-
Cervical cancer (CC) is one of the most common malignant tumors in gynecology, with a peak incidence between the ages of 50 and 55 years. The incidence of CC has slightly declined in recent years while the mortality rate has remained stable[1]. Since its pathogenesis is unclear, the current treatment and prognosis of patients with CC remain ineffective and poor, respectively[2]. The etiology of CC involves complex molecular processes, including activation of oncogenes and inhibition of tumor suppressor genes[3]. As researchers continue to study the molecular mechanisms of CC, an increasing number of genes/proteins and signaling pathways involved in the development and progression of CC are continuously being reported, thereby advancing the development of various treatment methods for CC, including gene-targeted therapy and hormone therapy[4]. Ghasemi et al. found that curcumin inhibits the invasion and proliferation ability of CC cells through suppression of the nuclear factor-kB and Wnt/β-catenin pathways and suggested further studies on the potential therapeutic effects of this compound in C[5]. Shafabakhsh et al. reviewed the anti-CC effects of melatonin and its underlying molecular mechanisms[6].
Accordingly, investigating the molecular mechanisms of CC development, progression, metastasis, and recurrence; discovering biomarkers for the clinical diagnosis and prognosis of CC; and identifying effective molecular targets for the treatment of CC have become research hotspots for medical researchers worldwide[7]. Recent advances in biological research on CC indicate that epigenetic changes are common in CC development and metastasis, especially those involving microRNA (miRNA/miR) regulation[8]. In particular, it has been found that an increasing number of transcription factors and miRNAs form a feedback loop through various mechanisms, thereby promoting or inhibiting tumor cell growth. In addition, in recent years, a large number of studies have shown that miRNAs are involved in the proliferation and invasion of CC cells, which are processes closely related to the development and progression of CC[9].
MiRNAs are a class of endogenous non-coding RNAs of approximately 22 nucleotides in length that are ubiquitously found in eukaryotes in various forms. To date, thousands of miRNAs have been predicted in the human genome, accounting for 3% of the total number of genes[10]. miRNAs induce the degradation or translation inhibition of target gene mRNAs by specifically pairing with the base site in the 3' untranslated region of each target mRNA, thereby regulating the expression of the target gene at the post-transcriptional level[11, 12]. According to statistics, miRNAs regulate at least 30% of human genes and play a key role in the normal development of the body. However, dysregulation of miRNA expression, which affects the proliferation, migration, and invasion of tumor cells, is also common in a variety of malignant tumors[13, 14]. Therefore, miRNAs have been considered as promising targets for the diagnosis, prognosis, and treatment of tumor progression and metastasis[15–18]. The expression of miR-378a-3p has been reported to be downregulated in clinical breast cancer tissues and cells. In addition, lower expression levels of miR-378a-3p have been found to be related to poor prognosis in patients with breast cancer, and its increased expression strengthens the tamoxifen sensitivity of tumor cells by targeting Golgi transport 1A[19]. MiR-378a-3p can also promote muscle cell differentiation by downregulating histone deacetylase 4[20]. In ovarian cancer, it can sensitize cells to cisplatin by targeting mitogen-activated protein kinase 1/growth factor receptor binding protein 2[21]. It has also been reported as a prognostic factor in colorectal cancer (CRC)[22]. However, the role and underlying molecular mechanism of action in CC remain unclear. Therefore, the aim of this study was to investigate the biological effects of miR-378a-3p on CC cell proliferation. To this end, in this study, we first determined the expression levels of miR-378a-3p in CC tissues and cells and then studied the relationship among its expression and clinicopathological features with the prognosis of CC patients. The effects of miR-378a-3p expression on CC cell proliferation, migration, and apoptosis were also examined in vitro.
-
Twelve CC tissue samples and matched non-tumor adjacent tissue samples were obtained from patients who underwent surgical resection at the Hospital of Integrated Chinese and Western Medicine between May 2017 and January 2019. Non-tumor adjacent tissue samples were collected for use as normal controls. All samples included in this study were from first-time cancer patients with no history of hormone therapy, radiation therapy, or chemotherapy before undergoing surgical treatment. All diagnoses of CC were confirmed by the Department of Pathology of the Beijing Hospital of Integrated Chinese and Western Medicine. The exclusion criteria were as follows: 1) patients who received cervical treatment due to inflammation or infection within 3 months before surgery; 2) pregnancy complications; (3) other malignant tumors of the female reproductive system, such as endometrial cancer, uterine leiomyosarcoma, or ovarian cancer; and 4) patients with other organ tumors, such as intestinal and breast cancer. During the surgery, tissue samples were collected and immediately stored in liquid nitrogen. The use of human tissues was approved by the Ethics Committee of the Hospital of Integrated Chinese and Western Medicine, and the study was conducted in accordance with the Declaration of Helsinki. Written informed consent was obtained from each patient.
-
From January 2015 to February 2018, 10 mL of whole blood was collected from 48 newly diagnosed fasting CC patients in the Hospital of Integrated Chinese and Western Medicine and used to prepare approximately 5 mL of serum. Serum was also prepared from blood collected from 48 healthy subjects admitted for check-ups during the same period and used as the control group. The study was approved by the Ethics Committee of the Hospital of Integrated Chinese and Western Medicine, and all participants were informed and volunteered to participate. Blood samples were centrifuged at 2,500 rpm for 10 min to prepare the serum. The serum was then placed in a cryogenic high-speed centrifuge at 4 °C and centrifuged at 12,000 rpm for 10 min. The serum supernatant was collected and stored separately in a freezer at −80 °C for later use.
-
The CC cell lines SiHa and Hela were obtained from the Chinese Academy of Sciences Cell Bank (Beijing, China). They were cultured in Dulbecco’s Modified Eagle’s Medium containing 10% fetal bovine serum at 37 °C in 5% CO2. The miR-378a-3p mimic and scramble negative control (NC) sequence were synthesized by Shanghai Jima Biotechnology Co. Ltd. (Shanghai, China). They were transfected into SiHa and HeLa cells using Lipofectamine 2000 according to the manufacturer’s instructions. After transfection, the cells were further cultured for 48 h.
