resulting immune complexes containing IgG or TRIM21 antibody were subsequently subjected to in-solution digestion and analyzed by LC-MS/MS.
Figure 3. Interaction between TRIM21 and YWHAZ. (A) Co-IP assay was used to validate the interaction between TRIM21 and YWHAZ. U2-OS cells transfected with HA-vector or HA-TRIM21 plasmid were employed to carry out Co-IP assay with HA antibody. The immune complexes were analyzed by immunoblotting with TRIM21 and YWHAZ antibodies. (B) BIFC assay. U2-OS cells were co-transfected with pHA-VC155 (VC-V), pMyc-VN155 (VN-V), VC-TRIM21, or VN-YWHAZ as indicated in B. After 24 hours, the fluorescence was visualized using an inverted fluorescence microscope with a 20 × objective. (C) Confocal microscopy assay. TRIM21 and YWHAZ were immunostained and then visualized using a laser scanning confocal microscope. DAPI staining was used to assess the morphology of cell nuclei. In the merged panel, yellow represents co-localization of two proteins. Scale bar: 10 μm.
Figure 4. RING domain of TRIM21 required for negative regulation of YWHAZ expression by TRIM21. (A) Knockdown of TRIM21 led to upregulation of YWHAZ expression. U2-OS cells were transfected with control (si-NC) or TRIM21-targeting short interfering RNA and then immunoblotted with the indicated antibodies. (B) Downregulation of YWHAZ induced by TRIM21 overexpression was inhibited by the proteasome inhibitor MG132. U2-OS cells stably expressing H125-V or H125-TRIM21 after induction with TET (10 mg/mL) for 18 hours were left untreated or treated with MG132 and then immunoblotted with the indicated antibodies. (C) Diagram displaying full-length TRIM21 and the TRIM21-ΔRING mutant. (D) Effect of the TRIM21 RING domain on YWHAZ expression. U2-OS cells were transfected with control HA-V, HA-TRIM21, or HA-TRIM21-ΔRING and then immunoblotted with the indicated antibodies.
Figure 5. Effects of YWHAZ overexpression on TRIM21 overexpression-induced osteosarcoma cell proliferation. (A) U2-OS cells were co-transfected with HA-TRIM21 or HA-TRIM21-ΔRING and Flag-YWHAZ as indicated and evaluated by MTT assays (upper panel) and Western Blotting (lower panel). Results for control cells co-transfected with HA-vector and Flag-vector were set at 100%. The asterisk indicates a statistically significant difference between the indicated cells and control cells. *: P < 0.05; **: P < 0.01; ***: P < 0.001. ###: P < 0.001. (B) U2-OS cells were transfected with Flag-vector or Flag-YWHAZ as indicated and performed standard IB assay with the TRIM21 and Flag antibodies. (C) U2-OS cells were co-transfected with HA-TRIM21 combined with Flag-YWHAZ as indicated and performed standard IB assay with antibodies against HA, Flag, or PCNA.
Table 1. Top Three TRIM21 Protein-interacting Partners Identified in this Study
Accession Gene Names Protein Names Molecular Weight (kD) Total Number of Peptides Assigned Sequence Coverage (%) Protein False Discovery Rate Confidence P63104 YWHAZ 14-3-3 protein zeta/delta 27.7 3 12.6531 High P26641 EEF1G Elongation factor 1-gamma 50.1 2 6.4073 High P17066 HSPA6 Heat-shock 70 kD protein 6 71.0 2 6.0653 High -
[1] Anderson ME. Update on Survival in Osteosarcoma. Orthop Clin North Am, 2016; 47, 283-92. doi: 10.1016/j.ocl.2015.08.022 [2] Berner K, Hall KS, Monge OR, et al. Prognostic factors and treatment results of high-grade osteosarcoma in norway:a scope beyond the 'classical' patient. Sarcoma, 2015; 2015, 516843. https://www.hindawi.com/journals/sarcoma/2015/516843/fig3/ [3] Ferrari S, Serra M. An update on chemotherapy for osteosarcoma. Expert Opin Pharmacother, 2015; 16, 2727-36. doi: 10.1517/14656566.2015.1102226 [4] Broadhead ML, Clark JC, Myers DE, et al. The molecular pathogenesis of osteosarcoma:a review. Sarcoma, 2011; 2011, 959248. http://www.oalib.com/paper/3080985 [5] Weissman AM. Regulating protein degradation by ubiquitination. Immunol Today, 1997; 18, 189-98. doi: 10.1016/S0167-5699(97)84666-X [6] Hatakeyama S. TRIM proteins and cancer. Nat Rev Cancer, 2011; 11, 792-804. doi: 10.1038/nrc3139 [7] Ozato K, Shin DM, Chang TH, et al. TRIM family proteins and their emerging roles in innate immunity. Nat Rev Immunol, 2008; 8, 849-60. doi: 10.1038/nri2413 [8] Zhang Z, Bao M, Lu N, et al. The E3 ubiquitin ligase TRIM21 negatively regulates the innate immune response to intracellular double-stranded DNA. Nat Immunol, 2013; 14, 172-8. https://www.sigmaaldrich.com/catalog/papers/23222971 [9] Reddy BA, van der Knaap JA, Bot AG, et al. Nucleotide biosynthetic enzyme GMP synthase is a TRIM21-controlled relay of p53 stabilization. Mol Cell, 2014; 53, 458-70. doi: 10.1016/j.molcel.2013.12.017 [10] McEwan WA, James LC. TRIM21-dependent intracellular antibody neutralization of virus infection. Prog Mol Biol Transl Sci, 2015; 129, 167-87. doi: 10.1016/bs.pmbts.2014.10.006 [11] Kyriakidis NC, Kapsogeorgou EK, Gourzi VC, et al. Toll-like receptor 3 stimulation promotes Ro52/TRIM21 synthesis and nuclear redistribution in salivary gland epithelial cells, partially via type Ⅰ interferon pathway. Clin Exp Immunol, 2014; 178, 548-60. doi: 10.1111/cei.2014.178.issue-3 [12] Nguyen JQ, Irby RB. TRIM21 is a novel regulator of Par-4 in colon and pancreatic cancer cells. Cancer Biol Ther, 2017; 18, 16-25. doi: 10.1080/15384047.2016.1252880 [13] Muller J, Maurer V, Reimers K, et al. TRIM21, a negative modulator of LFG in breast carcinoma MDA-MB-231 cells in vitro. Int J Oncol, 2015; 47, 1634-46. doi: 10.3892/ijo.2015.3169 [14] Wada K, Niida M, Tanaka M, et al. Ro52-mediated monoubiquitination of IKK{beta} down-regulates NF-{kappa}B signalling. J Biochem, 2009; 146, 821-32. doi: 10.1093/jb/mvp127 [15] Gao X, Xu F, Zhang HT, et al. PKCalpha-GSK3beta-NF-kappaB signaling pathway and the possible involvement of TRIM21 in TRAIL-induced apoptosis. Biochem Cell Biol, 2016; 94, 256-64. doi: 10.1139/bcb-2016-0009 [16] Jauharoh SN, Saegusa J, Sugimoto T, et al. SS-A/Ro52 promotes apoptosis by regulating Bcl-2 production. Biochem Biophys Res Commun, 2012; 417, 582-7. doi: 10.1016/j.bbrc.2011.12.010 [17] Shibata N, Ohoka N, Sugaki Y, et al. Degradation of Stop Codon Read-through Mutant Proteins via the Ubiquitin-Proteasome System Causes Hereditary Disorders. J Biol Chem, 2015; 290, 28428-37. doi: 10.1074/jbc.M115.670901 [18] Espinosa A, Zhou W, Ek M, et al. The Sjogren's syndrome-associated autoantigen Ro52 is an E3 ligase that regulates proliferation and cell death. J Immunol, 2006; 176, 6277-85. doi: 10.4049/jimmunol.176.10.6277 [19] Ding Q, He D, He K, et al. Downregulation of TRIM21 contributes to hepatocellular carcinoma carcinogenesis and indicates poor prognosis of cancers. Tumour Biol, 2015; 36, 8761-72. doi: 10.1007/s13277-015-3572-2 [20] Brauner S, Zhou W, Backlin C, et al. Reduced expression of TRIM21/Ro52 predicts poor prognosis in diffuse large B-cell lymphoma patients with and without rheumatic disease. J Intern Med, 2015; 278, 323-32. doi: 10.1111/joim.2015.278.issue-3 [21] Xue M, Tao W. Upregulation of MUC1 by its novel activator 14-3-3zeta promotes tumor invasion and indicates poor prognosis in lung adenocarcinoma. Oncol Rep, 2017; 38, 2637-46. doi: 10.3892/or.2017.5948 [22] Obsilova V, Kopecka M, Kosek D, et al. Mechanisms of the 14-3-3 protein function:regulation of protein function through