-
TRIzol reagent (Solarbio, Beijing, China) was used to extract total RNA from cells in the logarithmic growth stage according to the manufacturer’s instructions, and RNA integrity was confirmed by agarose gel electrophoresis. Total RNA was used to synthesize complementary DNA by reverse transcription according to the instructions of the reverse transcription kit. PCR conditions were as follows: initial denaturation at 95 °C for 5 min, followed by 40 cycles of denaturation at 95 °C for 30 s and annealing at 60 °C for 30 s. U6 non-coding small nuclear RNA (snRNA) was used as an internal reference. The relative expression of miR-378a-3p was calculated by the 2−ΔΔCt method. The miR-378a-3p forward primer was ACACTCCAGCTGGGACUGGACUUGGAGUCA, and the reverse primer was CTCAACTGGTGTCGTGGAGTCGGCAATTCAGTTGAGGCCTTCTG. The U6 snRNA forward primer was CTCGCTTCGGCAGCACA, and the reverse primer was AACGCTTCACGAATTTGCGT.
-
The Cell Counting Kit-8 (CCK-8) assay (Beyotime Biotechnology, Shanghai, China) was used to determine the proliferation rate of cells in each group. The cells were seeded into a 24-well plate with 2 × 103 cells/well. After culturing for 0 and 48 h, 20 μL of CCK-8 solution was added to each well and incubated for an additional 1 h. The absorbance of each well was measured at a wavelength of 450 nm.
-
Cells at the logarithmic growth stage were seeded into 24-well plates at a density of 3 × 104 cells/well. The next day, miR-378a-3p mimics or NCs were transfected according to the aforementioned method. After transfection for 24 h, the cells were collected and counted using a hemocytometer. The cells in each sample were seeded into 6-cm plates at a density of 1,500 cells/well. The cells were cultured for 10 days, and the culture was terminated after the appearance of the visible clone. Colonies were then fixed with 4% paraformaldehyde for 15 min, stained with crystal violet dye solution for 5 min, washed, dried, photographed, and recorded.
-
The scratch wound healing assay was used to determine the cell migration ability. When all of the cells reached confluence, lines were drawn with a 20 μL sterile tip spear, and the repair process was observed under the microscope at 0 h and after 48 h. The migration rate was calculated using the following formula: Migration rate (%) = (Area at 0 h − Area at 48 h)/(Area at 0 h) × 100%. The migration rate is proportional to the cell migration ability.
-
HeLa cells were infected with lentivirus-packaged miR-378a-3p mimics or NCs. Female BALB/C nude mice aged 4–6 weeks were used for the in vivo study. The infected cells were inoculated subcutaneously into the flanks of nude mice. All mice were sacrificed 8 weeks later. The tumors were removed under aseptic conditions, weighed, photographed, and the tumor volume was calculated. The experimental procedures involving animals were performed with the approval of the Ethics Committee of the Hospital of Integrated Chinese and Western Medicine. All efforts were made to reduce the suffering of animals in accordance with the Guidelines for the Care and Use of Laboratory Animals (Ministry of Science and Technology of China, 2006).
-
SPSS Statistics 19.0 software (IBM Corporation, Armonk, NY, USA) was used for statistical analysis. The Student’s t-test was used for comparisons between the two groups. Differences with P < 0.05 were considered to be statistically significant. For the survival curve, the data were analyzed using the log-rank test.
-
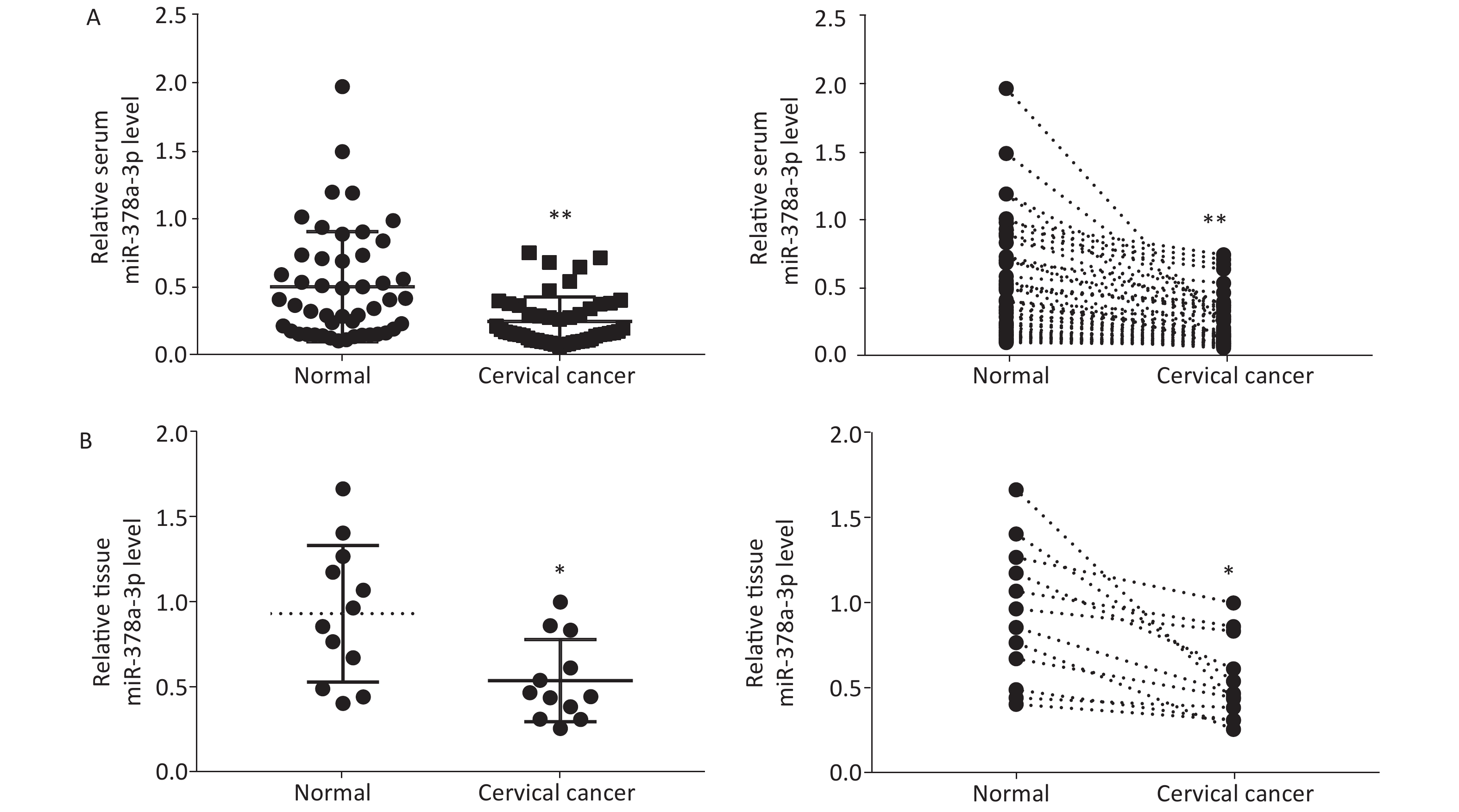
To investigatethe expression of miR-378a-3p in CC cells, we performed RT-qPCR analysis to determine the relative miR-378a-3p levels in the serum of 48 pairs of patients with CC and healthy controls. As shown in Figure 1A, serum miR-378a-3p expression was decreased in CC patients compared to that in normal subjects. Similarly, the analysis of the expression of miR-378a-3p in tissues from CC patients and their adjacent normal tissues revealed that miR-378a-3p expression was markedly decreased in the CC group (Figure 1B).