conformational modulation. Physiol Res, 2014; 63, S155-64. http://www.academia.edu/12485797/Mechanisms_of_the_14-3-3_protein_function_Regulation_of_protein_function_through_conformational_modulation [23] Wu YJ, Jan YJ, Ko BS, et al. Involvement of 14-3-3 Proteins in Regulating Tumor Progression of Hepatocellular Carcinoma. Cancers (Basel), 2015; 7, 1022-36. doi: 10.3390/cancers7020822 [24] Watanabe N, Komatsu S, Ichikawa D, et al. Overexpression of YWHAZ as an independent prognostic factor in adenocarcinoma of the esophago-gastric junction. Am J Cancer Res, 2016; 6, 2729-36. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5126287 [25] Ruenauver K, Menon R, Svensson MA, et al. Prognostic significance of YWHAZ expression in localized prostate cancer. Prostate Cancer Prostatic Dis, 2014; 17, 310-4. doi: 10.1038/pcan.2014.32 [26] Nishimura Y, Komatsu S, Ichikawa D, et al. Overexpression of YWHAZ relates to tumor cell proliferation and malignant outcome of gastric carcinoma. Br J Cancer, 2013; 108, 1324-31. doi: 10.1038/bjc.2013.65 [27] Neal CL, Yao J, Yang W, et al. 14-3-3zeta overexpression defines high risk for breast cancer recurrence and promotes cancer cell survival. Cancer Res, 2009; 69, 3425-32. doi: 10.1158/0008-5472.CAN-08-2765 [28] Gao X, Feng J, He Y, et al. hnRNPK inhibits GSK3beta Ser9 phosphorylation, thereby stabilizing c-FLIP and contributes to TRAIL resistance in H1299 lung adenocarcinoma cells. Sci Rep, 2016; 6, 22999. doi: 10.1038/srep22999 [29] Gao X, Wang JY, Gao LM, et al. Identification and analysis of glycogen synthase kinase 3 beta1 interactome. Cell Biol Int, 2013; 37, 768-79. doi: 10.1002/cbin.v37.8 [30] Varjosalo M, Sacco R, Stukalov A, et al. Interlaboratory reproducibility of large-scale human protein-complex analysis by standardized AP-MS. Nat Methods, 2013; 10, 307-14. doi: 10.1038/nmeth.2400 [31] Gao X, Dan S, Xie Y, et al. 14-3-3ζ reduces DNA damage by interacting with and stabilizing proliferating cell nuclear antigen. J Cell Biochem, 2015; 116, 158-69. doi: 10.1002/jcb.v116.1 [32] Huang WS, Xu FM, Zeng QZ, et al. ERK1/2-mediated Cytoplasmic Accumulation of hnRNPK Antagonizes TRAIL-induced Apoptosis through Upregulation of XIAP in H1299 Cells. Biomed Environ Sci, 2017; 30, 473-81. http://www.sciencedirect.com/science/article/pii/S0895398817300818 [33] Pratt EP, Owens JL, Hockerman GH, et al. Bimolecular Fluorescence Complementation (BiFC) Analysis of Protein-Protein Interactions and Assessment of Subcellular Localization in Live Cells. Methods Mol Biol, 2016; 1474, 153-70. doi: 10.1007/978-1-4939-6352-2 [34] Kodama Y, Hu CD. Bimolecular fluorescence complementation (BiFC):a 5-year update and future perspectives. Biotechniques, 2012; 53, 285-98. doi: 10.2144/000113943 [35] Joazeiro CA, Weissman AM. RING finger proteins:mediators of ubiquitin ligase activity. Cell, 2000; 102, 549-52. doi: 10.1016/S0092-8674(00)00077-5 [36] Kubben FJ, Peeters-Haesevoets A, Engels LG, et al. Proliferating cell nuclear antigen (PCNA):a new marker to study human colonic cell proliferation. Gut, 1994; 35, 530-5. doi: 10.1136/gut.35.4.530 [37] Lau JM, Wu C, Muslin AJ. Differential role of 14-3-3 family members in Xenopus development. Dev Dyn, 2006; 235, 1761-76. doi: 10.1002/dvdy.v235:7 [38] Muslin AJ, Lau JM. Differential functions of 14-3-3 isoforms in vertebrate development. Curr Top Dev Biol, 2005; 65, 211-28. [39] Joshi S, Yang J, Wang Q, et al. 14-3-3zeta loss impedes oncogene-induced mammary tumorigenesis and metastasis by attenuating oncogenic signaling. Am J Cancer Res, 2017; 7, 1654-64. http://www.ajcr.us/files/ajcr0026751.pdf -




 下载:
下载:



 Quick Links
Quick Links