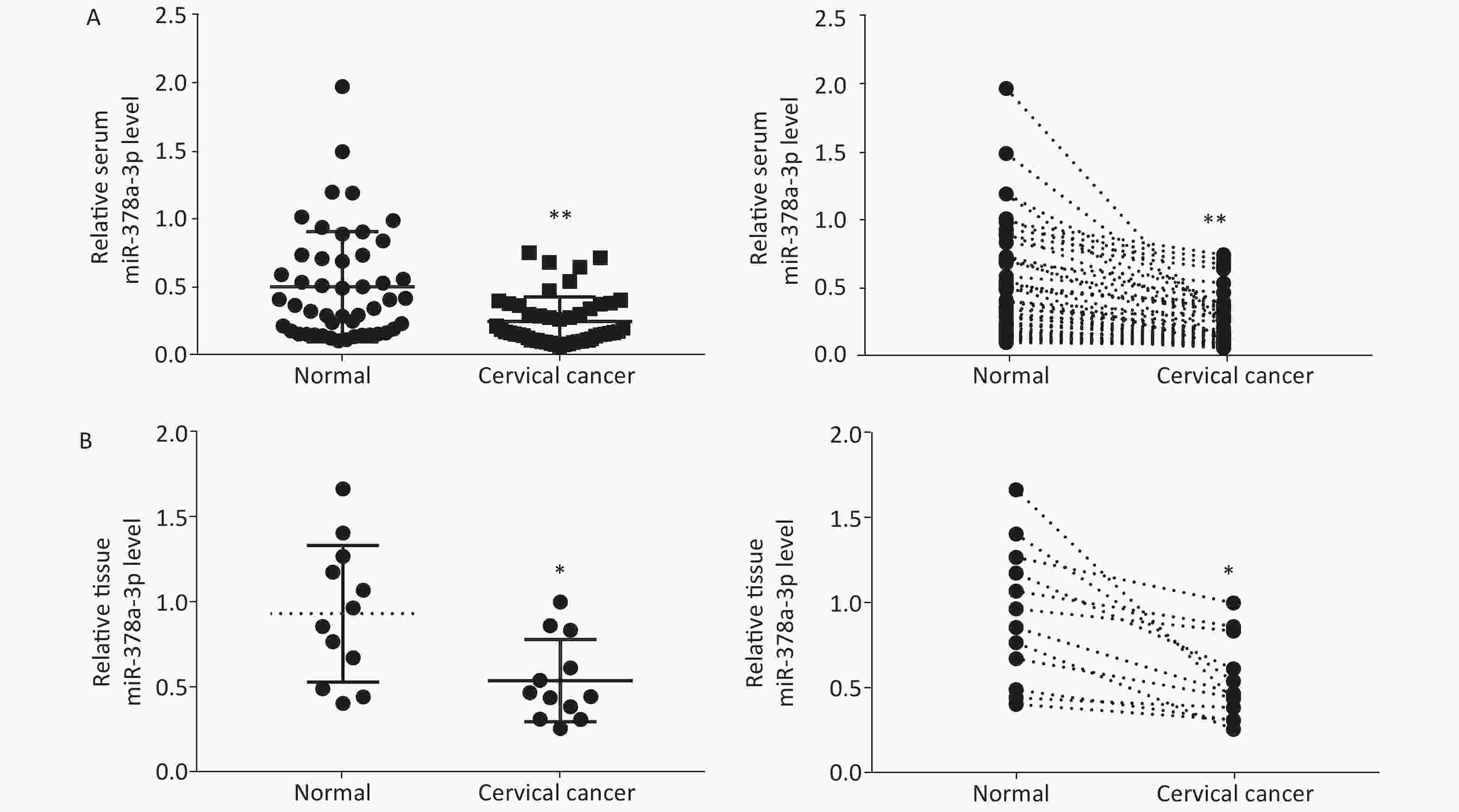
Figure 1. MiR-378a-3p is downregulated in cervical cancer (CC) serum and tissues. (A) Real-time quantitative polymerase chain reaction analysis was performed to evaluate the relative expression levels of miR-378a-3p in 48 tissue sample pairs of patients with CC and healthy control subjects. (B) The expression levels of miR-378a-3p in CC tissues and adjacent normal tissues were measured. *P < 0.05 and **P < 0.01 versus the control group indicate that the difference was significant.
-
The relationship between CC clinicopathological features and serummiR-378a-3p expression shown in Table 1 revealed that miR-378a-3p expression was closely related to tumor size, tumor-node-metastasis (TNM) stage, and lymph node metastasis. For the CC patient samples, patients with low miR-378a-3p expression had larger tumor sizes; more tumors in stages II, III, and IV; and lymph node metastasis. However, miR-378a-3p expression was not associated with age or pathological grading.
Table 1. Relevance between serum miR-378a-3p expression and clinical pathologic characteristics of cervical cancer
Variable No. of patients (n = 48) miR-378a-3p expression χ2 P High (n = 19) Low (n = 29) Age (years) ≤ 50 13 5 8 0.009 0.923 > 50 35 14 21 Tumor size (cm) ≤ 4 20 12 8 5.976 0.015 > 4 28 7 21 TNM stage I–II 18 11 7 5.581 0.018 III–IV 30 8 22 Pathological grading Well (I) 3 3 0 5.141 0.077 Moderate (II–III) 34 14 20 Poor (IV) 11 3 8 Lymph node metastasis Negative 23 13 10 5.298 0.022 Positive 25 6 19 -
The relationship between serum miR-378a-3p expression and the overall survival (OS) of patients with CC was analyzed using the Kaplan-Meier method. Patients with CC were separated into high and low miR-378a-3p expression groups. The results showed that low miR-378a-3p expression was related to poor OS in CC patients (Figure 2A). Moreover, the survival analysis of patients, based on miR-378a-3p expression using Kaplan-Meier Plotter visualization software[23] and The Cancer Genome Atlas database, showed that the mortality rates of patients with low miR-378a-3p expression were significantly higher compared to those with high expression (Figure 2B).
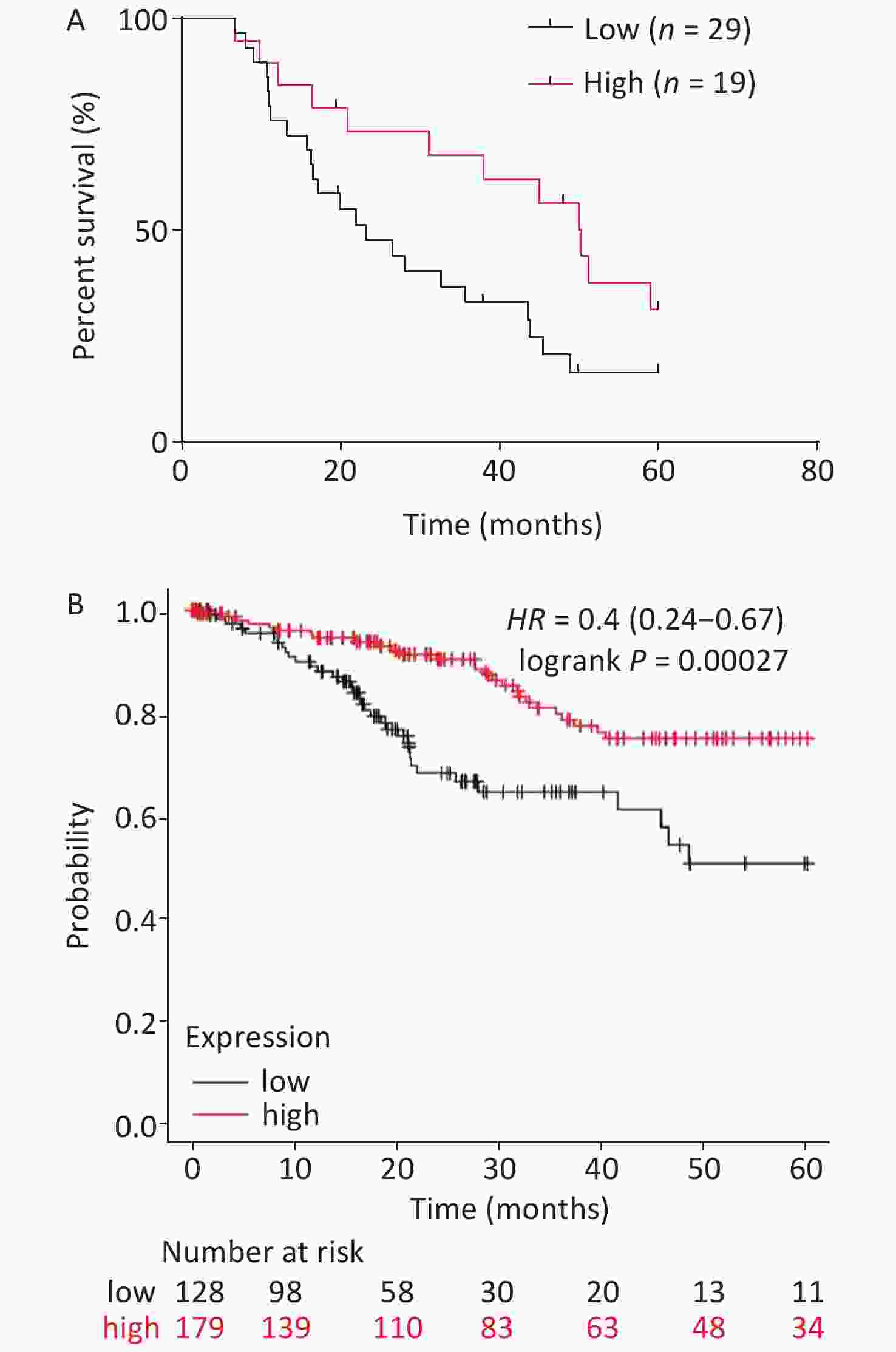
Figure 2. Analysis on the overall survival (OS) of cervical cancer (CC) patients with different miR-378a-3p expression levels. (A) The relationship between serum miR-378a-3p expression and the OS of 48 CC patients was analyzed by the Kaplan-Meier method. (B) OS analysis of miR-378a-3p was performed using Kaplan-Meier Plotter visualization software based on The Cancer Genome Atlas database (N = 306). The figure was downloaded from Kaplan-Meier Plotter.
-
After transfection with miR-378a-3p mimics, CCK-8, cell colony formation, and scratch wound healing assays were used to investigate the effect of miR-378a-3p expression on CC cell proliferation. Compared to the NC group, the viability of SiHa and HeLa cells in the miR-378a-3p mimic group was significantly decreased at 48 h (Figure 3A). The results of the cell colony formation assay revealed that the number and size of the colonies of the transfected SiHa and HeLa cells in the mimic group were suppressed (Figure 3B). In addition, the migration ability was dramatically inhibited after transfection with miR-378a-3p mimics (Figure 3C).
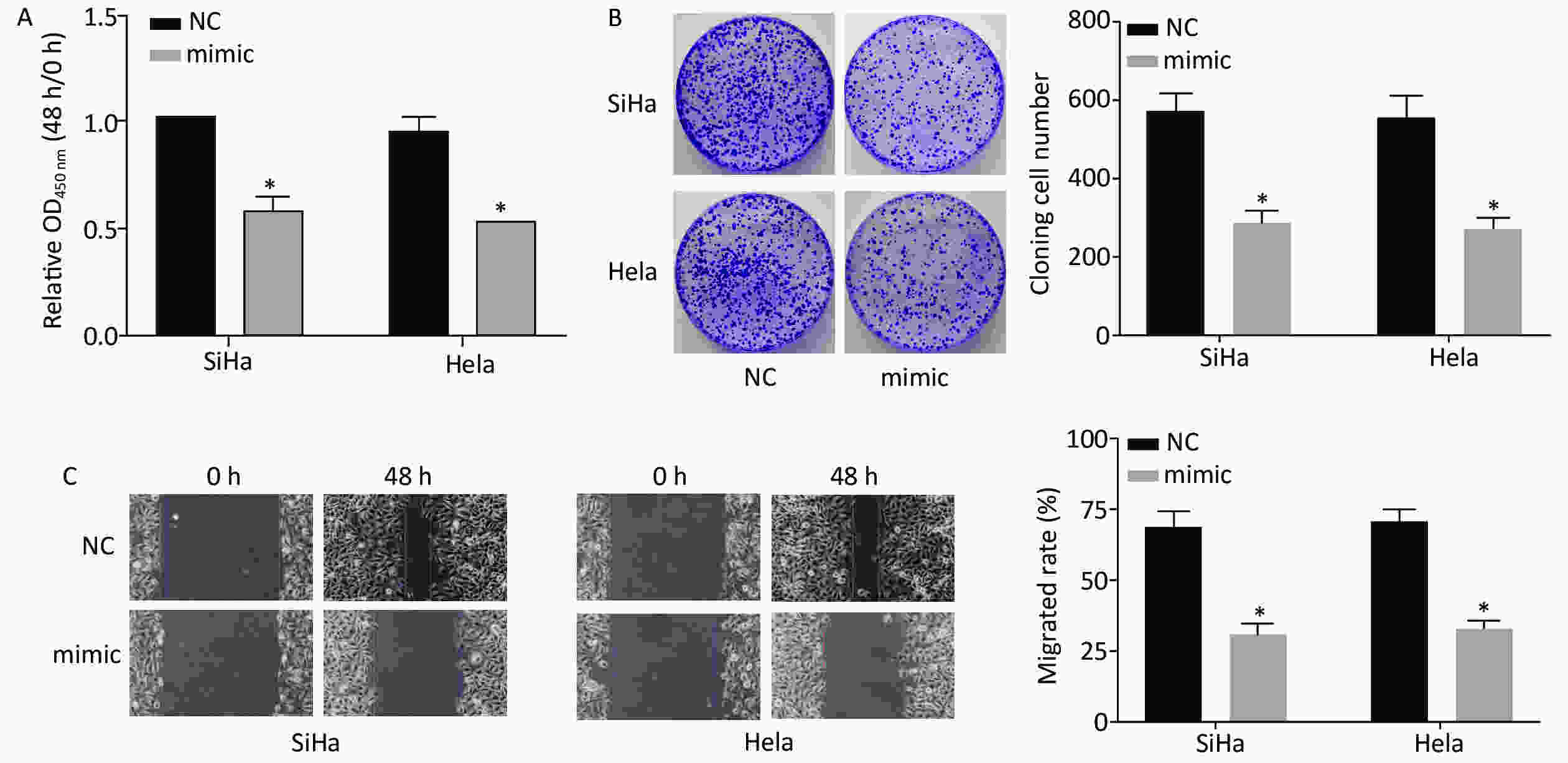
Figure 3. Overexpression of miR-378a-3p inhibits cervical cancer (CC) cell proliferation. (A) The optical density values in SiHa and HeLa cells in the miR-378a-3p mimic and NC groupswere determined by the Cell Counting Kit-8 assay. (B) The cell colony formation assay was performed to assess the proliferation of SiHa and HeLa cells. (C) The wound healing assay was performed to evaluate the migration of SiHa and HeLa cells. *P < 0.05 versus the control group indicate that the difference was significant.
-
Mice were subcutaneously injected with HeLa cells transfected with lentivirus-packaged miR-378a-3p mimics, and the tumor volume was measured every 7 days for 28 days. The results showed that the tumor volumein the miR-378a-3p mimic group was lower than that in the NC group (Figure 4A). We also measured the tumor weight in the two groups, and the results revealed that the tumor weight was dramatically reduced with overexpression of miR-378a-3p (Figure 4B). In addition, after injection of miR-378a-3p mimics, the isolated tumor tissues had a higher level of miR-378a-3p expression (Figure 4C). Moreover, immunohistochemical analysis of Ki-67 expression in isolated tumor tissues showed that Ki-67 expression was significantly decreased in the miR-378a-3p group (Figure 4D).
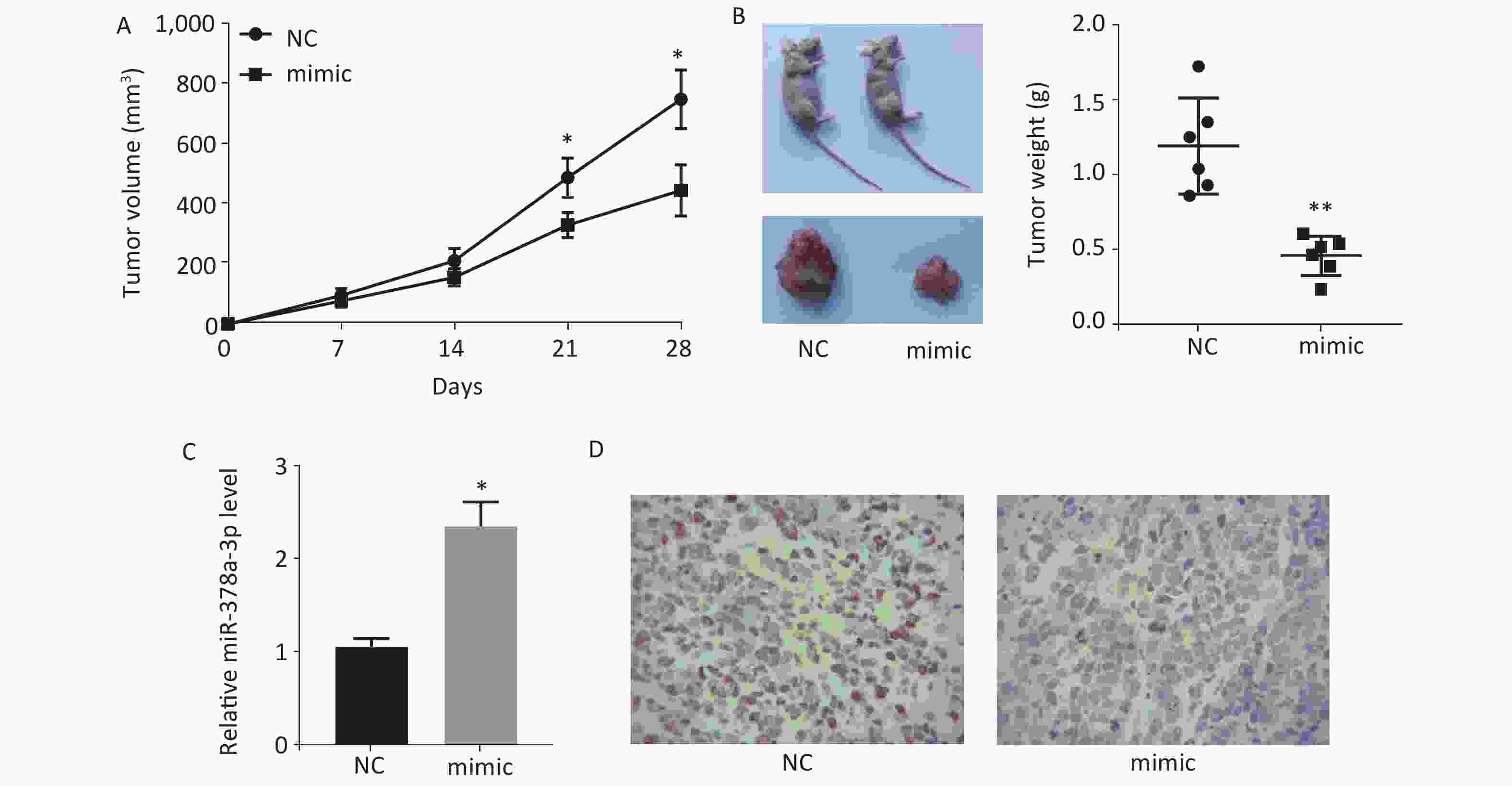
Figure 4. MiR-378a-3p inhibits tumor growth in vivo. (A) Tumor volume was measured every 7 days. (B) Tumors were isolated after 28 days of HeLa cell injection, and images for representative mice and tumors were captured (left). The tumor weight was also calculated (right). (C) The expressions levels of miR-378a-3p in isolated tumors were determined. (D) The Ki-67 expression level was determined by immunohistochemical analysis. *P < 0.05 and **P < 0.01 versus the small interfering RNA negative control group indicate that the difference was significant.
-
The development of CC is a complex pathophysiological process that is associated with dysregulation of the expression of many genes at different stages[24]. Dysregulation of gene expression in normal cervical epithelial cells, which involves numerous abnormalities in gene structure and expression, including coding and non-coding genes as well as regulatory genes, leads to atypical cell growth that in turn leads to carcinoma formation in situ, followed by cancer cell invasion and ultimately extensive spread and metastasis[25]. miRNAs can play a role in promoting or inhibiting various cancers and are involved in the occurrence, development, and progression of tumors. The in-depth study of tumor-associated miRNAs has also led researchers to examine the occurrence, development, and progression of CC from a new perspective. Searching for the specific expression profiles of miRNAs in CC and precancerous lesions and subsequently detecting miRNAs in CC samples presents a new direction in CC research[26, 27]. A study by Zhang et al. showed that miR-320 expression was downregulated in CC tissues compared to normal controls and also found that miR-320 induced tumor cell apoptosis and inhibited cell proliferation, invasion, and tumorigenesis by targeting myeloid cell leukemia 1[28]. Yu et al. used PCR to measure the serum miR-218 level in CC patients and normal control subjects and performed correlation analysis of its expression with the corresponding clinicopathological diagnosis. They found that the expression of miR-218 in CC patients was significantly lower than that in normal subjects, and there was a close correlation between the expression of miR-218 and the metastasis of CC[29]. Fan
et al. found markedly higher miR-20a expression in CC tissues compared to controls, and its expression was related to lymph node metastasis, histological grade, and tumor diameter[30]. MiR-501 was upregulated in CC tissues compared to normal cervical tissues, and it promoted cell proliferation, migration, and invasion in CC[31]. However, the expression levels of miR-378a-3p in CC are poorly characterized. To clarify the role of miR-378a-3p in CC, we first determined the expression levels of miR-378a-3p in the serum of patients with CC and healthy control subjects, and the results showed that miR-378a-3p was downregulated in the CC group. In addition, miR-378a-3p expression in CC tissues was also significantly reduced compared to that in adjacent normal tissues. Most studies on the effect of tumor size on the prognosis of CC have suggested a close relationship between them. This is mainly due to the increase in damage to the body with the increase in tumor size. Lymph node metastasis is one of the most important manifestations of CC. Cancer cells spread to the pelvic cavity and even the whole body through the lymph tissue. Lymph node dissection to excise the lymphatic metastasis is the primary means to reduce the metastasis and recurrence of cancer. In terms of the influence of the tumor stage on the prognosis of CC patients, most studies have concluded that it has an important influence on the prognosis of CC patients. In general, there is a significant correlation between the clinical stage and prognosis of the patient, and the later the stage, the worse the prognosis. Our study found that low expression levels of miR-378a-3p in serum were related to higher tumor stages, larger tumor size, and more lymph node metastasis. These findings indicate that miR-378a-3p may play a key role in CC progression and metastasis.
Recently, many studies have indicated that miRNAs can be used as diagnostic or prognostic biomarkers in many cancers. The expression of miR-378a-3p was downregulated in clinical breast cancer tissues, and lower expression levels of miR-378a-3p were related to poor prognosis in breast cancer patients. In CRC, miR-378a-3p expression was related to histological differentiation and TNM stage. CRC patients with low miR-378a-3p expression had a markedly shorter survival time than those with high expression[22]. We also examined the relationship between miR-378a-3p expression and prognosis of CC, and theresults of the Kaplan-Meier survival curve showed that patients with low miR-378a-3p expression had markedly poor OS.
Uncontrolled cell proliferation is an important characteristic that distinguishes malignant tumor cells from normal cells[32]. To investigate the role of miR-378a-3p in CC cell proliferation, miR-378a-3p mimicsand miR-378a-3p NCs were transfected into SiHa and HeLa cells, and the cell proliferation rate was determined. Our results showed that miR-378a-3p mimics inhibited CC cell proliferation compared to miR-378a-3p NCs. Another feature of malignancies is their metastatic nature. In the later stages of cancer development, tumor cells invade and migrate to surrounding tissues and can be further transferred to other tissues and organs through blood and lymph circulation. Tumor metastasis is the main cause of death in patients with tumors[33]. The ability of tumor cells to migrate is important for tumor cell metastasis. Our study found that overexpression of miR-378a-3p inhibited CC cell migration. In addition, a subcutaneous tumor formation experiment in nude mice was conducted to study the role of miR-378a-3p expression on tumor weight and volume. The results showed that overexpression of miR-378a-3p inhibited tumor growth in mice.
In conclusion, miR-378a-3p was found to be downregulated in CC serum and tissues. Thus, it may serve as an independent prognostic marker for OS in patients with CC, as patients with low expression of miR-378a-3p exhibit poorer prognosis than those with high expression. Additionally, overexpression of miR-378a-3p significantly inhibits the development of CC, indicating that miR-378a-3p is also a potential target for CC treatment.
doi: 10.3967/bes2021.026
MicroRNA-378a-3p Downregulation as a Novel Biomarker with Poor Clinical Outcomes in Cervical Cancer
-
Abstract:
Objective Cervical cancer (CC) is one of the most common malignant tumors in gynecology. This study aimed to investigate the prognostic significance of serum microRNA (miR)-378a-3p in CC and the effect of miR-378a-3p on tumor growth. Methods Real-time quantitative polymerase chain reaction analysis was used to measure the expression of miR-378a-3p in serum from patients with CC and healthy control subjects as well as from CC tissues and adjacent normal tissues. The association between serum miR-378a-3p levels and clinicopathological factors was analyzed. The correlation between miR-378a-3p levels and overall survival (OS) of CC patients was determined by Kaplan-Meier analysis. The CC cell proliferation and migration abilities after transfection of miR-378a-3p mimics were detected by Cell Counting Kit-8 and scratch wound healing assays, respectively. Tumor volume and weight in mice treated with miR-378a-3p were measured using a caliper and an electronic balance. Results MiR-378a-3p expression was downregulated in the serum and tissues of CC patients compared to that in healthy control subjects and normal tissues, respectively. Low expression of miR-378a-3p was positively correlated with large tumor size, advanced tumor stage, and lymph node metastasis. The OS of patients with low expression of miR-378a-3p was significantly lower than that of patients with high expression. Overexpression of miR-378a-3p suppressed the proliferation and migration of CC cells. In vivo studies indicated that overexpression of miR-378a-3p was associated with decreased tumor volume and weight in mice. Conclusion MiR-378a-3p downregulation is associated with the development and prognosis of CC, suggesting that it may be a potential biomarker for CC. -
Key words:
- Cervical cancer /
- miR-378a-3p /
- Biomarker /
- Prognosis /
- Tumor growth
-
Figure 1. MiR-378a-3p is downregulated in cervical cancer (CC) serum and tissues. (A) Real-time quantitative polymerase chain reaction analysis was performed to evaluate the relative expression levels of miR-378a-3p in 48 tissue sample pairs of patients with CC and healthy control subjects. (B) The expression levels of miR-378a-3p in CC tissues and adjacent normal tissues were measured. *P < 0.05 and **P < 0.01 versus the control group indicate that the difference was significant.
Figure 2. Analysis on the overall survival (OS) of cervical cancer (CC) patients with different miR-378a-3p expression levels. (A) The relationship between serum miR-378a-3p expression and the OS of 48 CC patients was analyzed by the Kaplan-Meier method. (B) OS analysis of miR-378a-3p was performed using Kaplan-Meier Plotter visualization software based on The Cancer Genome Atlas database (N = 306). The figure was downloaded from Kaplan-Meier Plotter.
Figure 3. Overexpression of miR-378a-3p inhibits cervical cancer (CC) cell proliferation. (A) The optical density values in SiHa and HeLa cells in the miR-378a-3p mimic and NC groupswere determined by the Cell Counting Kit-8 assay. (B) The cell colony formation assay was performed to assess the proliferation of SiHa and HeLa cells. (C) The wound healing assay was performed to evaluate the migration of SiHa and HeLa cells. *P < 0.05 versus the control group indicate that the difference was significant.
Figure 4. MiR-378a-3p inhibits tumor growth in vivo. (A) Tumor volume was measured every 7 days. (B) Tumors were isolated after 28 days of HeLa cell injection, and images for representative mice and tumors were captured (left). The tumor weight was also calculated (right). (C) The expressions levels of miR-378a-3p in isolated tumors were determined. (D) The Ki-67 expression level was determined by immunohistochemical analysis. *P < 0.05 and **P < 0.01 versus the small interfering RNA negative control group indicate that the difference was significant.
Table 1. Relevance between serum miR-378a-3p expression and clinical pathologic characteristics of cervical cancer
Variable No. of patients (n = 48) miR-378a-3p expression χ2 P High (n = 19) Low (n = 29) Age (years) ≤ 50 13 5 8 0.009 0.923 > 50 35 14 21 Tumor size (cm) ≤ 4 20 12 8 5.976 0.015 > 4 28 7 21 TNM stage I–II 18 11 7 5.581 0.018 III–IV 30 8 22 Pathological grading Well (I) 3 3 0 5.141 0.077 Moderate (II–III) 34 14 20 Poor (IV) 11 3 8 Lymph node metastasis Negative 23 13 10 5.298 0.022 Positive 25 6 19 -
[1] Torre LA, Bray F, Siegel RL, et al. Global cancer statistics, 2012. CA Cancer J Clin, 2015; 65, 87−108. doi: 10.3322/caac.21262 [2] Li HR, Wu XH, Cheng X. Advances in diagnosis and treatment of metastatic cervical cancer. J Gynecol Oncol, 2016; 27, e43. doi: 10.3802/jgo.2016.27.e43 [3] Luo H, Xu XH, Yang J, et al. Genome-wide somatic copy number alteration analysis and database construction for cervical cancer. Mol Genet Genomics, 2020; 295, 765−73. doi: 10.1007/s00438-019-01636-x [4] Balasubramaniam SD, Balakrishnan V, Oon CE, et al. Key Molecular Events in Cervical Cancer Development. Medicina (Kaunas), 2019; 55, 384. doi: 10.3390/medicina55070384 [5] Ghasemi F, Shafiee M, Banikazemi Z, et al. Curcumin inhibits NF-kB and Wnt/β-catenin pathways in cervical cancer cells. Pathol Res Pract, 2019; 215, 152556. doi: 10.1016/j.prp.2019.152556 [6] Shafabakhsh R, Reiter RJ, Mirzaei H, et al. Melatonin: a new inhibitor agent for cervical cancer treatment. J Cell Physiol, 2019; 234, 21670−82. doi: 10.1002/jcp.28865 [7] Chen YH, Hou YY, Yang Y, et al. Gene expression changes in cervical squamous cancers following neoadjuvant interventional chemoembolization. Clin Chim Acta, 2019; 493, 79−86. doi: 10.1016/j.cca.2019.02.012 [8] Aguda BD. Modeling microRNA-transcription factor networks in cancer. Adv Exp Med Biol, 2013; 774, 149−67. [9] Granados López AJ, López JA. Multistep model of cervical cancer: participation of miRNAs and coding genes. Int J Mol Sci, 2014; 15, 15700−33. doi: 10.3390/ijms150915700 [10] Tutar Y. miRNA and cancer; computational and experimental approaches. Curr Pharm Biotechnol, 2014; 15, 429. doi: 10.2174/138920101505140828161335 [11] Rupaimoole R, Slack FJ. MicroRNA therapeutics: towards a new era for the management of cancer and other diseases. Nat Rev Drug Discov, 2017; 16, 203−22. doi: 10.1038/nrd.2016.246 [12] Mirzaei H, Yazdi F, Salehi R, et al. SiRNA and epigenetic aberrations in ovarian cancer. J Can Res Ther, 2016; 12, 498−508. doi: 10.4103/0973-1482.153661 [13] Di Leva G, Garofalo M, Croce CM. MicroRNAs in cancer. Annu Rev Pathol, 2014; 9, 287−314. doi: 10.1146/annurev-pathol-012513-104715 [14] Aghdam AM, Amiri A, Salarinia R, et al. MicroRNAs as diagnostic, prognostic, and therapeutic biomarkers in prostate cancer. Crit Rev Eukaryot Gene Expr, 2019; 29, 127−39. doi: 10.1615/CritRevEukaryotGeneExpr.2019025273 [15] Lin SB, Gregory RI. MicroRNA biogenesis pathways in cancer. Nat Rev Cancer, 2015; 15, 321−33. doi: 10.1038/nrc3932 [16] Savardashtaki A, Shabaninejad Z, Movahedpour A, et al. miRNAs derived from cancer-associated fibroblasts in colorectal cancer. Epigenomics, 2019; 11, 1627−45. doi: 10.2217/epi-2019-0110 [17] Naeli P, Yousefi F, Ghasemi Y, et al. The role of MicroRNAs in lung cancer: implications for diagnosis and therapy. Curr Mol Med, 2020; 20, 90−101. doi: 10.2174/1566524019666191001113511 [18] Khani P, Nasri F, Khani Chamani F, et al. Genetic and epigenetic contribution to astrocytic gliomas pathogenesis. J Neurochem, 2019; 148, 188−203. doi: 10.1111/jnc.14616 [19] Ikeda K, Horie-Inoue K, Ueno T, et al. miR-378a-3p modulates tamoxifen sensitivity in breast cancer MCF-7 cells through targeting GOLT1A. Sci Rep, 2015; 5, 13170. doi: 10.1038/srep13170 [20] Wei XF, Li H, Zhang BW, et al. miR-378a-3p promotes differentiation and inhibits proliferation of myoblasts by targeting HDAC4 in skeletal muscle development. RNA Biol, 2016; 13, 1300−9. doi: 10.1080/15476286.2016.1239008 [21] Xu ZH, Yao TZ, Liu W. miR-378a-3p sensitizes ovarian cancer cells to cisplatin through targeting MAPK1/GRB2. Biomed Pharmacother, 2018; 107, 1410−7. doi: 10.1016/j.biopha.2018.08.132 [22] Li H, Dai SJ, Zhen TT, et al. Clinical and biological significance of miR-378a-3p and miR-378a-5p in colorectal cancer. Eur J Cancer, 2014; 50, 1207−21. doi: 10.1016/j.ejca.2013.12.010 [23] Nagy Á, Lánczky A, Menyhárt O, et al. Author correction: validation of miRNA prognostic power in hepatocellular carcinoma using expression data of independent datasets. Sci Rep, 2018; 8, 11515. doi: 10.1038/s41598-018-27521-y [24] Ditto A, Bogani G, Leone Roberti Maggiore U, et al. Oncologic effectiveness of nerve-sparing radical hysterectomy in cervical cancer. J Gynecol Oncol, 2018; 29, e41. doi: 10.3802/jgo.2018.29.e41 [25] Siegel EM, Riggs BM, Delmas AL, et al. Quantitative DNA methylation analysis of candidate genes in cervical cancer. PLoS One, 2015; 10, e0122495. doi: 10.1371/journal.pone.0122495 [26] Sadri Nahand J, Moghoofei M, Salmaninejad A, et al. Pathogenic role of exosomes and microRNAs in HPV-mediated inflammation and cervical cancer: a review. Int J Cancer, 2020; 146, 305−20. doi: 10.1002/ijc.32688 [27] Nahand JS, Taghizadeh-Boroujeni S, Karimzadeh M, et al. microRNAs: new prognostic, diagnostic, and therapeutic biomarkers in cervical cancer. J Cell Physiol, 2019; 234, 17064−99. doi: 10.1002/jcp.28457 [28] Zhang T, Zou P, Wang TJ, et al. Down-regulation of miR-320 associated with cancer progression and cell apoptosis via targeting Mcl-1 in cervical cancer. Tumor Biol, 2016; 37, 8931−40. doi: 10.1007/s13277-015-4771-6 [29] Yu JJ, Wang Y, Dong RF, et al. Circulating microRNA-218 was reduced in cervical cancer and correlated with tumor invasion. J Cancer Res Clin Oncol, 2012; 138, 671−4. doi: 10.1007/s00432-012-1147-9 [30] Zhao S, Yao D, Chen J, et al. MiR-20a promotes cervical cancer proliferation and metastasis in vitro and in vivo. PLoS One, 2015; 10, e0120905. doi: 10.1371/journal.pone.0120905 [31] Sanches JGP, Xu YC, Yabasin IB, et al. miR-501 is upregulated in cervical cancer and promotes cell proliferation, migration and invasion by targeting CYLD. Chem Biol Interact, 2018; 285, 85−95. doi: 10.1016/j.cbi.2018.02.024 [32] Alsina-Sanchís E, Figueras A, Lahiguera A, et al. TGFβ controls ovarian cancer cell proliferation. Int J Mol Sci, 2017; 18, 1658. doi: 10.3390/ijms18081658 [33] Robert J. Biologie de la métastase biology of cancer metastasis. Bull Cancer, 2013; 100, 333−42. doi: 10.1684/bdc.2013.1724 -




 下载:
下载:



 Quick Links
Quick Links